2023
Favate J S; Skalenko K S; Chiles E; Su X; Yadavalli S S; Shah P
Linking genotypic and phenotypic changes in the E. coli long-term evolution experiment using metabolomics Journal Article
eLife, vol. 12, pp. RP87039, 2023, ISSN: eLife.
Abstract | Links | BibTeX | Altmetric | Tags: Genotypes and Phenotypes, Parallelism and Divergence
@article{10.7554/eLife.87039,
title = {Linking genotypic and phenotypic changes in the \textit{E. coli} long-term evolution experiment using metabolomics},
author = {John S Favate and Kyle S Skalenko and Eric Chiles and Xiaoyang Su and Srujana Samhita Yadavalli and Premal Shah},
editor = {John McCutcheon and Christian R Landry},
url = {https://doi.org/10.7554/eLife.87039},
doi = {10.7554/eLife.87039},
issn = {eLife},
year = {2023},
date = {2023-11-01},
urldate = {2023-11-01},
journal = {eLife},
volume = {12},
pages = {RP87039},
publisher = {2050-084X},
abstract = {Changes in an organism's environment, genome, or gene expression patterns can lead to changes in its metabolism. The metabolic phenotype can be under selection and contributes to adaptation. However, the networked and convoluted nature of an organism's metabolism makes relating mutations, metabolic changes, and effects on fitness challenging. To overcome this challenge, we use the long-term evolution experiment (LTEE) with \textit{E. coli} as a model to understand how mutations can eventually affect metabolism and perhaps fitness. We used mass spectrometry to broadly survey the metabolomes of the ancestral strains and all 12 evolved lines. We combined this metabolic data with mutation and expression data to suggest how mutations that alter specific reaction pathways, such as the biosynthesis of nicotinamide adenine dinucleotide, might increase fitness in the system. Our work provides a better understanding of how mutations might affect fitness through the metabolic changes in the LTEE and thus provides a major step in developing a complete genotype-phenotype map for this experimental system. },
key = {E. coli; LTEE; adaptation; bacteria; evolutionary biology; metabolomics},
keywords = {Genotypes and Phenotypes, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
2022
Favate J S; Liang S; Cope A L; Yadavilli S S; Shah P
The landscape of transcriptional and translational changes over 22 years of bacterial adaptation Journal Article
eLife, vol. 11, pp. e81979, 2022.
Abstract | Links | BibTeX | Altmetric | Tags: Genotypes and Phenotypes, Parallelism and Divergence
@article{favate2022,
title = {The landscape of transcriptional and translational changes over 22 years of bacterial adaptation},
author = {John S Favate and Shun Liang and Alexander L Cope and Srujana S Yadavilli and Premal Shah},
editor = {Detlef Weigel},
url = {https://elifesciences.org/articles/81979},
doi = {10.7554/eLife.81979},
year = {2022},
date = {2022-10-10},
urldate = {2022-10-10},
journal = {eLife},
volume = {11},
pages = {e81979},
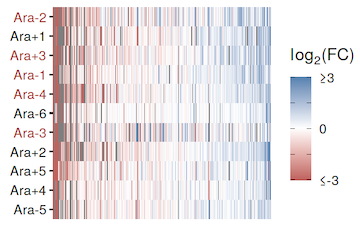
abstract = {Organisms can adapt to an environment by taking multiple mutational paths. This redundancy at the genetic level, where many mutations have similar phenotypic and fitness effects, can make untangling the molecular mechanisms of complex adaptations difficult. Here, we use the \textit{Escherichia coli} long-term evolution experiment (LTEE) as a model to address this challenge. To understand how different genomic changes could lead to parallel fitness gains, we characterize the landscape of transcriptional and translational changes across 12 replicate populations evolving in parallel for 50,000 generations. By quantifying absolute changes in mRNA abundances, we show that not only do all evolved lines have more mRNAs but that this increase in mRNA abundance scales with cell size. We also find that despite few shared mutations at the genetic level, clones from replicate populations in the LTEE are remarkably similar in their gene expression patterns at both the transcriptional and translational levels. Furthermore, we show that the majority of the expression changes are due to changes at the transcriptional level with very few translational changes. Finally, we show how mutations in transcriptional regulators lead to consistent and parallel changes in the expression levels of downstream genes. These results deepen our understanding of the molecular mechanisms underlying complex adaptations and provide insights into the repeatability of evolution.},
howpublished = {eLife},
keywords = {Genotypes and Phenotypes, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
Izutsu M; Lenski R E
Experimental test of the contributions of initial variation and new mutations to adaptive evolution in a novel environment Journal Article
Frontiers in Ecology and Evolution, vol. 10, pp. 958406, 2022, ISSN: 2296-701X.
Abstract | Links | BibTeX | Altmetric | Tags: Descendant Experiments, Historical Contingency, Parallelism and Divergence
@article{izutsu2022,
title = {Experimental test of the contributions of initial variation and new mutations to adaptive evolution in a novel environment},
author = {Minako Izutsu and Richard E. Lenski},
url = {https://www.frontiersin.org/articles/10.3389/fevo.2022.958406/full},
doi = {10.3389/fevo.2022.958406},
issn = {2296-701X},
year = {2022},
date = {2022-10-06},
urldate = {2022-10-01},
journal = {Frontiers in Ecology and Evolution},
volume = {10},
pages = {958406},
abstract = {Experimental evolution is an approach that allows researchers to study organisms as they evolve in controlled environments. Despite the growing popularity of this approach, there are conceptual gaps among projects that use different experimental designs. One such gap concerns the contributions to adaptation of genetic variation present at the start of an experiment and that of new mutations that arise during an experiment. The primary source of genetic variation has historically depended largely on the study organisms. In the long-term evolution experiment (LTEE) using \textit{Escherichia coli}, for example, each population started from a single haploid cell, and therefore, adaptation depended entirely on new mutations. Most other microbial evolution experiments have followed the same strategy. By contrast, evolution experiments using multicellular, sexually reproducing organisms typically start with preexisting variation that fuels the response to selection. New mutations may also come into play in later generations of these experiments, but it is generally difficult to quantify their contribution in these studies. Here, we performed an experiment using \textit{E. coli} to compare the contributions of initial genetic variation and new mutations to adaptation in a new environment. Our experiment had four treatments that varied in their starting diversity, with 18 populations in each treatment. One treatment depended entirely on new mutations, while the other three began with mixtures of clones, whole-population samples, or mixtures of whole-population samples from the LTEE. We tracked a genetic marker associated with different founders in two treatments. These data revealed significant variation in fitness among the founders, and that variation impacted evolution in the early generations of our experiment. However, there were no differences in fitness among the treatments after 500 or 2,000 generations in the new environment, despite the variation in fitness among the founders. These results indicate that new mutations quickly dominated, and eventually they contributed more to adaptation than did the initial variation. Our study thus shows that preexisting genetic variation can have a strong impact on early evolution in a new environment, but new beneficial mutations may contribute more to later evolution and can even drive some initially beneficial variants to extinction.},
keywords = {Descendant Experiments, Historical Contingency, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
2018
Blount Z D; Lenski R E; Losos J B
Contingency and determinism in evolution: Replaying life's tape Journal Article
Science, vol. 362, no. 6415, pp. eaam5979, 2018, ISSN: 0036-8075.
Abstract | Links | BibTeX | Altmetric | Tags: Historical Contingency, Parallelism and Divergence, Review Articles
@article{Blount2018,
title = {Contingency and determinism in evolution: Replaying life's tape},
author = {Zachary D. Blount and Richard E. Lenski and Jonathan B. Losos},
url = {https://www.sciencemag.org/lookup/doi/10.1126/science.aam5979},
doi = {10.1126/science.aam5979},
issn = {0036-8075},
year = {2018},
date = {2018-11-01},
urldate = {2018-11-01},
journal = {Science},
volume = {362},
number = {6415},
pages = {eaam5979},
abstract = {Historical processes display some degree of "contingency," meaning their outcomes are sensitive to seemingly inconsequential events that can fundamentally change the future. Contingency is what makes historical outcomes unpredictable. Unlike many other natural phenomena, evolution is a historical process. Evolutionary change is often driven by the deterministic force of natural selection, but natural selection works upon variation that arises unpredictably through time by random mutation, and even beneficial mutations can be lost by chance through genetic drift. Moreover, evolution has taken place within a planetary environment with a particular history of its own. This tension between determinism and contingency makes evolutionary biology a kind of hybrid between science and history. While philosophers of science examine the nuances of contingency, biologists have performed many empirical studies of evolutionary repeatability and contingency. Here, we review the experimental and comparative evidence from these studies. Replicate populations in evolutionary "replay" experiments often show parallel changes, especially in overall performance, although idiosyncratic outcomes show that the particulars of a lineage's history can affect which of several evolutionary paths is taken. Comparative biologists have found many notable examples of convergent adaptation to similar conditions, but quantification of how frequently such convergence occurs is difficult. On balance, the evidence indicates that evolution tends to be surprisingly repeatable among closely related lineages, but disparate outcomes become more likely as the footprint of history grows deeper. Ongoing research on the structure of adaptive landscapes is providing additional insight into the interplay of fate and chance in the evolutionary process.},
keywords = {Historical Contingency, Parallelism and Divergence, Review Articles},
pubstate = {published},
tppubtype = {article}
}
2017
Good B H; McDonald M J; Barrick J E; Lenski R E; Desai M M
The dynamics of molecular evolution over 60,000 generations Journal Article
Nature, vol. 551, no. 7678, pp. 45–50, 2017, ISSN: 14764687.
Abstract | Links | BibTeX | Altmetric | Tags: Demography and Ecology, Genome Evolution, Historical Contingency, Mutation Rates, Parallelism and Divergence
@article{Good2017,
title = {The dynamics of molecular evolution over 60,000 generations},
author = {Benjamin H. Good and Michael J. McDonald and Jeffrey E. Barrick and Richard E. Lenski and Michael M. Desai},
url = {http://dx.doi.org/10.1038/nature24287},
doi = {10.1038/nature24287},
issn = {14764687},
year = {2017},
date = {2017-01-01},
urldate = {2017-01-01},
journal = {Nature},
volume = {551},
number = {7678},
pages = {45--50},
publisher = {Nature Publishing Group},
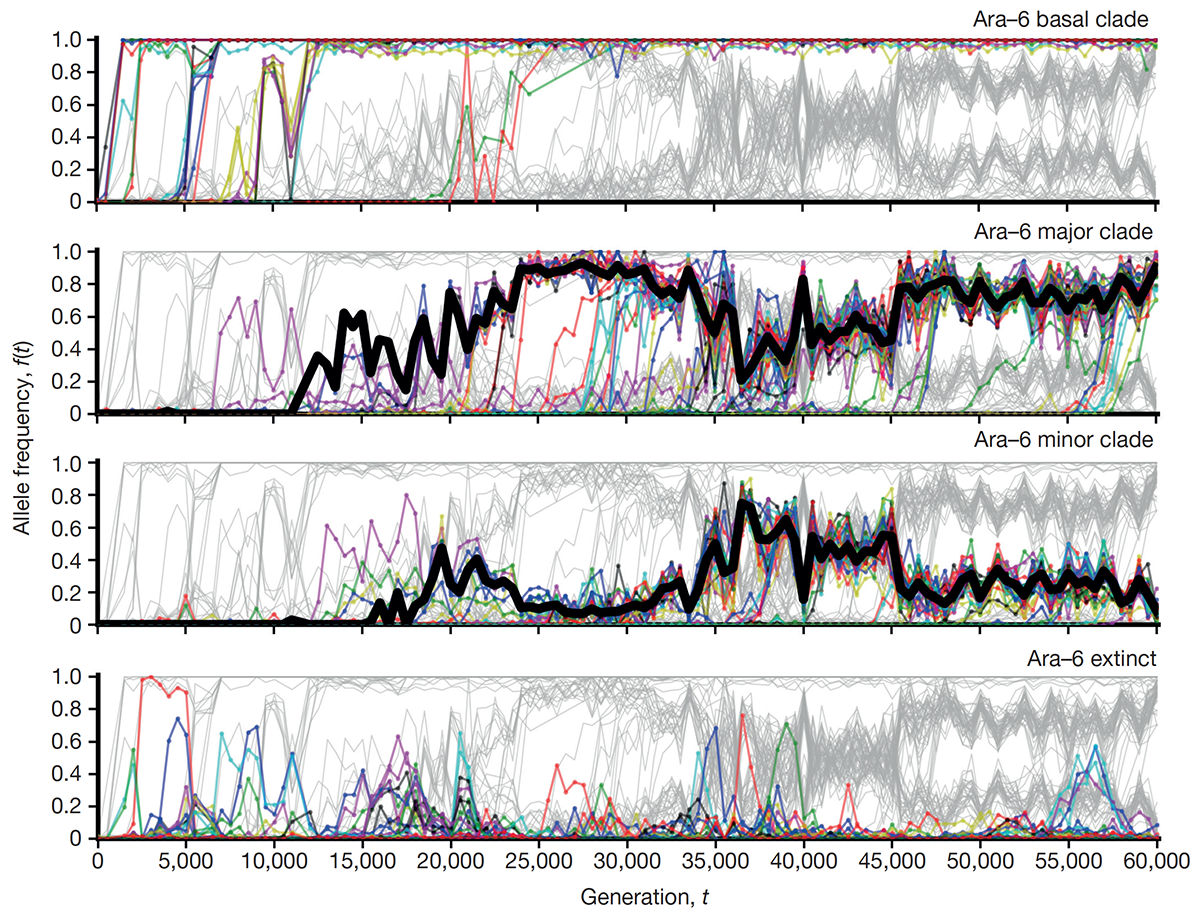
abstract = {The outcomes of evolution are determined by a stochastic dynamical process that governs how mutations arise and spread through a population. However, it is difficult to observe these dynamics directly over long periods and across entire genomes. Here we analyse the dynamics of molecular evolution in twelve experimental populations of \textit{Escherichia coli}, using whole-genome metagenomic sequencing at five hundred-generation intervals through sixty thousand generations. Although the rate of fitness gain declines over time, molecular evolution is characterized by signatures of rapid adaptation throughout the duration of the experiment, with multiple beneficial variants simultaneously competing for dominance in each population. Interactions between ecological and evolutionary processes play an important role, as long-term quasi-stable coexistence arises spontaneously in most populations, and evolution continues within each clade. We also present evidence that the targets of natural selection change over time, as epistasis and historical contingency alter the strength of selection on different genes. Together, these results show that long-term adaptation to a constant environment can be a more complex and dynamic process than is often assumed.},
keywords = {Demography and Ecology, Genome Evolution, Historical Contingency, Mutation Rates, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
2016
Tenaillon O; Barrick J E; Ribeck N; Deatherage D E; Blanchard J L; Dasgupta A; Wu G C; Wielgoss S; Cruveiller S; Medigue C; Schneider D; Lenski R E
Tempo and mode of genome evolution in a 50,000-generation experiment. Journal Article
Nature, vol. 536, no. 7615, pp. 165–170, 2016, ISSN: 1476-4687.
Abstract | Links | BibTeX | Altmetric | Tags: Genome Evolution, Mutation Rates, Parallelism and Divergence
@article{Tenaillon2016,
title = {Tempo and mode of genome evolution in a 50,000-generation experiment.},
author = {Olivier Tenaillon and Jeffrey E. Barrick and Noah Ribeck and Daniel E. Deatherage and Jeffrey L. Blanchard and Aurko Dasgupta and Gabriel C. Wu and Sebastien Wielgoss and Stephane Cruveiller and Claudine Medigue and Dominique Schneider and Richard E. Lenski},
url = {http://www.ncbi.nlm.nih.gov/pubmed/27479321},
doi = {10.1038/nature18959},
issn = {1476-4687},
year = {2016},
date = {2016-08-01},
urldate = {2016-08-01},
journal = {Nature},
volume = {536},
number = {7615},
pages = {165--170},
publisher = {Nature Publishing Group},
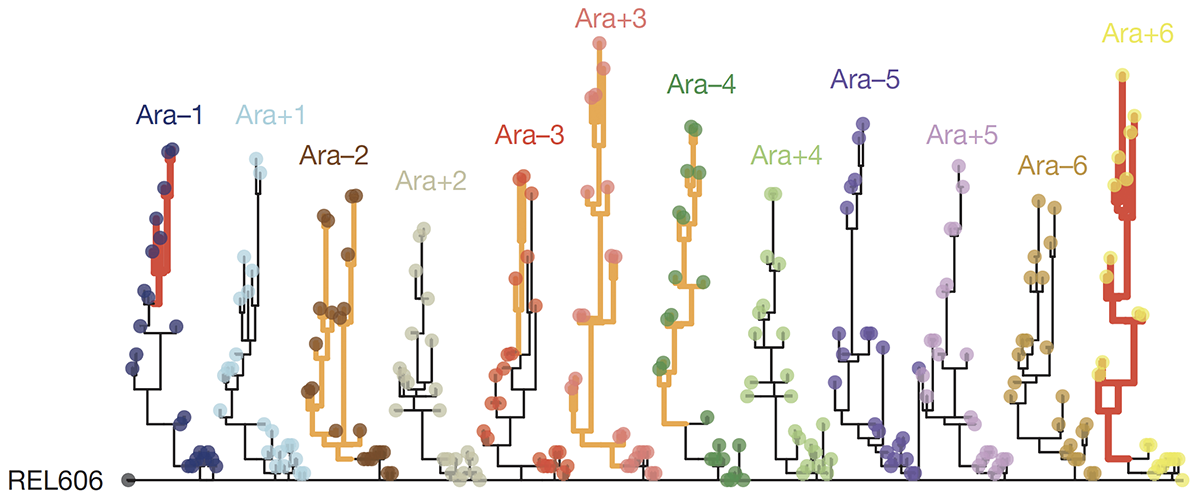
abstract = {Adaptation by natural selection depends on the rates, effects and interactions of many mutations, making it difficult to determine what proportion of mutations in an evolving lineage are beneficial. Here we analysed 264 complete genomes from 12 \textit{Escherichia coli} populations to characterize their dynamics over 50,000 generations. The populations that retained the ancestral mutation rate support a model in which most fixed mutations are beneficial, the fraction of beneficial mutations declines as fitness rises, and neutral mutations accumulate at a constant rate. We also compared these populations to mutation-accumulation lines evolved under a bottlenecking regime that minimizes selection. Nonsynonymous mutations, intergenic mutations, insertions and deletions are overrepresented in the long-term populations, further supporting the inference that most mutations that reached high frequency were favoured by selection. These results illuminate the shifting balance of forces that govern genome evolution in populations adapting to a new environment.},
keywords = {Genome Evolution, Mutation Rates, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
2015
Lenski R E; Wiser M J; Ribeck N; Blount Z D; Nahum J R; Morris J J; Zaman L; Turner C B; Wade B D; Maddamsetti R; Burmeister A R; Baird E J; Bundy J; Grant N A; Card K J; Rowles M; Weatherspoon K; Papoulis S E; Sullivan R; Clark C; Mulka J S; Hajela N
Sustained fitness gains and variability in fitness trajectories in the long-term evolution experiment with Escherichia coli Journal Article
Proceedings of the Royal Society B: Biological Sciences, vol. 282, no. 1821, pp. 20152292, 2015, ISSN: 0962-8452.
Abstract | Links | BibTeX | Altmetric | Tags: Fitness Trajectories, Mutation Rates, Parallelism and Divergence
@article{nokey,
title = {Sustained fitness gains and variability in fitness trajectories in the long-term evolution experiment with \textit{Escherichia coli}},
author = {Richard E. Lenski and Michael J. Wiser and Noah Ribeck and Zachary D. Blount and Joshua R. Nahum and J. Jeffrey Morris and Luis Zaman and Caroline B. Turner and Brian D. Wade and Rohan Maddamsetti and Alita R. Burmeister and Elizabeth J. Baird and Jay Bundy and Nkrumah A. Grant and Kyle J. Card and Maia Rowles and Kiyana Weatherspoon and Spiridon E. Papoulis and Rachel Sullivan and Colleen Clark and Joseph S. Mulka and Neerja Hajela},
url = {https://royalsocietypublishing.org/doi/10.1098/rspb.2015.2292},
doi = {10.1098/rspb.2015.2292},
issn = {0962-8452},
year = {2015},
date = {2015-12-22},
urldate = {2015-12-22},
journal = {Proceedings of the Royal Society B: Biological Sciences},
volume = {282},
number = {1821},
pages = {20152292},
abstract = {Many populations live in environments subject to frequent biotic and abiotic changes. Nonetheless, it is interesting to ask whether an evolving population's mean fitness can increase indefinitely, and potentially without any limit, even in a constant environment. A recent study showed that fitness trajectories of \textit{Escherichia coli} populations over 50 000 generations were better described by a power-law model than by a hyperbolic model. According to the power-law model, the rate of fitness gain declines over time but fitness has no upper limit, whereas the hyperbolic model implies a hard limit. Here, we examine whether the previously estimated power-law model predicts the fitness trajectory for an additional 10 000 generations. To that end, we conducted more than 1100 new competitive fitness assays. Consistent with the previous study, the power-law model fits the new data better than the hyperbolic model. We also analysed the variability in fitness among populations, finding subtle, but significant, heterogeneity in mean fitness. Some, but not all, of this variation reflects differences in mutation rate that evolved over time. Taken together, our results imply that both adaptation and divergence can continue indefinitely—or at least for a long time—even in a constant environment.},
keywords = {Fitness Trajectories, Mutation Rates, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
2010
Crozat E; Winkworth C L; Gaffé J; Hallin P F; Riley M A; Lenski R E; Schneider D
Parallel Genetic and Phenotypic Evolution of DNA Superhelicity in Experimental Populations of Escherichia coli Journal Article
Molecular Biology and Evolution, vol. 27, no. 9, pp. 2113–2128, 2010, ISSN: 0737-4038.
Abstract | Links | BibTeX | Altmetric | Tags: Genome Evolution, Parallelism and Divergence
@article{Crozat2010,
title = {Parallel Genetic and Phenotypic Evolution of DNA Superhelicity in Experimental Populations of \textit{Escherichia coli}},
author = {Estelle Crozat and Cynthia L. Winkworth and Joël Gaffé and Peter F. Hallin and Margaret A. Riley and Richard E. Lenski and Dominique Schneider},
url = {https://academic.oup.com/mbe/article-lookup/doi/10.1093/molbev/msq099},
doi = {10.1093/molbev/msq099},
issn = {0737-4038},
year = {2010},
date = {2010-09-01},
urldate = {2010-09-01},
journal = {Molecular Biology and Evolution},
volume = {27},
number = {9},
pages = {2113--2128},
abstract = {DNA supercoiling is the master function that interconnects chromosome structure and global gene transcription. This function has recently been shown to be under strong selection in \textit{Escherichia coli}. During the evolution of 12 initially identical populations propagated in a defined environment for 20,000 generations, parallel increases in DNA supercoiling were observed in ten populations. The genetic changes associated with the increased supercoiling were examined in one population, and beneficial mutations in the genes \textit{topA} (encoding topoisomerase I) and \textit{fis} (encoding a histone-like protein) were identified. To elucidate the molecular basis and impact of these changes, we quantified the level of genetic, phenotypic, and molecular parallelism linked to DNA supercoiling in all 12 evolving populations. First, sequence determination of DNA topology-related loci revealed strong genetic parallelism, with mutations concentrated in three genes (\textit{topA}, \textit{fis}, and \textit{dusB}), although the populations had different alleles at each locus. Statistical analyses of these polymorphisms implied the action of positive selection and, moreover, suggested that \textit{fis} and \textit{dusB}, which belong to the same operon, have related functions. Indeed, we demonstrated that DusB regulates the expression of \textit{fis} by both experimental and phylogenetic analyses. Second, molecular analyses of five mutations in \textit{fis} and \textit{dusB} affecting the transcription, translation, and protein activity of Fis also revealed strong parallelism in the resulting phenotypic effects. Third, artificially increasing DNA supercoiling in one of the two populations that lacked DNA topology changes led to a significant fitness increase. The high levels of molecular and genetic parallelism, targeting a small subset of the many genes involved in DNA supercoiling, indicate that changes in DNA superhelicity have been important in the evolution of these populations. Surprisingly, however, most of the evolved alleles we tested had either no detectable or slightly deleterious effects on fitness, despite these signatures of positive selection. },
keywords = {Genome Evolution, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
Meyer J R; Agrawal A A; Quick R T; Dobias D T; Schneider D; Lenski R E
Parallel changes in host resistance to viral infection during 45,000 generations of relaxed selection Journal Article
Evolution, vol. 64, no. 10, pp. no–no, 2010, ISSN: 00143820.
Abstract | Links | BibTeX | Altmetric | Tags: Correlated Responses, Genotypes and Phenotypes, Parallelism and Divergence
@article{Meyer2010,
title = {Parallel changes in host resistance to viral infection during 45,000 generations of relaxed selection},
author = {Justin R. Meyer and Anurag A. Agrawal and Ryan T. Quick and Devin T. Dobias and Dominique Schneider and Richard E. Lenski},
url = {https://onlinelibrary.wiley.com/doi/10.1111/j.1558-5646.2010.01049.x},
doi = {10.1111/j.1558-5646.2010.01049.x},
issn = {00143820},
year = {2010},
date = {2010-08-01},
urldate = {2010-08-01},
journal = {Evolution},
volume = {64},
number = {10},
pages = {no--no},
abstract = {The dynamics of host susceptibility to parasites are often influenced by trade-offs between the costs and benefits of resistance. We assayed changes in the resistance to three viruses in six lines of \textit{Escherichia coli} that had been evolving for almost 45,000 generations in their absence. The common ancestor of these lines was completely resistant to T6, partially resistant to T6* (a mutant of T6 with altered host range), and sensitive to lambda. None of the populations changed with respect to resistance to T6, whereas all six evolved increased susceptibility to T6*, probably ameliorating a cost of resistance. More surprisingly, however, the majority of lines evolved complete resistance to lambda, despite not encountering that virus during this period. By coupling our results with previous work, we infer that resistance to lambda evolved as a pleiotropic effect of a beneficial mutation that downregulated an unused metabolic pathway. The strong parallelism between the lines implies that selection had almost deterministic effects on the evolution of these patterns of host resistance. The opposite outcomes for resistance to T6* and lambda demonstrate that the evolution of host resistance under relaxed selection cannot be fully predicted by simple trade-off models.},
keywords = {Correlated Responses, Genotypes and Phenotypes, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
2009
Barrick J E; Yu D S; Yoon S H; Jeong H; Oh T K; Schneider D; Lenski R E; Kim J F
Genome evolution and adaptation in a long-term experiment with Escherichia coli. Journal Article
Nature, vol. 461, no. 7268, pp. 1243–7, 2009, ISSN: 1476-4687.
Abstract | Links | BibTeX | Altmetric | Tags: Fitness Trajectories, Genome Evolution, Genotypes and Phenotypes, Mutation Rates, Parallelism and Divergence
@article{Barrick2009,
title = {Genome evolution and adaptation in a long-term experiment with \textit{Escherichia coli}.},
author = {Jeffrey E. Barrick and Dong Su Yu and Sung Ho Yoon and Haeyoung Jeong and Tae Kwang Oh and Dominique Schneider and Richard E. Lenski and Jihyun F. Kim},
url = {http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=19838166&retmode=ref&cmd=prlinks papers2://publication/doi/10.1038/nature08480 http://www.ncbi.nlm.nih.gov/pubmed/19838166},
doi = {10.1038/nature08480},
issn = {1476-4687},
year = {2009},
date = {2009-10-01},
urldate = {2009-10-01},
journal = {Nature},
volume = {461},
number = {7268},
pages = {1243--7},
publisher = {Nature Publishing Group},
address = {Department of Microbiology and Molecular Genetics, Michigan State University, East Lansing, Michigan 48824, USA.},
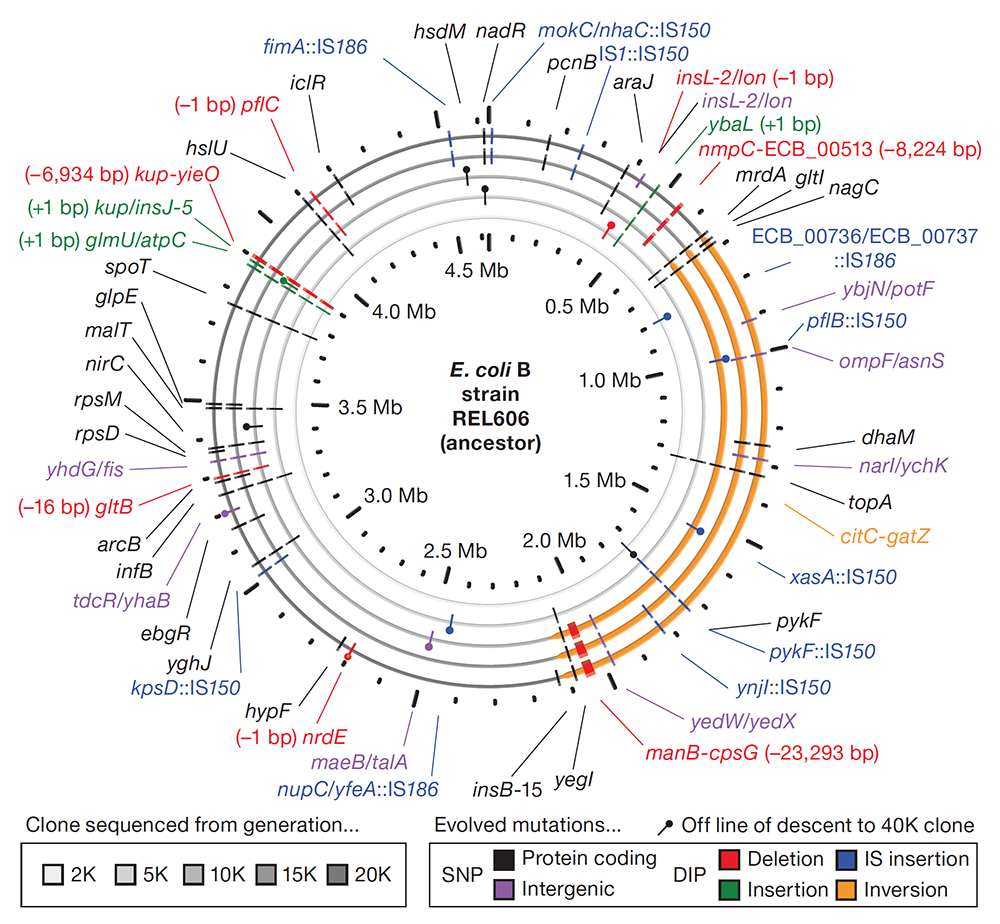
abstract = {The relationship between rates of genomic evolution and organismal adaptation remains uncertain, despite considerable interest. The feasibility of obtaining genome sequences from experimentally evolving populations offers the opportunity to investigate this relationship with new precision. Here we sequence genomes sampled through 40,000 generations from a laboratory population of \textit{Escherichia coli}. Although adaptation decelerated sharply, genomic evolution was nearly constant for 20,000 generations. Such clock-like regularity is usually viewed as the signature of neutral evolution, but several lines of evidence indicate that almost all of these mutations were beneficial. This same population later evolved an elevated mutation rate and accumulated hundreds of additional mutations dominated by a neutral signature. Thus, the coupling between genomic and adaptive evolution is complex and can be counterintuitive even in a constant environment. In particular, beneficial substitutions were surprisingly uniform over time, whereas neutral substitutions were highly variable.},
keywords = {Fitness Trajectories, Genome Evolution, Genotypes and Phenotypes, Mutation Rates, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
2008
Cooper T F; Remold S K; Lenski R E; Schneider D
Expression Profiles Reveal Parallel Evolution of Epistatic Interactions Involving the CRP Regulon in Escherichia coli Journal Article
PLoS Genetics, vol. 4, no. 2, pp. e35, 2008, ISSN: 1553-7404.
Abstract | Links | BibTeX | Altmetric | Tags: Genotypes and Phenotypes, Parallelism and Divergence
@article{Cooper2008,
title = {Expression Profiles Reveal Parallel Evolution of Epistatic Interactions Involving the CRP Regulon in \textit{Escherichia coli}},
author = {Tim F. Cooper and Susanna K. Remold and Richard E. Lenski and Dominique Schneider},
editor = {David S Guttman},
url = {https://dx.plos.org/10.1371/journal.pgen.0040035},
doi = {10.1371/journal.pgen.0040035},
issn = {1553-7404},
year = {2008},
date = {2008-02-01},
urldate = {2008-02-01},
journal = {PLoS Genetics},
volume = {4},
number = {2},
pages = {e35},
abstract = {The extent and nature of epistatic interactions between mutations are issues of fundamental importance in evolutionary biology. However, they are difficult to study and their influence on adaptation remains poorly understood. Here, we use a systems-level approach to examine epistatic interactions that arose during the evolution of \textit{Escherichia coli} in a defined environment. We used expression arrays to compare the effect on global patterns of gene expression of deleting a central regulatory gene, \textit{crp}. Effects were measured in two lineages that had independently evolved for 20,000 generations and in their common ancestor. We found that deleting \textit{crp} had a much more dramatic effect on the expression profile of the two evolved lines than on the ancestor. Because the sequence of the \textit{crp} gene was unchanged during evolution, these differences indicate epistatic interactions between \textit{crp} and mutations at other loci that accumulated during evolution. Moreover, a striking degree of parallelism was observed between the two independently evolved lines; 115 genes that were not \textit{crp}-dependent in the ancestor became dependent on \textit{crp} in both evolved lines. An analysis of changes in \textit{crp} dependence of well-characterized regulons identified a number of regulatory genes as candidates for harboring beneficial mutations that could account for these parallel expression changes. Mutations within three of these genes have previously been found and shown to contribute to fitness. Overall, these findings indicate that epistasis has been important in the adaptive evolution of these lines, and they provide new insight into the types of genetic changes through which epistasis can evolve. More generally, we demonstrate that expression profiles can be profitably used to investigate epistatic interactions. },
keywords = {Genotypes and Phenotypes, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
2006
Pelosi L; Kühn L; Guetta D; Garin J; Geiselmann J; Lenski R E; Schneider D
Parallel Changes in Global Protein Profiles During Long-Term Experimental Evolution in Escherichia coli Journal Article
Genetics, vol. 173, no. 4, pp. 1851-1869, 2006, ISSN: 1943-2631.
Abstract | Links | BibTeX | Altmetric | Tags: Genotypes and Phenotypes, Parallelism and Divergence
@article{nokey,
title = {Parallel Changes in Global Protein Profiles During Long-Term Experimental Evolution in \textit{Escherichia coli}},
author = {Ludovic Pelosi and Lauriane Kühn and Dorian Guetta and Jérôme Garin and Johannes Geiselmann and Richard E. Lenski and Dominique Schneider},
url = {https://academic.oup.com/genetics/article/173/4/1851/6061068},
doi = {10.1534/genetics.105.049619},
issn = {1943-2631},
year = {2006},
date = {2006-08-01},
urldate = {2006-08-01},
journal = {Genetics},
volume = {173},
number = {4},
pages = {1851-1869},
abstract = {Twelve populations of \textit{Escherichia coli} evolved in and adapted to a glucose-limited environment from a common ancestor. We used two-dimensional protein electrophoresis to compare two evolved clones, isolated from independently derived populations after 20,000 generations. Exceptional parallelism was detected. We compared the observed changes in protein expression profiles with previously characterized global transcription profiles of the same clones; this is the first time such a comparison has been made in an evolutionary context where these changes are often quite subtle. The two methodologies exhibited some remarkable similarities that highlighted two different levels of parallel regulatory changes that were beneficial during the evolution experiment. First, at the higher level, both methods revealed extensive parallel changes in the same global regulatory network, reflecting the involvement of beneficial mutations in genes that control the ppGpp regulon. Second, both methods detected expression changes of identical gene sets that reflected parallel changes at a lower level of gene regulation. The protein profiles led to the discovery of beneficial mutations affecting the \textit{malT} gene, with strong genetic parallelism across independently evolved populations. Functional and evolutionary analyses of these mutations revealed parallel phenotypic decreases in the maltose regulon expression and a high level of polymorphism at this locus in the evolved populations.},
keywords = {Genotypes and Phenotypes, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
Woods R J; Schneider D; Winkworth C L; Riley M A; Lenski R E
Tests of parallel molecular evolution in a long-term experiment with Escherichia coli Journal Article
Proceedings of the National Academy of Sciences of the United States of America, vol. 103, no. 24, pp. 9107–9112, 2006, ISSN: 0027-8424.
Abstract | Links | BibTeX | Altmetric | Tags: Genome Evolution, Genotypes and Phenotypes, Parallelism and Divergence
@article{Woods2006,
title = {Tests of parallel molecular evolution in a long-term experiment with \textit{Escherichia coli}},
author = {Robert J. Woods and Dominique Schneider and Cynthia L. Winkworth and Margaret A. Riley and Richard E. Lenski},
url = {http://www.pnas.org/cgi/doi/10.1073/pnas.0602917103},
doi = {10.1073/pnas.0602917103},
issn = {0027-8424},
year = {2006},
date = {2006-06-01},
urldate = {2006-06-01},
journal = {Proceedings of the National Academy of Sciences of the United States of America},
volume = {103},
number = {24},
pages = {9107--9112},
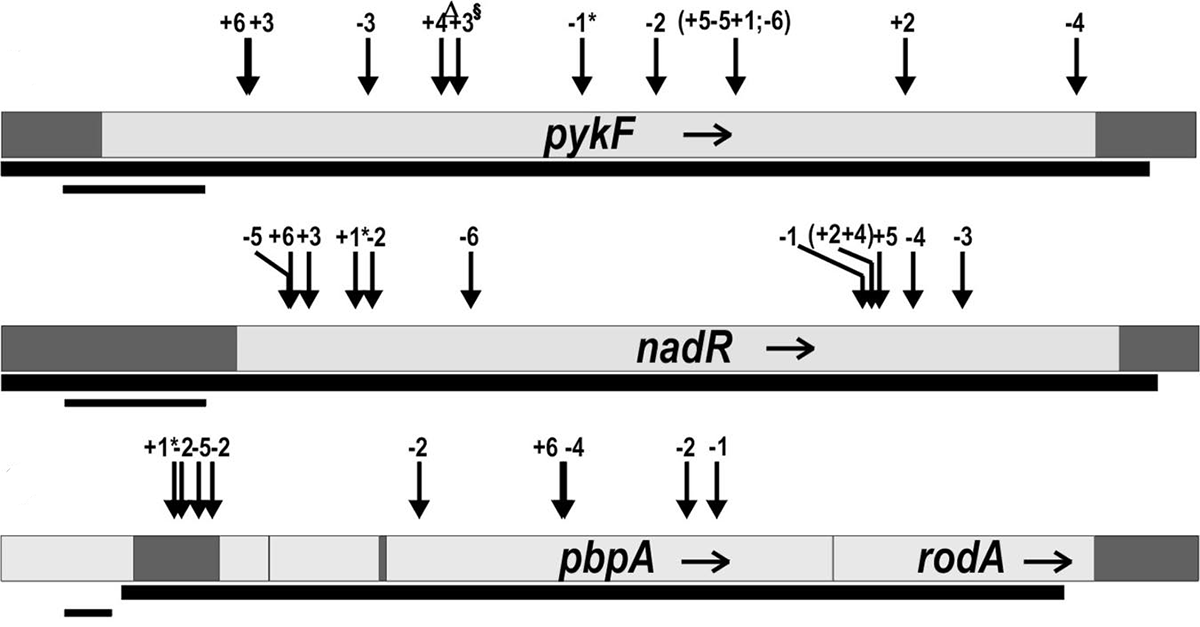
abstract = {The repeatability of evolutionary change is difficult to quantify because only a single outcome can usually be observed for any precise set of circumstances. In this study, however, we have quantified the frequency of parallel and divergent genetic changes in 12 initially identical populations of \textit{Escherichia coli} that evolved in identical environments for 20,000 cell generations. Unlike previous analyses in which candidate genes were identified based on parallel phenotypic changes, here we sequenced four loci (\textit{pykF}, \textit{nadR}, \textit{pbpA}-\textit{rodA}, and \textit{hokB}/\textit{sokB}) in which mutations of unknown effect had been discovered in one population, and then we compared the substitution pattern in these "blind" candidate genes with the pattern found in 36 randomly chosen genes. Two candidate genes, \textit{pykF} and \textit{nadR}, had substitutions in all 11 other populations, and the other 2 in several populations. There were very few cases, however, in which the exact same mutations were substituted, in contrast to the findings from conceptually related work performed with evolving virus populations. No random genes had any substitutions except in four populations that evolved defects in DNA repair. Tests of four different statistical aspects of the pattern of molecular evolution all indicate that adaptation by natural selection drove the parallel changes in these candidate genes.},
keywords = {Genome Evolution, Genotypes and Phenotypes, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
2005
Crozat E; Philippe N; Lenski R E; Geiselmann J; Schneider D
Long-Term Experimental Evolution in Escherichia coli. XII. DNA Topology as a Key Target of Selection Journal Article
Genetics, vol. 169, no. 2, pp. 523-532, 2005, ISSN: 1943-2631.
Abstract | Links | BibTeX | Altmetric | Tags: Genotypes and Phenotypes, Parallelism and Divergence
@article{nokey,
title = {Long-Term Experimental Evolution in \textit{Escherichia coli}. XII. DNA Topology as a Key Target of Selection},
author = {Estelle Crozat and Nadège Philippe and Richard E. Lenski and Johannes Geiselmann and Dominique Schneider},
url = {https://academic.oup.com/genetics/article/169/2/523/6060318},
doi = {10.1534/genetics.104.035717},
issn = {1943-2631},
year = {2005},
date = {2005-02-01},
urldate = {2005-02-01},
journal = {Genetics},
volume = {169},
number = {2},
pages = {523-532},
abstract = {The genetic bases of adaptation are being investigated in 12 populations of \textit{Escherichia coli}, founded from a common ancestor and serially propagated for 20,000 generations, during which time they achieved substantial fitness gains. Each day, populations alternated between active growth and nutrient exhaustion. DNA supercoiling in bacteria is influenced by nutritional state, and DNA topology helps coordinate the overall pattern of gene expression in response to environmental changes. We therefore examined whether the genetic controls over supercoiling might have changed during the evolution experiment. Parallel changes in topology occurred in most populations, with the level of DNA supercoiling increasing, usually in the first 2000 generations. Two mutations in the \textit{topA} and \textit{fis} genes that control supercoiling were discovered in a population that served as the focus for further investigation. Moving the mutations, alone and in combination, into the ancestral background had an additive effect on supercoiling, and together they reproduced the net change in DNA topology observed in this population. Moreover, both mutations were beneficial in competition experiments. Clonal interference involving other beneficial DNA topology mutations was also detected. These findings define a new class of fitness-enhancing mutations and indicate that the control of DNA supercoiling can be a key target of selection in evolving bacterial populations.},
keywords = {Genotypes and Phenotypes, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
2003
Cooper T F; Rozen D E; Lenski R E
Parallel changes in gene expression after 20,000 generations of evolution in Escherichia coli Journal Article
Proceedings of the National Academy of Sciences of the United States of America, vol. 100, no. 3, pp. 1072–1077, 2003, ISSN: 0027-8424.
Abstract | Links | BibTeX | Altmetric | Tags: Genotypes and Phenotypes, Parallelism and Divergence
@article{Cooper2003,
title = {Parallel changes in gene expression after 20,000 generations of evolution in \textit{Escherichia coli}},
author = {Tim F. Cooper and Daniel E. Rozen and Richard E. Lenski},
url = {http://www.pnas.org/cgi/doi/10.1073/pnas.0334340100},
doi = {10.1073/pnas.0334340100},
issn = {0027-8424},
year = {2003},
date = {2003-02-01},
urldate = {2003-02-01},
journal = {Proceedings of the National Academy of Sciences of the United States of America},
volume = {100},
number = {3},
pages = {1072--1077},
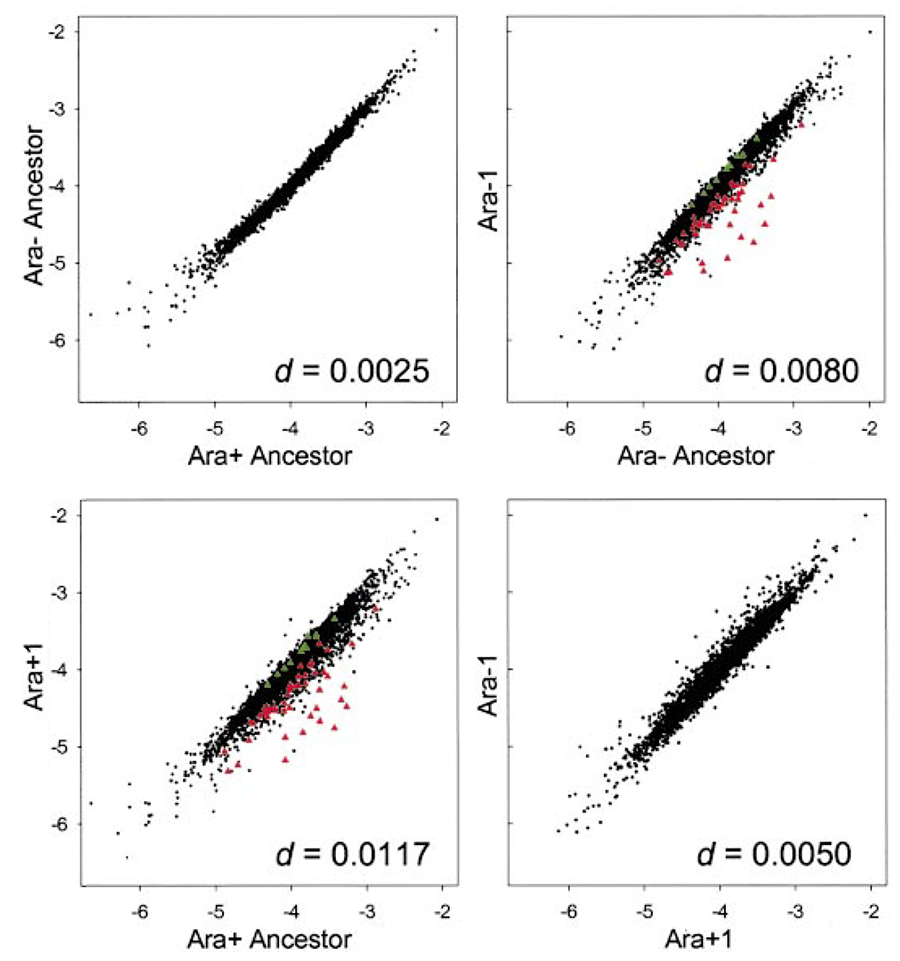
abstract = {Twelve populations of \textit{Escherichia coli}, derived from a common ancestor, evolved in a glucose-limited medium for 20,000 generations. Here we use DNA expression arrays to examine whether gene-expression profiles in two populations evolved in parallel, which would indicate adaptation, and to gain insight into the mechanisms underlying their adaptation. We compared the expression profile of the ancestor to that of clones sampled from both populations after 20,000 generations. The expression of 59 genes had changed significantly in both populations. Remarkably, all 59 were changed in the same direction relative to the ancestor. Many of these genes were members of the cAMP-cAMP receptor protein (CRP) and guanosine tetraphosphate (ppGpp) regulons. Sequencing of several genes controlling the effectors of these regulons found a nonsynonymous mutation in \textit{spoT} in one population. Moving this mutation into the ancestral background showed that it increased fitness and produced many of the expression changes manifest after 20,000 generations. The same mutation had no effect on fitness when introduced into the other evolved population, indicating that a mutation of similar effect was present already. Our study demonstrates the utility of expression arrays for addressing evolutionary issues including the quantitative measurement of parallel evolution in independent lineages and the identification of beneficial mutations.},
keywords = {Genotypes and Phenotypes, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
1996
Travisano M; Lenski R E
Long-Term Experimental Evolution in Escherichia coli. IV. Targets of Selection and the Specificity of Adaptation Journal Article
Genetics, vol. 143, no. 1, pp. 15–26, 1996, ISSN: 1943-2631.
Abstract | Links | BibTeX | Altmetric | Tags: Correlated Responses, Genotypes and Phenotypes, Parallelism and Divergence
@article{Travisano1996,
title = {Long-Term Experimental Evolution in \textit{Escherichia coli}. IV. Targets of Selection and the Specificity of Adaptation},
author = {Michael Travisano and Richard E. Lenski},
url = {https://academic.oup.com/genetics/article/143/1/15/6016887},
doi = {10.1093/genetics/143.1.15},
issn = {1943-2631},
year = {1996},
date = {1996-05-01},
urldate = {1996-05-01},
journal = {Genetics},
volume = {143},
number = {1},
pages = {15--26},
abstract = {This study investigates the physiological manifestation of adaptive evolutionary change in 12 replicate populations of \textit{Escherichia coli} that were propagated for 2000 generations in a glucose-limited environment. Representative genotypes from each population were assayed for fitness relative to their common ancestor in the experimental glucose environment and in 11 novel single-nutrient environments. After 2000 generations, the 12 derived genotypes had diverged into at least six distinct phenotypic classes. The nutrients were classified into four groups based upon their uptake physiology. All 12 derived genotypes improved in fitness by similar amounts in the glucose environment, and this pattern of parallel fitness gains was also seen in those novel environments where the limiting nutrient shared uptake mechanisms with glucose. Fitness showed little or no consistent improvement, but much greater genetic variation, in novel environments where the limiting nutrient differed from glucose in its uptake mechanisms. This pattern of fitness variation in the novel nutrient environments suggests that the independently derived genotypes adapted to the glucose environment by similar, but not identical, changes in the physiological mechanisms for moving glucose across both the inner and outer membranes.},
keywords = {Correlated Responses, Genotypes and Phenotypes, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
1995
Travisano M; Mongold J A; Bennett A F; Lenski R E
Experimental Tests of the Roles of Adaptation, Chance, and History Journal Article
Science, vol. 267, no. January, 1995.
Abstract | Links | BibTeX | Altmetric | Tags: Cell Morphology, Correlated Responses, Descendant Experiments, Fitness Trajectories, Historical Contingency, Methods and Miscellaneous, Parallelism and Divergence
@article{Travisano1995,
title = {Experimental Tests of the Roles of Adaptation, Chance, and History},
author = {Michael Travisano and Judith A. Mongold and Albert F. Bennett and Richard E. Lenski},
url = {https://www.science.org/lookup/doi/10.1126/science.7809610},
doi = {https://doi.org/10.1126/science.7809610},
year = {1995},
date = {1995-01-01},
urldate = {1995-01-01},
journal = {Science},
volume = {267},
number = {January},
abstract = {The contributions of adaptation, chance, and history to the evolution of fitness and cell size were measured in two separate experiments using bacteria. In both experiments, populations propagated in identical environments achieved similar fitnesses, regardless of prior history or subsequent chance events. In contrast, the evolution of cell size, a trait weakly correlated with fitness, was more strongly influenced by history and chance.},
keywords = {Cell Morphology, Correlated Responses, Descendant Experiments, Fitness Trajectories, Historical Contingency, Methods and Miscellaneous, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
Travisano M; Vasi F K; Lenski R E
Long-Term Experimental Evolution in Escherichia coli. III. Variation Among Replicate Populations in Correlated Responses to Novel Environments Journal Article
Evolution, vol. 49, no. 1, pp. 189–200, 1995.
Abstract | Links | BibTeX | Altmetric | Tags: Correlated Responses, Genotypes and Phenotypes, Parallelism and Divergence
@article{Travisano1995b,
title = {Long-Term Experimental Evolution in \textit{Escherichia coli}. III. Variation Among Replicate Populations in Correlated Responses to Novel Environments},
author = {Michael Travisano and Farida K. Vasi and Richard E. Lenski},
url = {https://onlinelibrary.wiley.com/doi/10.1111/j.1558-5646.1995.tb05970.x},
doi = {https://doi.org/10.1111/j.1558-5646.1995.tb05970.x},
year = {1995},
date = {1995-01-01},
urldate = {1995-01-01},
journal = {Evolution},
volume = {49},
number = {1},
pages = {189--200},
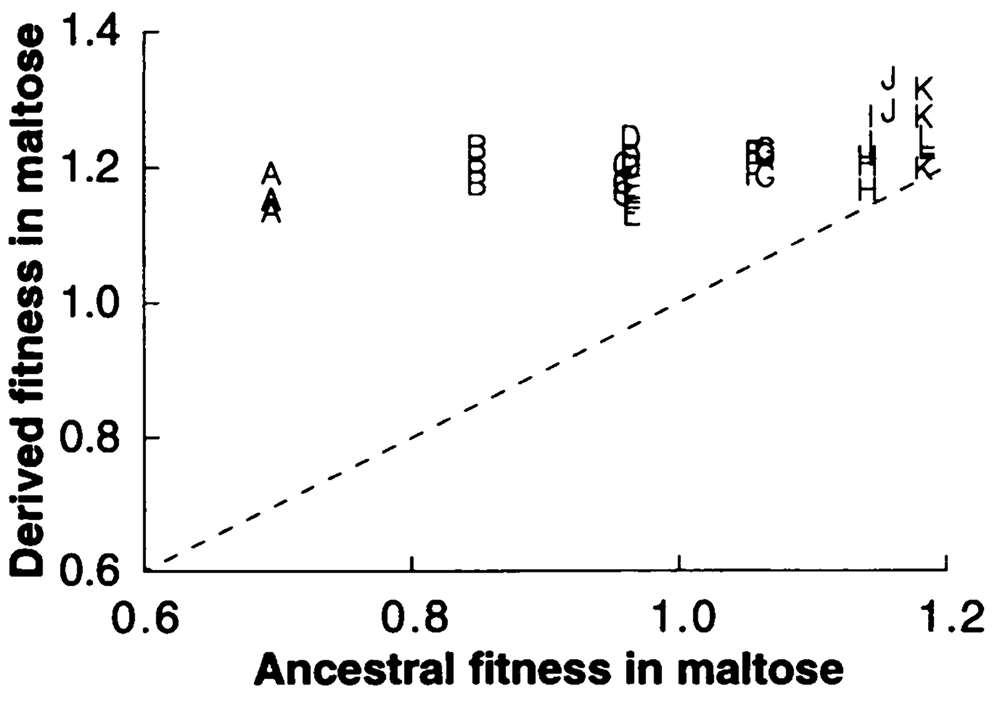
abstract = {Twelve populations of \textit{Escherichia coli} were founded from a single clone and propagated for 2000 gen- erations in identical glucose-limited environments. During this time, the mean fitnesses of the evolving populations relative to their common ancestor improved greatly, but their fitnesses relative to one another diverged only slightly. Although the populations showed similar fitness increases, they may have done so by different underlying adaptations, or they may have diverged in other respects by random genetic drift. Therefore, we examined the relative fitnesses of independently derived genotypes in two other sugars, maltose and lactose, to determine whether they were ho- mogeneous or heterogeneous in these environments. The genetic variation among the derived lines in fitness on maltose and lactose was more than 100-times greater than their variation in fitness on glucose. Moreover, the glucose-adapted genotypes, on average, showed significant adaptation to lactose, but not to maltose. That pathways for use of maltose and glucose are virtually identical in \textit{E. coli}, except for their distinct mechanisms of uptake, suggests that the derived genotypes have adapted primarily by improved glucose transport. From consideration of the number of generations of divergence, the mutation rate in \textit{E. coli}, and the proportion of its genome required for growth on maltose (but not glucose), we hypothesize that pleiotropy involving the selected alleles, rather than random genetic drift of alleles at other loci, was the major cause of the variation among the derived genotypes in fitness on these other sugars.},
keywords = {Correlated Responses, Genotypes and Phenotypes, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
Johnson P A; Lenski R E; Hoppensteadt F C
Theoretical Analysis of Divergence in Mean Fitness between Initially Identical Populations Journal Article
Proceedings of the Royal Society, London, pp. 125–130, 1995.
Abstract | Links | BibTeX | Altmetric | Tags: Fitness Trajectories, Parallelism and Divergence, Theory and Simulations
@article{Johnson1995,
title = {Theoretical Analysis of Divergence in Mean Fitness between Initially Identical Populations},
author = {Paul A. Johnson and Richard E. Lenski and Frank C. Hoppensteadt},
url = {https://royalsocietypublishing.org/doi/10.1098/rspb.1995.0019},
doi = {10.1098/rspb.1995.0019},
year = {1995},
date = {1995-01-01},
urldate = {1995-01-01},
journal = {Proceedings of the Royal Society, London},
pages = {125--130},
abstract = {Initially identical populations in identical environments may subsequently diverge from one another not only via the effects of genetic drift on neutral alleles, but also by selection on beneficial alleles that arise stochastically by mutation. In the simple case of one locus with two alleles in a haploid organism, a full range of combinations of population sizes, selection pressures, mutation rates and fixation probabilities reveals two qualitatively distinct dynamics of divergence among such initially identical populations. We define a non-dimensional parameter \textit{k} that describes conditions for the occurrence of these different dynamics. One dynamic (\textit{k} > 1) occurs when beneficial mutations are sufficiently common that substitutions within the populations are essentially simultaneous; the other dynamic (\textit{k} < 1) occurs when beneficial mutations are so rare that substitutions are likely to occur as isolated events. If there are more than two alleles, or multiple loci, divergence among the populations can be sustained indefinitely if \textit{k} < 1. The parameter \textit{k} pertains to the nature of biological evolution and its tendency to be gradual or punctuated.},
keywords = {Fitness Trajectories, Parallelism and Divergence, Theory and Simulations},
pubstate = {published},
tppubtype = {article}
}
1994
Vasi F K; Travisano M; Lenski R E
Long-Term Experimental Evolution in Escherichia coli. II. Changes in Life-History Traits During Adaptation to a Seasonal Environment Journal Article
The American Naturalist, vol. 144, no. 3, pp. 432–456, 1994, ISSN: 0003-0147.
Abstract | Links | BibTeX | Altmetric | Tags: Demography and Ecology, Parallelism and Divergence
@article{Vasi1994,
title = {Long-Term Experimental Evolution in \textit{Escherichia coli}. II. Changes in Life-History Traits During Adaptation to a Seasonal Environment},
author = {Farida K. Vasi and Michael Travisano and Richard E. Lenski},
url = {https://www.journals.uchicago.edu/doi/10.1086/285685},
doi = {10.1086/285685},
issn = {0003-0147},
year = {1994},
date = {1994-09-01},
urldate = {1994-09-01},
journal = {The American Naturalist},
volume = {144},
number = {3},
pages = {432--456},
abstract = {Twelve populations of the bacterium \textit{Escherichia coli} were propagated for 2,000 generations in a seasonal environment, which consisted of alternating periods of feast and famine. The mean fitness of the derived genotypes increased by ∼35% relative to their common ancestor, based on competition experiments in the same environment. The bacteria could have adapted, in principle, by decreasing their lag prior to growth upon transfer to fresh medium (L), increasing their maximum growth rate (V_{m}), reducing the concentration of resource required to support growth at half the maximum rate (K_{s}), and reducing their death rate after the limiting resource was exhausted (D). We estimated these parameters for the ancestor and then calculated the opportunity for selection on each parameter. The inferred selection gradients for V_{m} and L were much steeper than for K_{s} and D. The derived genotypes showed significant improvement in V_{m} and L but not in K_{s} or D. Also, the numerical yield in pure culture of the derived genotypes was significantly lower than the yield of the common ancestor, but the average cell size was much larger. The independently derived genotypes are somewhat more variable in these life-history traits than in their relative fitnesses, which indicates that they acquired different genetic adaptations to the seasonal environment. Nonetheless, the evolutionary changes in life-history traits exhibit substantial parallelism among the replicate populations.},
keywords = {Demography and Ecology, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}