2007
Beerenwinkel N; Pachter L; Sturmfels B; Elena S F; Lenski R E
Analysis of epistatic interactions and fitness landscapes using a new geometric approach Journal Article
BMC Evolutionary Biology, 7 (1), pp. 60, 2007, ISSN: 14712148.
Abstract | Links | BibTeX | Altmetric | Tags: Descendant Experiments
@article{Beerenwinkel2007,
title = {Analysis of epistatic interactions and fitness landscapes using a new geometric approach},
author = {Niko Beerenwinkel and Lior Pachter and Bernd Sturmfels and Santiago F. Elena and Richard E. Lenski},
url = {http://bmcevolbiol.biomedcentral.com/articles/10.1186/1471-2148-7-60},
doi = {10.1186/1471-2148-7-60},
issn = {14712148},
year = {2007},
date = {2007-01-01},
urldate = {2007-01-01},
journal = {BMC Evolutionary Biology},
volume = {7},
number = {1},
pages = {60},
abstract = {Background
Understanding interactions between mutations and how they affect fitness is a central problem in evolutionary biology that bears on such fundamental issues as the structure of fitness landscapes and the evolution of sex. To date, analyses of fitness landscapes have focused either on the overall directional curvature of the fitness landscape or on the distribution of pairwise interactions. In this paper, we propose and employ a new mathematical approach that allows a more complete description of multi-way interactions and provides new insights into the structure of fitness landscapes.
Results
We apply the mathematical theory of gene interactions developed by Beerenwinkel et al. to a fitness landscape for \textit{Escherichia coli} obtained by Elena and Lenski. The genotypes were constructed by introducing nine mutations into a wild-type strain and constructing a restricted set of 27 double mutants. Despite the absence of mutants higher than second order, our analysis of this genotypic space points to previously unappreciated gene interactions, in addition to the standard pairwise epistasis. Our analysis confirms Elena and Lenski's inference that the fitness landscape is complex, so that an overall measure of curvature obscures a diversity of interaction types. We also demonstrate that some mutations contribute disproportionately to this complexity. In particular, some mutations are systematically better than others at mixing with other mutations. We also find a strong correlation between epistasis and the average fitness loss caused by deleterious mutations. In particular, the epistatic deviations from multiplicative expectations tend toward more positive values in the context of more deleterious mutations, emphasizing that pairwise epistasis is a local property of the fitness landscape. Finally, we determine the geometry of the fitness landscape, which reflects many of these biologically interesting features.
Conclusion
A full description of complex fitness landscapes requires more information than the average curvature or the distribution of independent pairwise interactions. We have proposed a mathematical approach that, in principle, allows a complete description and, in practice, can suggest new insights into the structure of real fitness landscapes. Our analysis emphasizes the value of non-independent genotypes for these inferences.},
keywords = {Descendant Experiments},
pubstate = {published},
tppubtype = {article}
}
Understanding interactions between mutations and how they affect fitness is a central problem in evolutionary biology that bears on such fundamental issues as the structure of fitness landscapes and the evolution of sex. To date, analyses of fitness landscapes have focused either on the overall directional curvature of the fitness landscape or on the distribution of pairwise interactions. In this paper, we propose and employ a new mathematical approach that allows a more complete description of multi-way interactions and provides new insights into the structure of fitness landscapes.
Results
We apply the mathematical theory of gene interactions developed by Beerenwinkel et al. to a fitness landscape for Escherichia coli obtained by Elena and Lenski. The genotypes were constructed by introducing nine mutations into a wild-type strain and constructing a restricted set of 27 double mutants. Despite the absence of mutants higher than second order, our analysis of this genotypic space points to previously unappreciated gene interactions, in addition to the standard pairwise epistasis. Our analysis confirms Elena and Lenski's inference that the fitness landscape is complex, so that an overall measure of curvature obscures a diversity of interaction types. We also demonstrate that some mutations contribute disproportionately to this complexity. In particular, some mutations are systematically better than others at mixing with other mutations. We also find a strong correlation between epistasis and the average fitness loss caused by deleterious mutations. In particular, the epistatic deviations from multiplicative expectations tend toward more positive values in the context of more deleterious mutations, emphasizing that pairwise epistasis is a local property of the fitness landscape. Finally, we determine the geometry of the fitness landscape, which reflects many of these biologically interesting features.
Conclusion
A full description of complex fitness landscapes requires more information than the average curvature or the distribution of independent pairwise interactions. We have proposed a mathematical approach that, in principle, allows a complete description and, in practice, can suggest new insights into the structure of real fitness landscapes. Our analysis emphasizes the value of non-independent genotypes for these inferences.
Cooper T F
Recombination speeds adaptation by reducing competition between beneficial mutations in populations of Escherichia coli Journal Article
PLoS Biology, 5 (9), pp. e225, 2007.
Abstract | Links | BibTeX | Altmetric | Tags: Descendant Experiments, Fitness Trajectories
@article{Cooper2007,
title = {Recombination speeds adaptation by reducing competition between beneficial mutations in populations of \textit{Escherichia coli}},
author = {Tim F. Cooper},
doi = {10.1371/journal.pbio.0050225},
year = {2007},
date = {2007-01-01},
urldate = {2007-01-01},
journal = {PLoS Biology},
volume = {5},
number = {9},
pages = {e225},
abstract = {Identification of the selective forces contributing to the origin and maintenance of sex is a fundamental problem in biology. The Fisher-Muller model proposes that sex is advantageous because it allows beneficial mutations that arise in different lineages to recombine, thereby reducing clonal interference and speeding adaptation. I used the F plasmid to mediate recombination in the bacterium \textit{Escherichia coli} and measured its effect on adaptation at high and low mutation rates. Recombination increased the rate of adaptation approximately 3-fold more in the high mutation rate treatment, where beneficial mutations had to compete for fixation. Sequencing of candidate loci revealed the presence of a beneficial mutation in six high mutation rate lines. In the absence of recombination, this mutation took longer to fix and, over the course of its substitution, conferred a reduced competitive advantage, indicating interference between competing beneficial mutations. Together, these results provide experimental support for the Fisher-Muller model and demonstrate that plasmid-mediated gene transfer can accelerate bacterial adaptation.},
keywords = {Descendant Experiments, Fitness Trajectories},
pubstate = {published},
tppubtype = {article}
}
2006
Cooper T F; Morby A P; Gunn A; Schneider D
Effect of random and hub gene disruptions on environmental and mutational robustness in Escherichia coli Journal Article
BMC Genomics, 7 (1), pp. 237, 2006, ISSN: 1471-2164.
Abstract | Links | BibTeX | Altmetric | Tags: Genotypes and Phenotypes
@article{Cooper2006,
title = {Effect of random and hub gene disruptions on environmental and mutational robustness in \textit{Escherichia coli}},
author = {Tim F. Cooper and Andrew P. Morby and Annabel Gunn and Dominique Schneider},
url = {https://bmcgenomics.biomedcentral.com/articles/10.1186/1471-2164-7-237},
doi = {10.1186/1471-2164-7-237},
issn = {1471-2164},
year = {2006},
date = {2006-12-01},
urldate = {2006-12-01},
journal = {BMC Genomics},
volume = {7},
number = {1},
pages = {237},
abstract = {Background
Genome-wide profiling has allowed the regulatory interaction networks of many organisms to be visualised and the pattern of connections between genes to be studied. These networks are non-random, following a power-law distribution with a small number of well-connected 'hubs' and many genes with only one or a few connections. Theoretical work predicts that power-law networks display several unique properties. One of the most biologically interesting of these is an intrinsic robustness to disturbance such that removal of a random gene will have little effect on network function. Conversely, targeted removal of a hub gene is expected to have a large effect.
Results
We compared the response of \textit{Escherichia coli} to environmental and mutational stress following disruption of random or hub genes. We found that disruption of random genes had less effect on robustness to environmental stress than did the targeted disruption of hub genes. In contrast, random disruption strains were slightly less robust to the effect of mutational stress than were hub disruption strains. When we compared the effect of each disruption on environmental and mutational stress, we found a negative relationship, such that strains that were more environmentally robust tended to be less robust to mutational stress.
Conclusion
Our results demonstrate that mutant strains of \textit{E. coli} respond differently to stress, depending on whether random or hub genes are disrupted. This difference indicates that the power-law distribution of regulatory interactions has biological significance, making random disruptions less deleterious to organisms facing environmental stress. That \textit{E. coli} can reduce the effect of environmental stress without reducing the phenotypic effect of additional mutations, indicates that robustness and evolvability need not be antagonistic.},
keywords = {Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}
Genome-wide profiling has allowed the regulatory interaction networks of many organisms to be visualised and the pattern of connections between genes to be studied. These networks are non-random, following a power-law distribution with a small number of well-connected 'hubs' and many genes with only one or a few connections. Theoretical work predicts that power-law networks display several unique properties. One of the most biologically interesting of these is an intrinsic robustness to disturbance such that removal of a random gene will have little effect on network function. Conversely, targeted removal of a hub gene is expected to have a large effect.
Results
We compared the response of Escherichia coli to environmental and mutational stress following disruption of random or hub genes. We found that disruption of random genes had less effect on robustness to environmental stress than did the targeted disruption of hub genes. In contrast, random disruption strains were slightly less robust to the effect of mutational stress than were hub disruption strains. When we compared the effect of each disruption on environmental and mutational stress, we found a negative relationship, such that strains that were more environmentally robust tended to be less robust to mutational stress.
Conclusion
Our results demonstrate that mutant strains of E. coli respond differently to stress, depending on whether random or hub genes are disrupted. This difference indicates that the power-law distribution of regulatory interactions has biological significance, making random disruptions less deleterious to organisms facing environmental stress. That E. coli can reduce the effect of environmental stress without reducing the phenotypic effect of additional mutations, indicates that robustness and evolvability need not be antagonistic.
Sleight S C; Wigginton N S; Lenski R E
Increased susceptibility to repeated freeze-thaw cycles in Escherichia coli following long-term evolution in a benign environment. Journal Article
BMC evolutionary biology, 6 (February), pp. 104, 2006, ISSN: 1471-2148.
Abstract | Links | BibTeX | Altmetric | Tags: Correlated Responses
@article{Sleight2006,
title = {Increased susceptibility to repeated freeze-thaw cycles in \textit{Escherichia coli} following long-term evolution in a benign environment.},
author = {Sean C. Sleight and Nicholas S. Wigginton and Richard E. Lenski},
url = {http://www.ncbi.nlm.nih.gov/pubmed/17147797
http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=PMC1698501},
doi = {10.1186/1471-2148-6-104},
issn = {1471-2148},
year = {2006},
date = {2006-12-01},
urldate = {2006-12-01},
journal = {BMC evolutionary biology},
volume = {6},
number = {February},
pages = {104},
abstract = {Background
In order to study the dynamics of evolutionary change, 12 populations of \textit{E. coli} B were serially propagated for 20,000 generations in minimal glucose medium at constant 37 degrees C. Correlated changes in various other traits have been previously associated with the improvement in competitive fitness in the selective environment. This study examines whether these evolved lines changed in their ability to tolerate the stresses of prolonged freezing and repeated freeze-thaw cycles during adaptation to a benign environment.
Results
All 12 lines that evolved in the benign environment for 20,000 generations are more sensitive to freeze-thaw cycles than their ancestor. The evolved lines have an average mortality rate of 54% per daily cycle, compared to the ancestral rate of 34%. By contrast, there was no significant difference between the evolved lines and their ancestor in mortality during prolonged freezing. There was also some variability among the evolved lines in susceptibility to repeated freeze-thaw cycles. Those lines that had evolved higher competitive fitness in the minimal glucose medium at 37 degrees C also had higher mortality during freeze-thaw cycles. This variability was not associated, however, with differences among lines in DNA repair functionality and mutability.
Conclusion
The consistency of the evolutionary declines in freeze-thaw tolerance, the correlation between fitness in glucose medium at 37 degrees C and mortality during freeze-thaw cycles, and the absence of greater declines in freeze-thaw survival among the hypermutable lines all indicate a trade-off between performance in minimal glucose medium at 37 degrees C and the capacity to tolerate this stress. Analyses of the mutations that enhance fitness at 37 degrees C may shed light on the physiological basis of this trade-off.},
keywords = {Correlated Responses},
pubstate = {published},
tppubtype = {article}
}
In order to study the dynamics of evolutionary change, 12 populations of E. coli B were serially propagated for 20,000 generations in minimal glucose medium at constant 37 degrees C. Correlated changes in various other traits have been previously associated with the improvement in competitive fitness in the selective environment. This study examines whether these evolved lines changed in their ability to tolerate the stresses of prolonged freezing and repeated freeze-thaw cycles during adaptation to a benign environment.
Results
All 12 lines that evolved in the benign environment for 20,000 generations are more sensitive to freeze-thaw cycles than their ancestor. The evolved lines have an average mortality rate of 54% per daily cycle, compared to the ancestral rate of 34%. By contrast, there was no significant difference between the evolved lines and their ancestor in mortality during prolonged freezing. There was also some variability among the evolved lines in susceptibility to repeated freeze-thaw cycles. Those lines that had evolved higher competitive fitness in the minimal glucose medium at 37 degrees C also had higher mortality during freeze-thaw cycles. This variability was not associated, however, with differences among lines in DNA repair functionality and mutability.
Conclusion
The consistency of the evolutionary declines in freeze-thaw tolerance, the correlation between fitness in glucose medium at 37 degrees C and mortality during freeze-thaw cycles, and the absence of greater declines in freeze-thaw survival among the hypermutable lines all indicate a trade-off between performance in minimal glucose medium at 37 degrees C and the capacity to tolerate this stress. Analyses of the mutations that enhance fitness at 37 degrees C may shed light on the physiological basis of this trade-off.
Pelosi L; Kühn L; Guetta D; Garin J; Geiselmann J; Lenski R E; Schneider D
Parallel Changes in Global Protein Profiles During Long-Term Experimental Evolution in Escherichia coli Journal Article
Genetics, 173 (4), pp. 1851-1869, 2006, ISSN: 1943-2631.
Abstract | Links | BibTeX | Altmetric | Tags: Genotypes and Phenotypes, Parallelism and Divergence
@article{nokey,
title = {Parallel Changes in Global Protein Profiles During Long-Term Experimental Evolution in \textit{Escherichia coli}},
author = {Ludovic Pelosi and Lauriane Kühn and Dorian Guetta and Jérôme Garin and Johannes Geiselmann and Richard E. Lenski and Dominique Schneider},
url = {https://academic.oup.com/genetics/article/173/4/1851/6061068},
doi = {10.1534/genetics.105.049619},
issn = {1943-2631},
year = {2006},
date = {2006-08-01},
urldate = {2006-08-01},
journal = {Genetics},
volume = {173},
number = {4},
pages = {1851-1869},
abstract = {Twelve populations of \textit{Escherichia coli} evolved in and adapted to a glucose-limited environment from a common ancestor. We used two-dimensional protein electrophoresis to compare two evolved clones, isolated from independently derived populations after 20,000 generations. Exceptional parallelism was detected. We compared the observed changes in protein expression profiles with previously characterized global transcription profiles of the same clones; this is the first time such a comparison has been made in an evolutionary context where these changes are often quite subtle. The two methodologies exhibited some remarkable similarities that highlighted two different levels of parallel regulatory changes that were beneficial during the evolution experiment. First, at the higher level, both methods revealed extensive parallel changes in the same global regulatory network, reflecting the involvement of beneficial mutations in genes that control the ppGpp regulon. Second, both methods detected expression changes of identical gene sets that reflected parallel changes at a lower level of gene regulation. The protein profiles led to the discovery of beneficial mutations affecting the \textit{malT} gene, with strong genetic parallelism across independently evolved populations. Functional and evolutionary analyses of these mutations revealed parallel phenotypic decreases in the maltose regulon expression and a high level of polymorphism at this locus in the evolved populations.},
keywords = {Genotypes and Phenotypes, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
Woods R J; Schneider D; Winkworth C L; Riley M A; Lenski R E
Tests of parallel molecular evolution in a long-term experiment with Escherichia coli Journal Article
Proceedings of the National Academy of Sciences of the United States of America, 103 (24), pp. 9107–9112, 2006, ISSN: 0027-8424.
Abstract | Links | BibTeX | Altmetric | Tags: Genome Evolution, Genotypes and Phenotypes, Parallelism and Divergence
@article{Woods2006,
title = {Tests of parallel molecular evolution in a long-term experiment with \textit{Escherichia coli}},
author = {Robert J. Woods and Dominique Schneider and Cynthia L. Winkworth and Margaret A. Riley and Richard E. Lenski},
url = {http://www.pnas.org/cgi/doi/10.1073/pnas.0602917103},
doi = {10.1073/pnas.0602917103},
issn = {0027-8424},
year = {2006},
date = {2006-06-01},
urldate = {2006-06-01},
journal = {Proceedings of the National Academy of Sciences of the United States of America},
volume = {103},
number = {24},
pages = {9107--9112},
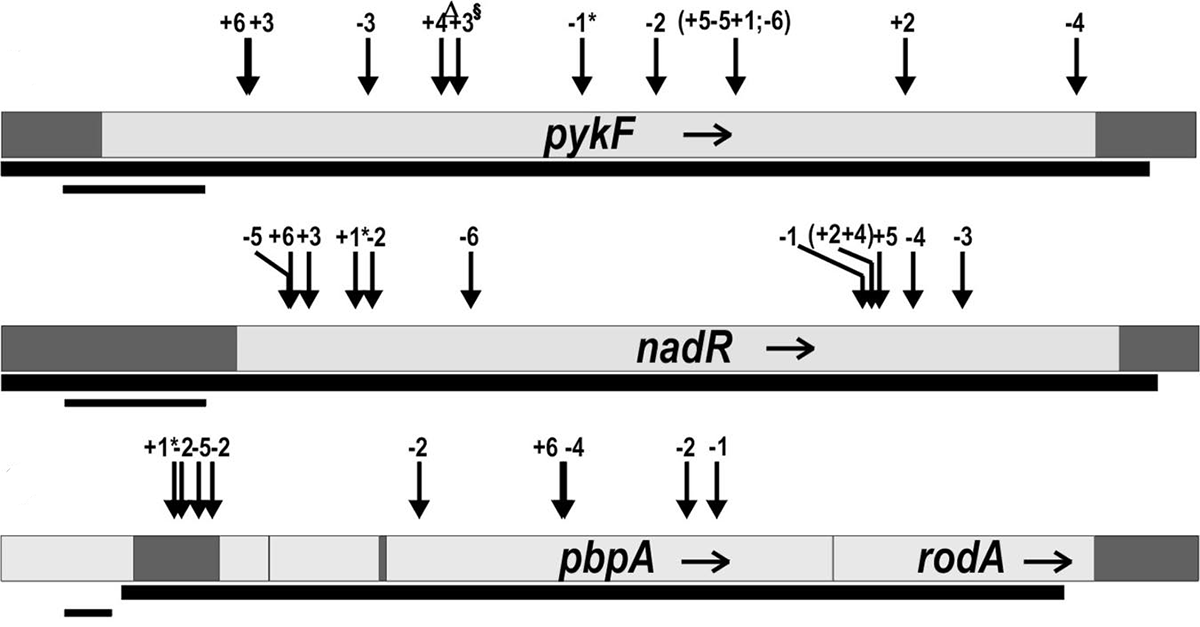
abstract = {The repeatability of evolutionary change is difficult to quantify because only a single outcome can usually be observed for any precise set of circumstances. In this study, however, we have quantified the frequency of parallel and divergent genetic changes in 12 initially identical populations of \textit{Escherichia coli} that evolved in identical environments for 20,000 cell generations. Unlike previous analyses in which candidate genes were identified based on parallel phenotypic changes, here we sequenced four loci (\textit{pykF}, \textit{nadR}, \textit{pbpA}-\textit{rodA}, and \textit{hokB}/\textit{sokB}) in which mutations of unknown effect had been discovered in one population, and then we compared the substitution pattern in these "blind" candidate genes with the pattern found in 36 randomly chosen genes. Two candidate genes, \textit{pykF} and \textit{nadR}, had substitutions in all 11 other populations, and the other 2 in several populations. There were very few cases, however, in which the exact same mutations were substituted, in contrast to the findings from conceptually related work performed with evolving virus populations. No random genes had any substitutions except in four populations that evolved defects in DNA repair. Tests of four different statistical aspects of the pattern of molecular evolution all indicate that adaptation by natural selection drove the parallel changes in these candidate genes.},
keywords = {Genome Evolution, Genotypes and Phenotypes, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
Novak M; Pfeiffer T; Lenski R E; Sauer U; Bonhoeffer S
Experimental Tests for an Evolutionary Trade-Off between Growth Rate and Yield in E. coli Journal Article
The American naturalist, 168 , pp. 242-51, 2006.
Abstract | Links | BibTeX | Altmetric | Tags: Demography and Ecology
@article{articled,
title = {Experimental Tests for an Evolutionary Trade-Off between Growth Rate and Yield in \textit{E. coli}},
author = {Maja Novak and Thomas Pfeiffer and Richard E. Lenski and Uwe Sauer and Sebastian Bonhoeffer},
url = {https://www.journals.uchicago.edu/doi/full/10.1086/506527},
doi = {10.1086/506527},
year = {2006},
date = {2006-01-01},
urldate = {2006-01-01},
journal = {The American naturalist},
volume = {168},
pages = {242-51},
abstract = {Theoretical studies have predicted a trade-off between growth rate and yield in heterotrophic organisms. Here we test for the existence of this trade-off by analyzing the growth characteristics of 12 \textit{E. coli} B populations that evolved for 20,000 generations under a constant selection regime. We performed three different tests. First, we analyzed changes in growth rate and yield over evolutionary time for each population. Second, we tested for a negative correlation between rate and yield across the 12 populations. Finally, we isolated clones from four selected populations and tested for a negative correlation between rate and yield within these populations. We did not find evidence for a trade-off based on the first two tests. However, we did observe a trade-off based on the within-population correlation of yield and rate. Our results indicate that, at least for the populations studied here, an analysis of the within-population diversity might be the most sensitive test for the existence of a trade-off. The observation of a trade-off within, but not between, populations suggests that the populations evolved different genetic solutions for growth in the selective environment, which in turn led to different physiological constraints.},
keywords = {Demography and Ecology},
pubstate = {published},
tppubtype = {article}
}
2005
Ostrowski E A; Rozen D E; Lenski R E
Pleiotropic effects of beneficial mutations in Escherichia coli Journal Article
Evolution, 59 (11), pp. 2343–2352, 2005, ISSN: 0014-3820.
Abstract | Links | BibTeX | Altmetric | Tags: Correlated Responses, Descendant Experiments
@article{Ostrowski2005,
title = {Pleiotropic effects of beneficial mutations in \textit{Escherichia coli}},
author = {Elizabeth A. Ostrowski and Daniel E. Rozen and Richard E. Lenski},
url = {https://onlinelibrary.wiley.com/doi/10.1111/j.0014-3820.2005.tb00944.x},
doi = {10.1111/j.0014-3820.2005.tb00944.x},
issn = {0014-3820},
year = {2005},
date = {2005-11-01},
urldate = {2005-11-01},
journal = {Evolution},
volume = {59},
number = {11},
pages = {2343--2352},
abstract = {Micromutational models of adaptation have placed considerable weight on antagonistic pleiotropy as a mechanism that prevents mutations of large effect from achieving fixation. However, there are few empirical studies of the distribution of pleiotropic effects, and no studies that have examined this distribution for a large number of adaptive mutations. Here we examine the form and extent of pleiotropy associated with beneficial mutations in \textit{Escherichia coli}. To do so, we used a collection of independently evolved genotypes, each of which contains a beneficial mutation that confers increased fitness in a glucose-limited environment. To determine the pleiotropic effects of these mutations, we examined the fitnesses of the mutants in five novel resource environments. Our results show that the majority of mutations had significant fitness effects in alternative resources, such that pleiotropy was common. The predominant form of this pleiotropy was positive - that is, most mutations that conferred increased fitness in glucose also conferred increased fitness in novel resources. We did detect some deleterious pleiotropic effects, but they were primarily limited to one of the five resources, and within this resource, to only a subset of mutants. Although pleiotropic effects were generally positive, fitness levels were lower and more variable on resources that differed most in their mechanisms of uptake and catabolism from that of glucose. Positive pleiotropic effects were strongly correlated in magnitude with their direct effects, but no such correlation was found among mutants with deleterious pleiotropic effects. Whereas previous studies of populations evolved on glucose for longer periods of time showed consistent declines on some of the resources used here, our results suggest that deleterious pleiotropic effects were limited to only a subset of the beneficial mutations available. },
keywords = {Correlated Responses, Descendant Experiments},
pubstate = {published},
tppubtype = {article}
}
Rozen D E; Schneider D; Lenski R E
Long-Term Experimental Evolution in Escherichia coli. XIII. Phylogenetic History of a Balanced Polymorphism Journal Article
Journal of Molecular Evolution, 61 (2), pp. 171–180, 2005, ISSN: 0022-2844.
Abstract | Links | BibTeX | Altmetric | Tags: Demography and Ecology, Genome Evolution
@article{Rozen2005,
title = {Long-Term Experimental Evolution in \textit{Escherichia coli}. XIII. Phylogenetic History of a Balanced Polymorphism},
author = {Daniel E. Rozen and Dominique Schneider and Richard E. Lenski},
url = {http://link.springer.com/10.1007/s00239-004-0322-2},
doi = {10.1007/s00239-004-0322-2},
issn = {0022-2844},
year = {2005},
date = {2005-08-01},
urldate = {2005-08-01},
journal = {Journal of Molecular Evolution},
volume = {61},
number = {2},
pages = {171--180},
abstract = {We investigated the phylogenetic history of a balanced polymorphism that evolved in an experimental population of \textit{Escherichia coli}. Previous work showed that two ecologically and morphologically distinct types, designated L (large) and S (small), arose by generation 6000 and coexisted for more than 12,000 generations thereafter. Here, we performed RFLP analyses using Insertion Sequence elements to resolve the phylogenetic history of L and S. Specifically, we sought to determine whether the derived S morph was monophyletic, indicating a long history of coexistence with L or, alternatively, S was repeatedly regenerated from L, indicating a series of periods with only transiently stable coexistence. Phylogenetic analysis of some 200 clones collected throughout the history of this population demonstrates that S is monophyletic. We then performed competition assays using clones of both morphs from different generations to determine whether either or both lineages continued to undergo genetic adaptation. Indeed, both lineages continued to adapt, and their continued evolution contributed to fluctuations in their relative abundance over evolutionary time. Based on their phylogenetic history and independent evolutionary trajectories, S and L fulfill Cohan's criteria for being different asexual species. },
keywords = {Demography and Ecology, Genome Evolution},
pubstate = {published},
tppubtype = {article}
}
Crozat E; Philippe N; Lenski R E; Geiselmann J; Schneider D
Long-Term Experimental Evolution in Escherichia coli. XII. DNA Topology as a Key Target of Selection Journal Article
Genetics, 169 (2), pp. 523-532, 2005, ISSN: 1943-2631.
Abstract | Links | BibTeX | Altmetric | Tags: Genotypes and Phenotypes, Parallelism and Divergence
@article{nokey,
title = {Long-Term Experimental Evolution in \textit{Escherichia coli}. XII. DNA Topology as a Key Target of Selection},
author = {Estelle Crozat and Nadège Philippe and Richard E. Lenski and Johannes Geiselmann and Dominique Schneider},
url = {https://academic.oup.com/genetics/article/169/2/523/6060318},
doi = {10.1534/genetics.104.035717},
issn = {1943-2631},
year = {2005},
date = {2005-02-01},
urldate = {2005-02-01},
journal = {Genetics},
volume = {169},
number = {2},
pages = {523-532},
abstract = {The genetic bases of adaptation are being investigated in 12 populations of \textit{Escherichia coli}, founded from a common ancestor and serially propagated for 20,000 generations, during which time they achieved substantial fitness gains. Each day, populations alternated between active growth and nutrient exhaustion. DNA supercoiling in bacteria is influenced by nutritional state, and DNA topology helps coordinate the overall pattern of gene expression in response to environmental changes. We therefore examined whether the genetic controls over supercoiling might have changed during the evolution experiment. Parallel changes in topology occurred in most populations, with the level of DNA supercoiling increasing, usually in the first 2000 generations. Two mutations in the \textit{topA} and \textit{fis} genes that control supercoiling were discovered in a population that served as the focus for further investigation. Moving the mutations, alone and in combination, into the ancestral background had an additive effect on supercoiling, and together they reproduced the net change in DNA topology observed in this population. Moreover, both mutations were beneficial in competition experiments. Clonal interference involving other beneficial DNA topology mutations was also detected. These findings define a new class of fitness-enhancing mutations and indicate that the control of DNA supercoiling can be a key target of selection in evolving bacterial populations.},
keywords = {Genotypes and Phenotypes, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
2004
Schneider D; Lenski R E
Dynamics of insertion sequence elements during experimental evolution of bacteria Journal Article
Research in Microbiology, 155 (5), pp. 319–327, 2004, ISSN: 09232508.
Abstract | Links | BibTeX | Altmetric | Tags: Review Articles
@article{Schneider2004,
title = {Dynamics of insertion sequence elements during experimental evolution of bacteria},
author = {Dominique Schneider and Richard E. Lenski},
url = {https://linkinghub.elsevier.com/retrieve/pii/S0923250804000671},
doi = {10.1016/j.resmic.2003.12.008},
issn = {09232508},
year = {2004},
date = {2004-06-01},
urldate = {2004-06-01},
journal = {Research in Microbiology},
volume = {155},
number = {5},
pages = {319--327},
abstract = {We review the intersection between two areas of microbial evolution that were research foci of Michel Blot. One focus is the behavior of insertion sequence (IS) elements, including their role in promoting the evolutionary adaptation of their hosts. The other focus is experimental evolution, an approach that allows the dynamics of genomic and phenotypic change to be observed in the laboratory. This review shows that IS elements are useful as markers for detecting genomic change over experimental time scales and, moreover, that IS elements generate some of the beneficial mutations that increase organismal fitness. },
keywords = {Review Articles},
pubstate = {published},
tppubtype = {article}
}
Remold S K; Lenski R E
Pervasive joint influence of epistasis and plasticity on mutational effects in Escherichia coli Journal Article
Nature Genetics, 36 (4), pp. 423–426, 2004, ISSN: 1061-4036.
Abstract | Links | BibTeX | Altmetric | Tags: Descendant Experiments
@article{Remold2004,
title = {Pervasive joint influence of epistasis and plasticity on mutational effects in \textit{Escherichia coli}},
author = {Susanna K. Remold and Richard E. Lenski},
url = {http://www.nature.com/articles/ng1324},
doi = {10.1038/ng1324},
issn = {1061-4036},
year = {2004},
date = {2004-04-01},
urldate = {2004-04-01},
journal = {Nature Genetics},
volume = {36},
number = {4},
pages = {423--426},
abstract = {The effects of mutations on phenotype and fitness may depend on the environment (phenotypic plasticity), other mutations (genetic epistasis) or both. Here we examine the fitness effects of 18 random insertion mutations in \textit{E. coli} in two resource environments and five genetic backgrounds. We tested each mutation for plasticity and epistasis by comparing its fitness effects across these ecological and genetic contexts. Some mutations had no measurable effect in any of these contexts. None of the mutations had effects on phenotypic plasticity that were independent of genetic background. However, half the mutations had epistatic interactions such that their effects differed among genetic backgrounds, usually in an environment-dependent manner. Also, the pattern of mutational effects across backgrounds indicated that epistasis had been shaped primarily by unique events in the evolutionary history of a population rather than by repeatable events associated with shared environmental history.},
keywords = {Descendant Experiments},
pubstate = {published},
tppubtype = {article}
}
Lenski R E
Phenotypic and genomic evolution during a 20,000-generation experiment with the bacterium Escherichia coli. Incollection
Plant Breeding Reviews, pp. 225 – 265, 2004.
Abstract | Links | BibTeX | Altmetric | Tags: Review Articles
@incollection{Lenski2004,
title = {Phenotypic and genomic evolution during a 20,000-generation experiment with the bacterium \textit{Escherichia coli}.},
author = {Richard E. Lenski},
url = {https://onlinelibrary.wiley.com/doi/10.1002/9780470650288.ch8},
doi = {http://dx.doi.org/10.1002/9780470650288.ch8},
year = {2004},
date = {2004-01-01},
urldate = {2004-01-01},
booktitle = {Plant Breeding Reviews},
pages = {225 -- 265},
edition = {24},
chapter = {8},
abstract = {This paper reviews another long-term selection experiment, one that is both shorter and longer than the 100 years of selection for oil and protein content in maize that is the main focus of this volume. In 1988, 12 populations of the bacterium \textit{Escherichia coli} were founded from the same strain, and these have been propagated in a simple, defined laboratory environment ever since. Each day, the populations are diluted in fresh medium, undergo about 6.6 generations of binary fission before they exhaust the limiting resource, and then must wait until their “springtime” appears again the next day.
This review summarizes some interesting changes and dynamics, both phenotypic and genomic, that occurred in these populations through generation 20,000. For an annual plant 20,000 generations would of course require some 20,000 years; for humans 20,000 generations would span 400,000 years, assuming an average generation of 20 years. This ability to observe evolution in action over many generations is an obvious benefit of studying bacteria. In fact, the experiment recently passed 30,000 generations, but the bacteria evolve faster than we can study them, and generation 20,000 represents the last milestone at which we systematically studied the populations. Rather than taking the same measurements at intervals, we have repeatedly pushed our analyses in new directions. Thus, some changes were analyzed through 2,000 generations, others through 10,000 generations, and still other changes through 20,000 generations. Another wonderful feature of bacteria for evolutionary research is that we can store entire populations frozen, and resurrect them whenever we wish, such that we can gather more data about earlier generations if we later find other aspects of the evolving bacteria that we want to measure and study.},
keywords = {Review Articles},
pubstate = {published},
tppubtype = {incollection}
}
This review summarizes some interesting changes and dynamics, both phenotypic and genomic, that occurred in these populations through generation 20,000. For an annual plant 20,000 generations would of course require some 20,000 years; for humans 20,000 generations would span 400,000 years, assuming an average generation of 20 years. This ability to observe evolution in action over many generations is an obvious benefit of studying bacteria. In fact, the experiment recently passed 30,000 generations, but the bacteria evolve faster than we can study them, and generation 20,000 represents the last milestone at which we systematically studied the populations. Rather than taking the same measurements at intervals, we have repeatedly pushed our analyses in new directions. Thus, some changes were analyzed through 2,000 generations, others through 10,000 generations, and still other changes through 20,000 generations. Another wonderful feature of bacteria for evolutionary research is that we can store entire populations frozen, and resurrect them whenever we wish, such that we can gather more data about earlier generations if we later find other aspects of the evolving bacteria that we want to measure and study.
2003
Elena S F; Lenski R E
Evolution experiments with microorganisms: the dynamics and genetic bases of adaptation Journal Article
Nature Reviews Genetics, 4 (6), pp. 457–469, 2003, ISSN: 1471-0056.
Abstract | Links | BibTeX | Altmetric | Tags: Review Articles
@article{Elena2003,
title = {Evolution experiments with microorganisms: the dynamics and genetic bases of adaptation},
author = {Santiago F. Elena and Richard E. Lenski},
url = {http://www.nature.com/articles/nrg1088},
doi = {10.1038/nrg1088},
issn = {1471-0056},
year = {2003},
date = {2003-06-01},
urldate = {2003-06-01},
journal = {Nature Reviews Genetics},
volume = {4},
number = {6},
pages = {457--469},
abstract = {Microorganisms have been mutating and evolving on Earth for billions of years. Now, a field of research has developed around the idea of using microorganisms to study evolution in action. Controlled and replicated experiments are using viruses, bacteria and yeast to investigate how their genomes and phenotypic properties evolve over hundreds and even thousands of generations. Here, we examine the dynamics of evolutionary adaptation, the genetic bases of adaptation, tradeoffs and the environmental specificity of adaptation, the origin and evolutionary consequences of mutators, and the process of drift decay in very small populations.},
keywords = {Review Articles},
pubstate = {published},
tppubtype = {article}
}
Riehle M M; Bennett A F; Lenski R E; Long A D
Evolutionary changes in heat-inducible gene expression in lines of Escherichia coli adapted to high temperature Journal Article
Physiological Genomics, 14 (1), pp. 47–58, 2003, ISSN: 1094-8341.
Abstract | Links | BibTeX | Altmetric | Tags: Descendant Experiments
@article{Riehle2003,
title = {Evolutionary changes in heat-inducible gene expression in lines of \textit{Escherichia coli} adapted to high temperature},
author = {Michelle M. Riehle and Albert F. Bennett and Richard E. Lenski and Anthony D. Long},
url = {https://www.physiology.org/doi/10.1152/physiolgenomics.00034.2002},
doi = {10.1152/physiolgenomics.00034.2002},
issn = {1094-8341},
year = {2003},
date = {2003-06-01},
urldate = {2003-06-01},
journal = {Physiological Genomics},
volume = {14},
number = {1},
pages = {47--58},
abstract = {The involvement of heat-inducible genes, including the heat-shock genes, in the acute response to temperature stress is well established. However, their importance in genetic adaptation to long-term temperature stress is less clear. Here we use high-density arrays to examine changes in expression for 35 heat-inducible genes in three independent lines of \textit{Escherichia coli} that evolved at high temperature (41.5°C) for 2,000 generations. These lines exhibited significant changes in heat-inducible gene expression relative to their ancestor, including parallel changes in \textit{fkpA}, \textit{gapA}, and \textit{hslT}. As a group, the heat-inducible genes were significantly more likely than noncandidate genes to have evolved changes in expression. Genes encoding molecular chaperones and ATP-dependent proteases, key components of the cytoplasmic stress response, exhibit relatively little expression change; whereas genes with periplasmic functions exhibit significant expression changes suggesting a key role for the extracytoplasmic stress response in the adaptation to high temperature. Following acclimation at 41.5°C, two of the three lines exhibited significantly improved survival at 50°C, indicating changes in inducible thermotolerance. Thus evolution at high temperature led to significant changes at the molecular level in heat-inducible gene expression and at the organismal level in inducible thermotolerance and fitness.},
keywords = {Descendant Experiments},
pubstate = {published},
tppubtype = {article}
}
Lenski R E; Winkworth C L; Riley M A
Rates of DNA Sequence Evolution in Experimental Populations of Escherichia coli During 20,000 Generations Journal Article
Journal of Molecular Evolution, 56 (4), pp. 498–508, 2003, ISSN: 0022-2844.
Abstract | Links | BibTeX | Altmetric | Tags: Genome Evolution, Mutation Rates
@article{Lenski2003,
title = {Rates of DNA Sequence Evolution in Experimental Populations of \textit{Escherichia coli} During 20,000 Generations},
author = {Richard E. Lenski and Cynthia L. Winkworth and Margaret A. Riley},
url = {http://link.springer.com/10.1007/s00239-002-2423-0},
doi = {10.1007/s00239-002-2423-0},
issn = {0022-2844},
year = {2003},
date = {2003-04-01},
urldate = {2003-04-01},
journal = {Journal of Molecular Evolution},
volume = {56},
number = {4},
pages = {498--508},
abstract = {We examined rates of DNA sequence evolution in 12 populations of \textit{Escherichia coli} propagated in a glucose minimal medium for 20,000 generations. Previous work saw mutations mediated by mobile elements in these populations, but the extent of other genomic changes was not investigated. Four of the populations evolved defects in DNA repair and became mutators. Some 500 bp was sequenced in each of 36 genes for 50 clones, including 2 ancestral variants, 2 clones from each population at generation 10,000, and 2 from each at generation 20,000. Ten mutations were found in total, all point mutations including mostly synonymous substitutions and nonsynonymous polymorphisms; all 10 were found in mutator populations. We compared the observed sequence evolution to predictions based on different scenarios. The number of synonymous substitutions is lower than predicted from measured mutation rates in \textit{E. coli}, but the number is higher than rates based on comparing \textit{E. coli} and \textit{Salmonella} genomes. Extrapolating to the entire genome, these data predict about 250 synonymous substitutions on average per mutator population, but only about 3 synonymous substitutions per nonmutator population, during 20,000 generations. These data illustrate the challenge of finding sequence variation among bacterial isolates that share such a recent ancestor. However, this limited variation also provides a useful baseline for research aimed at finding the beneficial substitutions in these populations.},
keywords = {Genome Evolution, Mutation Rates},
pubstate = {published},
tppubtype = {article}
}
Cooper T F; Rozen D E; Lenski R E
Parallel changes in gene expression after 20,000 generations of evolution in Escherichia coli Journal Article
Proceedings of the National Academy of Sciences of the United States of America, 100 (3), pp. 1072–1077, 2003, ISSN: 0027-8424.
Abstract | Links | BibTeX | Altmetric | Tags: Genotypes and Phenotypes, Parallelism and Divergence
@article{Cooper2003,
title = {Parallel changes in gene expression after 20,000 generations of evolution in \textit{Escherichia coli}},
author = {Tim F. Cooper and Daniel E. Rozen and Richard E. Lenski},
url = {http://www.pnas.org/cgi/doi/10.1073/pnas.0334340100},
doi = {10.1073/pnas.0334340100},
issn = {0027-8424},
year = {2003},
date = {2003-02-01},
urldate = {2003-02-01},
journal = {Proceedings of the National Academy of Sciences of the United States of America},
volume = {100},
number = {3},
pages = {1072--1077},
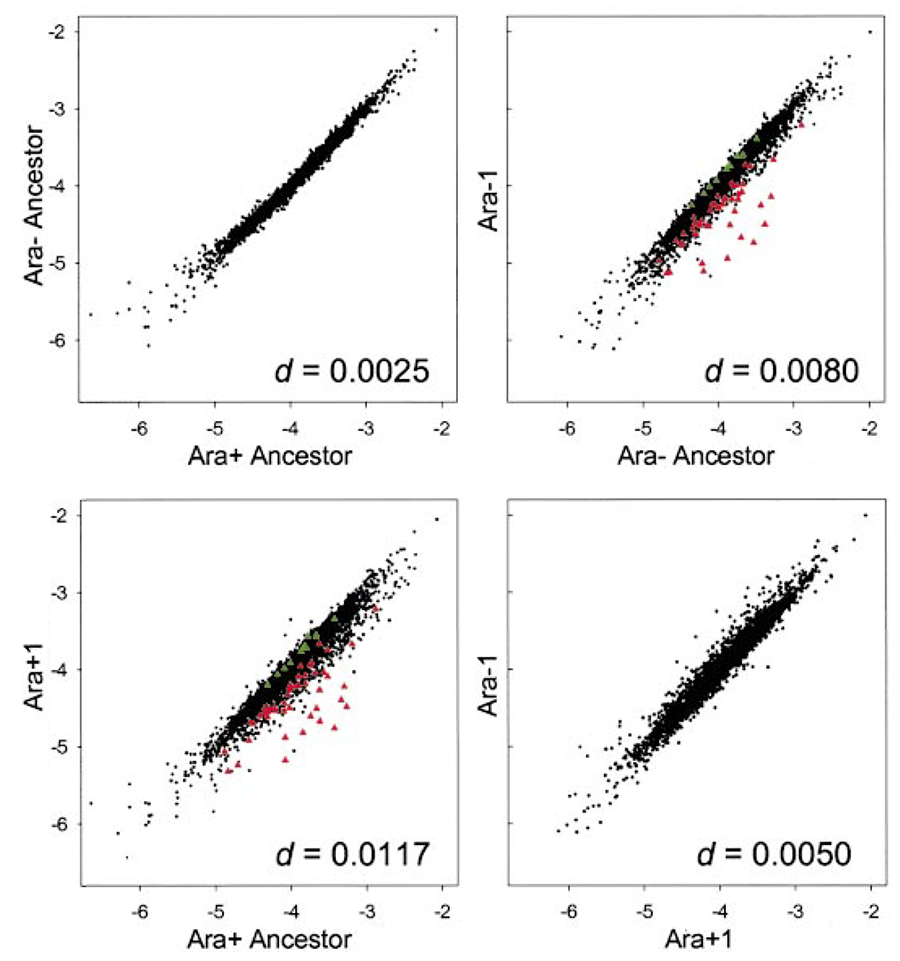
abstract = {Twelve populations of \textit{Escherichia coli}, derived from a common ancestor, evolved in a glucose-limited medium for 20,000 generations. Here we use DNA expression arrays to examine whether gene-expression profiles in two populations evolved in parallel, which would indicate adaptation, and to gain insight into the mechanisms underlying their adaptation. We compared the expression profile of the ancestor to that of clones sampled from both populations after 20,000 generations. The expression of 59 genes had changed significantly in both populations. Remarkably, all 59 were changed in the same direction relative to the ancestor. Many of these genes were members of the cAMP-cAMP receptor protein (CRP) and guanosine tetraphosphate (ppGpp) regulons. Sequencing of several genes controlling the effectors of these regulons found a nonsynonymous mutation in \textit{spoT} in one population. Moving this mutation into the ancestral background showed that it increased fitness and produced many of the expression changes manifest after 20,000 generations. The same mutation had no effect on fitness when introduced into the other evolved population, indicating that a mutation of similar effect was present already. Our study demonstrates the utility of expression arrays for addressing evolutionary issues including the quantitative measurement of parallel evolution in independent lineages and the identification of beneficial mutations.},
keywords = {Genotypes and Phenotypes, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
2002
de Visser J A G M; Lenski R E
Long-term experimental evolution in Escherichia coli. XI. Rejection of non-transitive interactions as cause of declining rate of adaptation. Journal Article
BMC evolutionary biology, 2 (May 2014), pp. 19, 2002, ISSN: 1471-2148.
Abstract | Links | BibTeX | Altmetric | Tags: Fitness Trajectories
@article{DeVisser2002,
title = {Long-term experimental evolution in \textit{Escherichia coli}. XI. Rejection of non-transitive interactions as cause of declining rate of adaptation.},
author = {J. Arjan G. M. {de Visser} and Richard E. Lenski},
url = {http://www.ncbi.nlm.nih.gov/pubmed/12443537
http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=PMC134600},
doi = {10.1186/1471-2148-2-19},
issn = {1471-2148},
year = {2002},
date = {2002-10-01},
urldate = {2002-10-01},
journal = {BMC evolutionary biology},
volume = {2},
number = {May 2014},
pages = {19},
abstract = {Background
Experimental populations of \textit{Escherichia coli} have evolved for 20,000 generations in a uniform environment. Their rate of improvement, as measured in competitions with the ancestor in that environment, has declined substantially over this period. This deceleration has been interpreted as the bacteria approaching a peak or plateau in a fitness landscape. Alternatively, this deceleration might be caused by non-transitive competitive interactions, in particular such that the measured advantage of later genotypes relative to earlier ones would be greater if they competed directly.
Results
To distinguish these two hypotheses, we performed a large set of competitions using one of the evolved lines. Twenty-one samples obtained at 1,000-generation intervals each competed against five genetically marked clones isolated at 5,000-generation intervals, with three-fold replication. The pattern of relative fitness values for these 315 pairwise competitions was compared with expectations under transitive and non-transitive models, the latter structured to produce the observed deceleration in fitness relative to the ancestor. In general, the relative fitness of later and earlier generations measured by direct competition agrees well with the fitness inferred from separately competing each against the ancestor. These data thus support the transitive model.
Conclusion
Non-transitive competitive interactions were not a major feature of evolution in this population. Instead, the pronounced deceleration in its rate of fitness improvement indicates that the population early on incorporated most of those mutations that provided the greatest gains, and subsequently relied on beneficial mutations that were fewer in number, smaller in effect, or both.},
keywords = {Fitness Trajectories},
pubstate = {published},
tppubtype = {article}
}
Experimental populations of Escherichia coli have evolved for 20,000 generations in a uniform environment. Their rate of improvement, as measured in competitions with the ancestor in that environment, has declined substantially over this period. This deceleration has been interpreted as the bacteria approaching a peak or plateau in a fitness landscape. Alternatively, this deceleration might be caused by non-transitive competitive interactions, in particular such that the measured advantage of later genotypes relative to earlier ones would be greater if they competed directly.
Results
To distinguish these two hypotheses, we performed a large set of competitions using one of the evolved lines. Twenty-one samples obtained at 1,000-generation intervals each competed against five genetically marked clones isolated at 5,000-generation intervals, with three-fold replication. The pattern of relative fitness values for these 315 pairwise competitions was compared with expectations under transitive and non-transitive models, the latter structured to produce the observed deceleration in fitness relative to the ancestor. In general, the relative fitness of later and earlier generations measured by direct competition agrees well with the fitness inferred from separately competing each against the ancestor. These data thus support the transitive model.
Conclusion
Non-transitive competitive interactions were not a major feature of evolution in this population. Instead, the pronounced deceleration in its rate of fitness improvement indicates that the population early on incorporated most of those mutations that provided the greatest gains, and subsequently relied on beneficial mutations that were fewer in number, smaller in effect, or both.
Schneider D; Duperchy E; Depeyrot J; Coursange E; Lenski R E; Blot M
Genomic comparisons among Escherichia coli strains B, K-12, and O157:H7 using IS elements as molecular markers. Journal Article
BMC microbiology, 2 , pp. 18, 2002, ISSN: 1471-2180.
Abstract | Links | BibTeX | Altmetric | Tags: Methods and Miscellaneous
@article{Schneider2002,
title = {Genomic comparisons among \textit{Escherichia coli} strains B, K-12, and O157:H7 using IS elements as molecular markers.},
author = {Dominique Schneider and Esther Duperchy and Joelle Depeyrot and Evelyne Coursange and Richard E. Lenski and Michel Blot},
url = {http://www.ncbi.nlm.nih.gov/pubmed/12106505
http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=PMC117601},
doi = {10.1186/1471-2180-2-18},
issn = {1471-2180},
year = {2002},
date = {2002-07-01},
urldate = {2002-07-01},
journal = {BMC microbiology},
volume = {2},
pages = {18},
abstract = {Background
Insertion Sequence (IS) elements are mobile genetic elements widely distributed among bacteria. Their activities cause mutations, promoting genetic diversity and sometimes adaptation. Previous studies have examined their copy number and distribution in \textit{Escherichia coli} K-12 and natural isolates. Here, we map most of the IS elements in \textit{E. coli} B and compare their locations with the published genomes of K-12 and O157:H7.
Results
The genomic locations of IS elements reveal numerous differences between B, K-12, and O157:H7. IS elements occur in \textit{hok-sok} loci (homologous to plasmid stabilization systems) in both B and K-12, whereas these same loci lack IS elements in O157:H7. IS elements in B and K-12 are often found in locations corresponding to O157:H7-specific sequences, which suggests IS involvement in chromosomal rearrangements including the incorporation of foreign DNA. Some sequences specific to B are identified, as reported previously for O157:H7. The extent of nucleotide sequence divergence between B and K-12 is < 2% for most sequences adjacent to IS elements. By contrast, B and K-12 share only a few IS locations besides those in \textit{hok-sok} loci. Several phenotypic features of B are explained by IS elements, including differential porin expression from K-12.
Conclusions
These data reveal a high level of IS activity since \textit{E. coli} B, K-12, and O157:H7 diverged from a common ancestor, including IS association with deletions and incorporation of horizontally acquired genes as well as transpositions. These findings indicate the important role of IS elements in genome plasticity and divergence.},
keywords = {Methods and Miscellaneous},
pubstate = {published},
tppubtype = {article}
}
Insertion Sequence (IS) elements are mobile genetic elements widely distributed among bacteria. Their activities cause mutations, promoting genetic diversity and sometimes adaptation. Previous studies have examined their copy number and distribution in Escherichia coli K-12 and natural isolates. Here, we map most of the IS elements in E. coli B and compare their locations with the published genomes of K-12 and O157:H7.
Results
The genomic locations of IS elements reveal numerous differences between B, K-12, and O157:H7. IS elements occur in hok-sok loci (homologous to plasmid stabilization systems) in both B and K-12, whereas these same loci lack IS elements in O157:H7. IS elements in B and K-12 are often found in locations corresponding to O157:H7-specific sequences, which suggests IS involvement in chromosomal rearrangements including the incorporation of foreign DNA. Some sequences specific to B are identified, as reported previously for O157:H7. The extent of nucleotide sequence divergence between B and K-12 is < 2% for most sequences adjacent to IS elements. By contrast, B and K-12 share only a few IS locations besides those in hok-sok loci. Several phenotypic features of B are explained by IS elements, including differential porin expression from K-12.
Conclusions
These data reveal a high level of IS activity since E. coli B, K-12, and O157:H7 diverged from a common ancestor, including IS association with deletions and incorporation of horizontally acquired genes as well as transpositions. These findings indicate the important role of IS elements in genome plasticity and divergence.
Lenski R E
Experimental evolution: a long-term study with E. coli. Miscellaneous
2002, ISBN: 9780195122008.
Abstract | Links | BibTeX | Altmetric | Tags: Review Articles
@misc{Lenski2002,
title = {Experimental evolution: a long-term study with \textit{E. coli}.},
author = {Richard E. Lenski},
editor = {M. Pagel and S. Frank and C. Godfray and B. K. Hall and K. Hawkes and D. M. Hillis and A. Kodric-Brown and R. E. Lenski and A. Pomiankowski},
url = {https://www.oxfordreference.com/view/10.1093/acref/9780195122008.001.0001/acref-9780195122008-e-137?rskey=hBoe2z&result=128},
doi = {10.1093/acref/9780195122008.001.0001},
isbn = {9780195122008},
year = {2002},
date = {2002-01-01},
urldate = {2002-01-01},
booktitle = {Encyclopedia of Evolution},
pages = {342--344},
publisher = {Oxford University Press},
address = {New York},
abstract = {The paleontologist Stephen Jay Gould envisioned a thought experiment of “replaying life’s tape” to explore the predictability, or repeatability, of evolution. Gould (1989) argued that evolution is not repeatable: “Any replay of the tape would lead evolution down a pathway radically different from the road actually taken ... no finale can be specified at the start, and none would ever occur a second time in the same way, because any pathway proceeds through thousands of improbable stages.” No one can replay evolution on the vast scale imagined by Gould. But on a much smaller scale, the following experiment examines the same issue by monitoring multiple populations, all founded all founded from the same ancestor, as they evolve in identical environments for thousands of generations.},
keywords = {Review Articles},
pubstate = {published},
tppubtype = {misc}
}