2026
Donofrio M A; Blasius H L; Nguyen C C; Schnell A L; Turner C B
Antibiotic susceptibility of Escherichia coli is affected by evolutionary history but not by history of elemental limitation Journal Article Forthcoming
mSphere, pp. e00538-25, Forthcoming.
Abstract | Links | BibTeX | Altmetric | Tags: Correlated Responses, Descendant Experiments, Genotypes and Phenotypes
@article{Donofrio2026,
title = {Antibiotic susceptibility of \textit{Escherichia coli} is affected by evolutionary history but not by history of elemental limitation},
author = {Marissa A. Donofrio and Heather L. Blasius and Catherine C. Nguyen and Alexa L. Schnell and Caroline B. Turner},
url = {https://journals.asm.org/doi/10.1128/msphere.00538-25},
doi = {10.1128/msphere.00538-25},
year = {2026},
date = {2026-03-23},
journal = {mSphere},
pages = {e00538-25},
abstract = {ABSTRACT: Antibiotic resistance in bacteria is a global public health threat. To understand how the evolution of antibiotic susceptibility is affected by environmental conditions and prior evolutionary history, we worked with populations from the Long-Term Evolution Experiment (LTEE) with \textit{Escherichia coli}. These populations previously evolved independently for 50,000 generations in an environment without antibiotics, making them an ideal system for studying the effect of evolutionary history on adaptation to new selective pressures. We further evolved five of the LTEE populations, as well as their shared ancestor, under either carbon- or nitrogen-limited conditions and then tested intrinsic resistance to four antibiotics. Evolution under elemental limitation did not have a significant impact on resistance to any of the tested antibiotics. However, some LTEE populations did have higher resistance than other populations. Susceptibility also varied within one population, which had the lowest level of resistance to all four antibiotics. We hypothesized that resistance levels might differ between two clades of bacteria that have coexisted within this population for more than 40,000 generations. Interestingly, although antibiotic susceptibility varied within the population, there was no consistent difference between clades. Instead, one particular clone isolated from the population exhibited higher resistance than the other clones sampled. These findings indicate that antibiotic resistance can vary both within and between experimentally evolved populations, even in the absence of direct selection on resistance. Our results also show that measured levels of susceptibility may depend on stochastic sampling effects during isolation of clones. IMPORTANCE: Antibiotic resistance is one of the most pressing health challenges worldwide, and understanding how bacteria evolve resistance, even when not directly exposed to antibiotics, is critical for managing and predicting emerging threats. Our study leverages the unique Long-Term Evolution Experiment with \textit{Escherichia coli} to show that both the evolutionary history of bacterial populations and random variation among individual clones can significantly influence intrinsic antibiotic susceptibility. Our results also suggest that elemental limitation, while a critical environmental variable, may not be an important driver of intrinsic antibiotic susceptibility, at least over short time frames.},
keywords = {Correlated Responses, Descendant Experiments, Genotypes and Phenotypes},
pubstate = {forthcoming},
tppubtype = {article}
}
2024
Mori M; Patsalo V; Euler C; Williamson J R; Scott M
Proteome partitioning constraints in long-term laboratory evolution Journal Article
Nature Communications, 15 , pp. 4087, 2024, ISSN: 2041-1723.
Abstract | Links | BibTeX | Altmetric | Tags: experimental evolution, Genotypes and Phenotypes, Microbial genetics, Proteomic analysis
@article{Mori2024,
title = {Proteome partitioning constraints in long-term laboratory evolution},
author = {Matteo Mori and Vadim Patsalo and Christian Euler and James R. Williamson and Matthew Scott},
url = {https://www.nature.com/articles/s41467-024-48447-2},
doi = {10.1038/s41467-024-48447-2},
issn = {2041-1723},
year = {2024},
date = {2024-05-01},
urldate = {2024-05-01},
journal = {Nature Communications},
volume = {15},
pages = {4087},
abstract = {Adaptive laboratory evolution experiments provide a controlled context in which the dynamics of selection and adaptation can be followed in real-time at the single-nucleotide level. And yet this precision introduces hundreds of degrees-of-freedom as genetic changes accrue in parallel lineages over generations. On short timescales, physiological constraints have been leveraged to provide a coarse-grained view of bacterial gene expression characterized by a small set of phenomenological parameters. Here, we ask whether this same framework, operating at a level between genotype and fitness, informs physiological changes that occur on evolutionary timescales. Using a strain adapted to growth in glucose minimal medium, we find that the proteome is substantially remodeled over 40 000 generations. The most striking change is an apparent increase in enzyme efficiency, particularly in the enzymes of lower-glycolysis. We propose that deletion of metabolic flux-sensing regulation early in the adaptation results in increased enzyme saturation and can account for the observed proteome remodeling.},
keywords = {experimental evolution, Genotypes and Phenotypes, Microbial genetics, Proteomic analysis},
pubstate = {published},
tppubtype = {article}
}
Ascensao J A; Denk J; Lok K; Yu Q; Wetmore K M; Hallatschek O
Rediversification following ecotype isolation reveals hidden adaptive potential Journal Article
Current Biology, 34 (4), pp. 855–867, 2024, ISSN: 0960-9822.
Abstract | Links | BibTeX | Altmetric | Tags: Demography and Ecology, Genotypes and Phenotypes, Historical Contingency
@article{Ascensao2024,
title = {Rediversification following ecotype isolation reveals hidden adaptive potential},
author = {Joao A. Ascensao and Jonas Denk and Kristen Lok and QinQin Yu and Kelly M. Wetmore and Oskar Hallatschek},
url = {https://www.sciencedirect.com/science/article/pii/S0960982224000290},
doi = {10.1016/j.cub.2024.01.029},
issn = {0960-9822},
year = {2024},
date = {2024-02-01},
urldate = {2024-02-01},
journal = {Current Biology},
volume = {34},
number = {4},
pages = {855--867},
abstract = {Microbial communities play a critical role in ecological processes, and their diversity is key to their functioning. However, little is known about whether communities can regenerate ecological diversity following ecotype removal or extinction and how the rediversified communities would compare to the original ones. Here, we show that simple two-ecotype communities from the \textit{E. coli} long-term evolution experiment (LTEE) consistently rediversified into two ecotypes following the isolation of one of the ecotypes, coexisting via negative frequency-dependent selection. Communities separated by more than 30,000 generations of evolutionary time rediversify in similar ways. The rediversified ecotype appears to share a number of growth traits with the ecotype it replaces. However, the rediversified community is also different from the original community in ways relevant to the mechanism of ecotype coexistence—for example, in stationary phase response and survival. We found substantial variation in the transcriptional states between the two original ecotypes, whereas the differences within the rediversified community were comparatively smaller, although the rediversified community showed unique patterns of differential expression. Our results suggest that evolution may leave room for alternative diversification processes even in a maximally reduced community of only two strains. We hypothesize that the presence of alternative evolutionary pathways may be even more pronounced in communities of many species where there are even more potential niches, highlighting an important role for perturbations, such as species removal, in evolving ecological communities.},
keywords = {Demography and Ecology, Genotypes and Phenotypes, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
2023
Favate J S; Skalenko K S; Chiles E; Su X; Yadavalli S S; Shah P
Linking genotypic and phenotypic changes in the E. coli long-term evolution experiment using metabolomics Journal Article
eLife, 12 , pp. RP87039, 2023, ISSN: eLife.
Abstract | Links | BibTeX | Altmetric | Tags: Genotypes and Phenotypes, Parallelism and Divergence
@article{10.7554/eLife.87039,
title = {Linking genotypic and phenotypic changes in the \textit{E. coli} long-term evolution experiment using metabolomics},
author = {John S Favate and Kyle S Skalenko and Eric Chiles and Xiaoyang Su and Srujana Samhita Yadavalli and Premal Shah},
editor = {John McCutcheon and Christian R Landry},
url = {https://doi.org/10.7554/eLife.87039},
doi = {10.7554/eLife.87039},
issn = {eLife},
year = {2023},
date = {2023-11-01},
urldate = {2023-11-01},
journal = {eLife},
volume = {12},
pages = {RP87039},
publisher = {2050-084X},
abstract = {Changes in an organism's environment, genome, or gene expression patterns can lead to changes in its metabolism. The metabolic phenotype can be under selection and contributes to adaptation. However, the networked and convoluted nature of an organism's metabolism makes relating mutations, metabolic changes, and effects on fitness challenging. To overcome this challenge, we use the long-term evolution experiment (LTEE) with \textit{E. coli} as a model to understand how mutations can eventually affect metabolism and perhaps fitness. We used mass spectrometry to broadly survey the metabolomes of the ancestral strains and all 12 evolved lines. We combined this metabolic data with mutation and expression data to suggest how mutations that alter specific reaction pathways, such as the biosynthesis of nicotinamide adenine dinucleotide, might increase fitness in the system. Our work provides a better understanding of how mutations might affect fitness through the metabolic changes in the LTEE and thus provides a major step in developing a complete genotype-phenotype map for this experimental system. },
key = {E. coli; LTEE; adaptation; bacteria; evolutionary biology; metabolomics},
keywords = {Genotypes and Phenotypes, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
Ascensao J A; Wetmore K M; Good B H; Arkin A P; Hallatschek O
Quantifying the adaptive potential of a nascent bacterial community Journal Article
Nature Communications, 14 (1), pp. 248, 2023.
Abstract | Links | BibTeX | Altmetric | Tags: Demography and Ecology, Descendant Experiments, Genome Evolution, Genotypes and Phenotypes
@article{ascensao2023,
title = {Quantifying the adaptive potential of a nascent bacterial community},
author = {Joao A. Ascensao and Kelly M. Wetmore and Benjamin H. Good and Adam P. Arkin and Oskar Hallatschek},
url = {https://www.nature.com/articles/s41467-022-35677-5},
doi = {10.1038/s41467-022-35677-5},
year = {2023},
date = {2023-01-16},
urldate = {2022-01-01},
journal = {Nature Communications},
volume = {14},
number = {1},
pages = {248},
publisher = {Cold Spring Harbor Laboratory},
abstract = {The fitness effects of all possible mutations available to an organism largely shape the dynamics of evolutionary adaptation. Yet, whether and how this adaptive landscape changes over evolutionary times, especially upon ecological diversification and changes in community composition, remains poorly understood. We sought to fill this gap by analyzing a stable community of two closely related ecotypes (“L” and “S”) shortly after they emerged within the \textit{E. coli} Long-Term Evolution Experiment (LTEE). We engineered genome-wide barcoded transposon libraries to measure the invasion fitness effects of all possible gene knockouts in the coexisting strains as well as their ancestor, for many different, ecologically relevant conditions. We find consistent statistical patterns of fitness effect variation across both genetic background and community composition, despite the idiosyncratic behavior of individual knockouts. Additionally, fitness effects are correlated with evolutionary outcomes for a number of conditions, possibly revealing shifting patterns of adaptation. Together, our results reveal how ecological and epistatic effects combine to shape the adaptive landscape in a nascent ecological community.},
keywords = {Demography and Ecology, Descendant Experiments, Genome Evolution, Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}
2022
Laurin D; Mercier C; Quansah N; Robert J S; Usson Y; Schneider D; Hindré T; Schaack B
Extracellular vesicles from 50,000 generation clones of the Escherichia coli long-term evolution experiment Journal Article
International Journal of Molecular Sciences, 23 , pp. 14580, 2022, ISSN: 1422-0067.
Abstract | Links | BibTeX | Altmetric | Tags: Cell Morphology, Genotypes and Phenotypes
@article{Laurin2022,
title = {Extracellular vesicles from 50,000 generation clones of the \textit{Escherichia coli} long-term evolution experiment},
author = {David Laurin and Corinne Mercier and Nyamekye Quansah and Julie Suzanne Robert and Yves Usson and Dominique Schneider and Thomas Hindré and Béatrice Schaack },
url = {https://www.mdpi.com/1422-0067/23/23/14580},
doi = {10.3390/ijms232314580},
issn = {1422-0067},
year = {2022},
date = {2022-11-01},
urldate = {2022-11-01},
journal = {International Journal of Molecular Sciences},
volume = {23},
pages = {14580},
abstract = {Extracellular vesicles (EVs) are critical elements of cell–cell communication. Here, we characterized the outer membrane vesicles (OMVs) released by specific clones of Escherichia coli isolated from the Long-Term Evolution Experiment after 50,000 generations (50K) of adaptation to glucose minimal medium. Compared with their ancestor, the evolved clones produce small OMVs but also larger ones which display variable amounts of both OmpA and LPS. Tracking ancestral, fluorescently labelled OMVs revealed that they fuse with both ancestral- and 50K-evolved cells, albeit in different proportions. We quantified that less than 2% of the cells from one 50K-evolved clone acquired the fluorescence delivered by OMVs from the ancestral strain but that one cell concomitantly fuses with several OMVs. Globally, our results showed that OMV production in E. coli is a phenotype that varies along bacterial evolution and question the contribution of OMVs-mediated interactions in bacterial adaptation.},
keywords = {Cell Morphology, Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}
Favate J S; Liang S; Cope A L; Yadavilli S S; Shah P
The landscape of transcriptional and translational changes over 22 years of bacterial adaptation Journal Article
eLife, 11 , pp. e81979, 2022.
Abstract | Links | BibTeX | Altmetric | Tags: Genotypes and Phenotypes, Parallelism and Divergence
@article{favate2022,
title = {The landscape of transcriptional and translational changes over 22 years of bacterial adaptation},
author = {John S Favate and Shun Liang and Alexander L Cope and Srujana S Yadavilli and Premal Shah},
editor = {Detlef Weigel},
url = {https://elifesciences.org/articles/81979},
doi = {10.7554/eLife.81979},
year = {2022},
date = {2022-10-10},
urldate = {2022-10-10},
journal = {eLife},
volume = {11},
pages = {e81979},
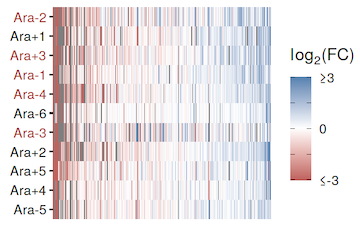
abstract = {Organisms can adapt to an environment by taking multiple mutational paths. This redundancy at the genetic level, where many mutations have similar phenotypic and fitness effects, can make untangling the molecular mechanisms of complex adaptations difficult. Here, we use the \textit{Escherichia coli} long-term evolution experiment (LTEE) as a model to address this challenge. To understand how different genomic changes could lead to parallel fitness gains, we characterize the landscape of transcriptional and translational changes across 12 replicate populations evolving in parallel for 50,000 generations. By quantifying absolute changes in mRNA abundances, we show that not only do all evolved lines have more mRNAs but that this increase in mRNA abundance scales with cell size. We also find that despite few shared mutations at the genetic level, clones from replicate populations in the LTEE are remarkably similar in their gene expression patterns at both the transcriptional and translational levels. Furthermore, we show that the majority of the expression changes are due to changes at the transcriptional level with very few translational changes. Finally, we show how mutations in transcriptional regulators lead to consistent and parallel changes in the expression levels of downstream genes. These results deepen our understanding of the molecular mechanisms underlying complex adaptations and provide insights into the repeatability of evolution.},
howpublished = {eLife},
keywords = {Genotypes and Phenotypes, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
Smith C E; Smith A N H; Cooper T F; Moore F B -G
Fitness of evolving bacterial populations is contingent on deep and shallow history but only shallow history creates predictable patterns Journal Article
Proceedings of the Royal Society B: Biological Sciences, 289 (1982), pp. 20221292, 2022.
Abstract | Links | BibTeX | Altmetric | Tags: Descendant Experiments, Fitness Trajectories, Genotypes and Phenotypes, Historical Contingency
@article{smith2022,
title = {Fitness of evolving bacterial populations is contingent on deep and shallow history but only shallow history creates predictable patterns},
author = {Chelsea E. Smith AND Adam N. H. Smith AND Tim F. Cooper AND Francisco B.-G. Moore},
url = {https://royalsocietypublishing.org/doi/10.1098/rspb.2022.1292},
doi = {10.1098/rspb.2022.1292},
year = {2022},
date = {2022-09-14},
journal = {Proceedings of the Royal Society B: Biological Sciences},
volume = {289},
number = {1982},
pages = {20221292},
abstract = {Long-term evolution experiments have tested the importance of genetic and environmental factors in influencing evolutionary outcomes. Differences in phylogenetic history, recent adaptation to distinct environments and chance events, all influence the fitness of a population. However, the interplay of these factors on a population's evolutionary potential remains relatively unexplored. We tracked the outcome of 2000 generations of evolution of four natural isolates of \textit{Escherichia coli} bacteria that were engineered to also create differences in shallow history by adding previously identified mutations selected in a separate long-term experiment. Replicate populations started from each progenitor evolved in four environments. We found that deep and shallow phylogenetic histories both contributed significantly to differences in evolved fitness, though by different amounts in different selection environments. With one exception, chance effects were not significant. Whereas the effect of deep history did not follow any detectable pattern, effects of shallow history followed a pattern of diminishing returns whereby fitter ancestors had smaller fitness increases. These results are consistent with adaptive evolution being contingent on the interaction of several evolutionary forces but demonstrate that the nature of these interactions is not fixed and may not be predictable even when the role of chance is small.},
keywords = {Descendant Experiments, Fitness Trajectories, Genotypes and Phenotypes, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
Mathé-Hubert H; Amie R; Martin M; Gaffé J; Schneider D
Evolution of bacterial persistence to antibiotics during a 50,000-generation experiment in an antibiotic-free environment Journal Article
Antibiotics, 11 (4), pp. 451, 2022, ISSN: 2079-6382.
Abstract | Links | BibTeX | Altmetric | Tags: Correlated Responses, Genotypes and Phenotypes
@article{mathe-hubert2022,
title = {Evolution of bacterial persistence to antibiotics during a 50,000-generation experiment in an antibiotic-free environment},
author = {Hugo Mathé-Hubert and Rafika Amie and Mikaël Martin and Joël Gaffé and Dominique Schneider},
url = {https://www.mdpi.com/2079-6382/11/4/451},
doi = {10.3390/antibiotics11040451},
issn = {2079-6382},
year = {2022},
date = {2022-03-27},
urldate = {2022-03-27},
journal = {Antibiotics},
volume = {11},
number = {4},
pages = {451},
abstract = {Failure of antibiotic therapies causes >700,000 deaths yearly and involves both bacterial resistance and persistence. Persistence results in the relapse of infections by producing a tiny fraction of pathogen survivors that stay dormant during antibiotic exposure. From an evolutionary perspective, persistence is either a ‘bet-hedging strategy’ that helps to cope with stochastically changing environments or an unavoidable minimal rate of ‘cellular errors’ that lock the cells in a low activity state. Here, we analyzed the evolution of persistence over 50,000 bacterial generations in a stable environment by improving a published method that estimates the number of persister cells based on the growth of the reviving population. Our results challenged our understanding of the factors underlying persistence evolution. In one case, we observed a substantial decrease in persistence proportion, suggesting that the naturally observed persistence level is not an unavoidable minimal rate of ‘cellular errors’. However, although there was no obvious environmental stochasticity, in 11 of the 12 investigated populations, the persistence level was maintained during 50,000 bacterial generations.},
keywords = {Correlated Responses, Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}
2021
van Raay K; Stolyar S; Sevigny J; Draghi J; Lenski R E; Marx C J; Kerr B; Zaman L
Evolution with private resources reverses some changes from long-term evolution with public resources Unpublished
bioRxiv, 2021.
Abstract | Links | BibTeX | Altmetric | Tags: Cell Morphology, Correlated Responses, Demography and Ecology, Descendant Experiments, Genotypes and Phenotypes, Historical Contingency, Methods and Miscellaneous
@unpublished{Raay2021,
title = {Evolution with private resources reverses some changes from long-term evolution with public resources},
author = {Katrina {van Raay} and Sergey Stolyar and Jordana Sevigny and Jeremy Draghi and Richard E. Lenski and Christopher J. Marx and Benjamin Kerr and Luis Zaman},
url = {https://www.biorxiv.org/content/10.1101/2021.07.11.451942v1},
doi = {https://doi.org/10.1101/2021.07.11.451942},
year = {2021},
date = {2021-07-12},
urldate = {2021-07-12},
journal = {bioRxiv},
pages = {2021.07.11.451942},
abstract = {A population under selection to improve one trait may evolve a sub-optimal state for another trait due to tradeoffs and other evolutionary constraints. How this evolution affects the capacity of a population to adapt when conditions change to favor the second trait is an open question. We investigated this question using isolates from a lineage spanning 60,000 generations of the Long-Term Evolution Experiment (LTEE) with \textit{Escherichia coli}, where cells have access to a shared pool of resources, and have evolved increased competitive ability and a concomitant reduction in numerical yield. Using media-in oil emulsions we shifted the focus of selection to numerical yield, where cells grew in isolated patches with private resources. We found that the time spent evolving under shared resources did not affect the ability to re-evolve toward higher numerical yield. The evolution of numerical yield commonly occurred through mutations in the phosphoenolpyruvate phosphotransferase system. These mutants exhibit slower uptake of glucose, making them poorer competitors for public resources, and produce smaller cells that release less carbon as overflow metabolites. Our results demonstrate that mutations that were not part of adaptation under one selective regime may enable access to ancestral phenotypes when selection changes to favor evolutionary reversion. },
howpublished = {bioRxiv},
keywords = {Cell Morphology, Correlated Responses, Demography and Ecology, Descendant Experiments, Genotypes and Phenotypes, Historical Contingency, Methods and Miscellaneous},
pubstate = {published},
tppubtype = {unpublished}
}
Grant N A; Maddamsetti R; Lenski R E
Maintenance of Metabolic Plasticity despite Relaxed Selection in a Long-Term Evolution Experiment with Escherichia coli Journal Article
The American Naturalist, 198 (1), pp. 93–112, 2021, ISSN: 0003-0147.
Abstract | Links | BibTeX | Altmetric | Tags: Correlated Responses, Fitness Trajectories, Genome Evolution, Genotypes and Phenotypes
@article{Grant2021b,
title = {Maintenance of Metabolic Plasticity despite Relaxed Selection in a Long-Term Evolution Experiment with \textit{Escherichia coli}},
author = {Nkrumah A. Grant and Rohan Maddamsetti and Richard E. Lenski},
url = {https://www.journals.uchicago.edu/doi/10.1086/714530},
doi = {10.1086/714530},
issn = {0003-0147},
year = {2021},
date = {2021-07-01},
urldate = {2021-07-01},
journal = {The American Naturalist},
volume = {198},
number = {1},
pages = {93--112},
abstract = {Traits that are unused in a given environment are subject to processes that tend to erode them, leading to reduced fitness in other environments. Although this general tendency is clear, we know much less about why some traits are lost while others are retained and about the roles of mutation and selection in generating different responses. We addressed these issues by examining populations of a facultative anaerobe, \textit{Escherichia coli}, that have evolved for 130 years in the presence of oxygen, with relaxed selection for anaerobic growth and the associated metabolic plasticity. We asked whether evolution led to the loss, improvement, or maintenance of anaerobic growth, and we analyzed gene expression and mutational data sets to understand the outcomes. We identified genomic signatures of both positive and purifying selection on aerobic-specific genes, while anaerobic-specific genes showed clear evidence of relaxed selection. We also found parallel evolution at two interacting loci that regulate anaerobic growth. We competed the ancestor and evolved clones from each population in an anoxic environment, and we found that anaerobic fitness had not decayed, despite relaxed selection. In summary, relaxed selection does not necessarily reduce an organism's fitness in other environments. Instead, the genetic architecture of the traits under relaxed selection and their correlations with traits under positive and purifying selection may sometimes determine evolutionary outcomes.},
keywords = {Correlated Responses, Fitness Trajectories, Genome Evolution, Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}
Karkare K; Lai H; Azevedo R B R; Cooper T F
Historical Contingency Causes Divergence in Adaptive Expression of the lac Operon Journal Article
Molecular Biology and Evolution, 38 (7), pp. 2869–2879, 2021, ISSN: 1537-1719.
Abstract | Links | BibTeX | Altmetric | Tags: Descendant Experiments, Genotypes and Phenotypes
@article{10.1093/molbev/msab077,
title = {Historical Contingency Causes Divergence in Adaptive Expression of the lac Operon},
author = {Kedar Karkare and Huei-Yi Lai and Ricardo B. R. Azevedo and Tim F. Cooper},
editor = {Patricia Wittkopp},
url = {https://academic.oup.com/mbe/article/38/7/2869/6179336},
doi = {10.1093/molbev/msab077},
issn = {1537-1719},
year = {2021},
date = {2021-06-01},
urldate = {2021-06-01},
journal = {Molecular Biology and Evolution},
volume = {38},
number = {7},
pages = {2869--2879},
abstract = {Populations of \textit{Escherichia coli} selected in constant and fluctuating environments containing lactose often adapt by substituting mutations in the lacI repressor that cause constitutive expression of the lac operon. These mutations occur at a high rate and provide a significant benefit. Despite this, eight of 24 populations evolved for 8,000 generations in environments containing lactose contained no detectable repressor mutations. We report here on the basis of this observation. We find that, given relevant mutation rates, repressor mutations are expected to have fixed in all evolved populations if they had maintained the same fitness effect they confer when introduced to the ancestor. In fact, reconstruction experiments demonstrate that repressor mutations have become neutral or deleterious in those populations in which they were not detectable. Populations not fixing repressor mutations nevertheless reached the same fitness as those that did fix them, indicating that they followed an alternative evolutionary path that made redundant the potential benefit of the repressor mutation, but involved unique mutations of equivalent benefit. We identify a mutation occurring in the promoter region of the uspB gene as a candidate for influencing the selective choice between these paths. Our results detail an example of historical contingency leading to divergent evolutionary outcomes.},
keywords = {Descendant Experiments, Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}
Grant N A; abdel Magid A; Franklin J; Dufour Y; Lenski R E
Changes in cell size and shape during 50,000 generations of experimental evolution with Escherichia coli Journal Article
Journal of Bacteriology, 203 (10), pp. e00469-20, 2021.
Abstract | Links | BibTeX | Altmetric | Tags: Cell Morphology, Genotypes and Phenotypes
@article{Grant2021,
title = {Changes in cell size and shape during 50,000 generations of experimental evolution with \textit{Escherichia coli}},
author = {Nkrumah A. Grant and Ali {abdel Magid} and Joshua Franklin and Yann Dufour and Richard E. Lenski},
url = {https://journals.asm.org/doi/10.1128/JB.00469-20},
doi = {10.1128/JB.00469-20},
year = {2021},
date = {2021-01-01},
urldate = {2021-01-01},
journal = {Journal of Bacteriology},
volume = {203},
number = {10},
pages = {e00469-20},
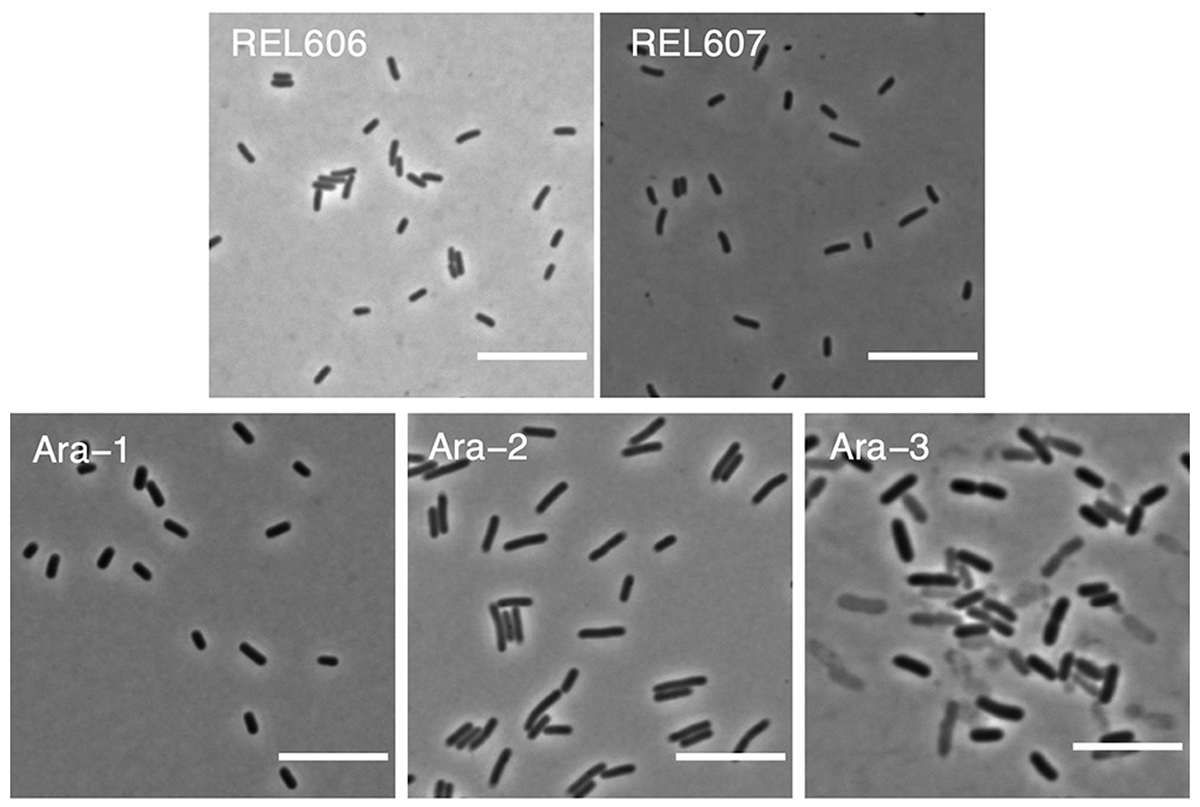
abstract = {Bacteria adopt a wide variety of sizes and shapes, with many species exhibiting stereotypical morphologies. How morphology changes, and over what timescales, is less clear. Previous work examining cell morphology in an experiment with \textit{Escherichia coli} showed that populations evolved larger cells and, in some cases, cells that were less rod-like. That experiment has now run for over two more decades. Meanwhile, genome sequence data are available for these populations, and new computational methods enable high-throughput microscopic analyses. In this study, we measured stationary-phase cell volumes for the ancestor and 12 populations at 2,000, 10,000, and 50,000 generations, including measurements during exponential growth at the last time point. We measured the distribution of cell volumes for each sample using a Coulter counter and microscopy, the latter of which also provided data on cell shape. Our data confirm the trend toward larger cells while also revealing substantial variation in size and shape across replicate populations. Most populations first evolved wider cells but later reverted to the ancestral length-to-width ratio. All but one population evolved mutations in rod shape maintenance genes. We also observed many ghost-like cells in the only population that evolved the novel ability to grow on citrate, supporting the hypothesis that this lineage struggles with maintaining balanced growth. Lastly, we show that cell size and fitness remain correlated across 50,000 generations. Our results suggest that larger cells are beneficial in the experimental environment, while the reversion toward ancestral length-to-width ratios suggests partial compensation for the less favorable surface area-to-volume ratios of the evolved cells.},
keywords = {Cell Morphology, Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}
Maddamsetti R
Universal Constraints on Protein Evolution in the Long-Term Evolution Experiment with Escherichia coli Journal Article
Genome Biology and Evolution, 13 (6), pp. evab070, 2021.
Abstract | Links | BibTeX | Altmetric | Tags: Genome Evolution, Genotypes and Phenotypes
@article{Maddamsetti2021b,
title = {Universal Constraints on Protein Evolution in the Long-Term Evolution Experiment with \textit{Escherichia coli}},
author = {Rohan Maddamsetti},
url = {https://academic.oup.com/gbe/article/13/6/evab070/6226398},
doi = {https://doi.org/10.1093/gbe/evab070},
year = {2021},
date = {2021-01-01},
urldate = {2021-01-01},
journal = {Genome Biology and Evolution},
volume = {13},
number = {6},
pages = {evab070},
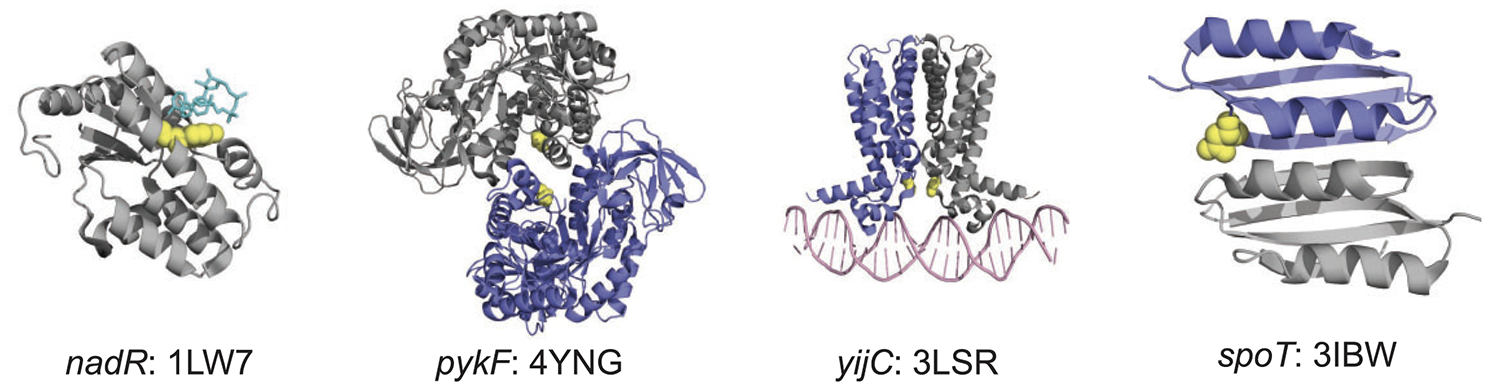
abstract = {Although it is well known that abundant proteins evolve slowly across the tree of life, there is little consensus for why this is true. Here, I report that abundant proteins evolve slowly in the hypermutator populations of Lenski’s long-term evolution experiment with \textit{Escherichia coli} (LTEE). Specifically, the density of all observed mutations per gene, as measured in metagenomic time series covering 60,000 generations of the LTEE, significantly anticorrelates with mRNA abundance, protein abundance, and degree of protein–protein interaction. The same pattern holds for nonsynonymous mutation density. However, synonymous mutation density, measured across the LTEE hypermutator populations, positively correlates with protein abundance. These results show that universal constraints on protein evolution are visible in data spanning three decades of experimental evolution. Therefore, it should be possible to design experiments to answer why abundant proteins evolve slowly.},
keywords = {Genome Evolution, Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}
Maddamsetti R
Selection Maintains Protein Interactome Resilience in the Long-Term Evolution Experiment with Escherichia coli Journal Article
Genome Biology and Evolution, 13 (6), pp. evab074, 2021.
Abstract | Links | BibTeX | Altmetric | Tags: Genome Evolution, Genotypes and Phenotypes
@article{Maddamsetti2021,
title = {Selection Maintains Protein Interactome Resilience in the Long-Term Evolution Experiment with \emph{Escherichia coli}},
author = {Rohan Maddamsetti},
url = {https://academic.oup.com/gbe/article/13/6/evab074/6240992},
doi = {https://doi.org/10.1093/gbe/evab074},
year = {2021},
date = {2021-01-01},
urldate = {2021-01-01},
journal = {Genome Biology and Evolution},
volume = {13},
number = {6},
pages = {evab074},
abstract = {Most cellular functions are carried out by a dynamic network of interacting proteins. An open question is whether the network properties of protein interactomes represent phenotypes under natural selection. One proposal is that protein interactomes have evolved to be resilient, such that they tend to maintain connectivity when proteins are removed from the network. This hypothesis predicts that interactome resilience should be maintained by natural selection during long-term experimental evolution. I tested this prediction by modeling the evolution of protein–protein interaction (PPI) networks in Lenski’s long-term evolution experiment with \textit{Escherichia coli} (LTEE). In this test, I removed proteins affected by nonsense, insertion, deletion, and transposon mutations in evolved LTEE strains, and measured the resilience of the resulting networks. I compared the rate of change of network resilience in each LTEE population to the rate of change of network resilience for corresponding randomized networks. The evolved PPI networks are significantly more resilient than networks in which random proteins have been deleted. Moreover, the evolved networks are generally more resilient than networks in which the random deletion of proteins was restricted to those disrupted in LTEE. These results suggest that evolution in the LTEE has favored PPI networks that are, on average, more resilient than expected from the genetic variation across the evolved strains. My findings therefore support the hypothesis that selection maintains protein interactome resilience over evolutionary time.},
keywords = {Genome Evolution, Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}
2020
Blount Z D; Maddamsetti R; Grant N A; Ahmed S T; Jagdish T; Baxter J A; Sommerfeld B A; Tillman A; Moore J; Slonczewski J L; Barrick J E; Lenski R E
Genomic and phenotypic evolution of Escherichia coli in a novel citrate-only resource environment Journal Article
eLife, 9 , pp. 1–64, 2020, ISSN: 2050-084X.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Demography and Ecology, Descendant Experiments, Genotypes and Phenotypes
@article{Blount2020,
title = {Genomic and phenotypic evolution of \textit{Escherichia coli} in a novel citrate-only resource environment},
author = {Zachary D. Blount and Rohan Maddamsetti and Nkrumah A. Grant and Sumaya T. Ahmed and Tanush Jagdish and Jessica A. Baxter and Brooke A. Sommerfeld and Alice Tillman and Jeremy Moore and Joan L. Slonczewski and Jeffrey E. Barrick and Richard E. Lenski},
url = {https://elifesciences.org/articles/55414},
doi = {10.7554/eLife.55414},
issn = {2050-084X},
year = {2020},
date = {2020-05-01},
urldate = {2020-05-01},
journal = {eLife},
volume = {9},
pages = {1--64},
abstract = {Evolutionary innovations allow populations to colonize new ecological niches. We previously reported that aerobic growth on citrate (Cit^{+}) evolved in an \textit{Escherichia coli} population during adaptation to a minimal glucose medium containing citrate (DM25). Cit^{+} variants can also grow in citrate-only medium (DM0), a novel environment for \textit{E. coli}. To study adaptation to this niche, we founded two sets of Cit^{+} populations and evolved them for 2500 generations in DM0 or DM25. The evolved lineages acquired numerous parallel mutations, many mediated by transposable elements. Several also evolved amplifications of regions containing the \textit{maeA} gene. Unexpectedly, some evolved populations and clones show apparent declines in fitness. We also found evidence of substantial cell death in Cit^{+} clones. Our results thus demonstrate rapid trait refinement and adaptation to the new citrate niche, while also suggesting a recalcitrant mismatch between \textit{E. coli} physiology and growth on citrate.},
keywords = {Citrate Evolution, Demography and Ecology, Descendant Experiments, Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}
2018
Peng F; Widmann S; Wünsche A; Duan K; Donovan K A; Dobson R C J; Lenski R E; Cooper T F
Effects of Beneficial Mutations in pykF Gene Vary over Time and across Replicate Populations in a Long-Term Experiment with Bacteria Journal Article
Molecular Biology and Evolution, 35 (1), pp. 202–210, 2018, ISSN: 0737-4038.
Abstract | Links | BibTeX | Altmetric | Tags: Genotypes and Phenotypes, Historical Contingency
@article{Peng2018,
title = {Effects of Beneficial Mutations in \textit{pykF} Gene Vary over Time and across Replicate Populations in a Long-Term Experiment with Bacteria},
author = {Fen Peng and Scott Widmann and Andrea Wünsche and Kristina Duan and Katherine A. Donovan and Renwick C J Dobson and Richard E. Lenski and Tim F. Cooper},
url = {https://academic.oup.com/mbe/article/35/1/202/4562833},
doi = {10.1093/molbev/msx279},
issn = {0737-4038},
year = {2018},
date = {2018-01-01},
urldate = {2018-01-01},
journal = {Molecular Biology and Evolution},
volume = {35},
number = {1},
pages = {202--210},
abstract = {The fitness effects of mutations can depend on the genetic backgrounds in which they occur and thereby influence future opportunities for evolving populations. In particular, mutations that fix in a population might change the selective benefit of subsequent mutations, giving rise to historical contingency. We examine these effects by focusing on mutations in a key metabolic gene, \textit{pykF}, that arose independently early in the history of 12 \textit{Escherichia coli} populations during a long-Term evolution experiment. Eight different evolved nonsynonymous mutations conferred similar fitness benefits of ~10% when transferred into the ancestor, and these benefits were greater than the one conferred by a deletion mutation. In contrast, the same mutations had highly variable fitness effects, ranging from ~0% to 25%, in evolved clones isolated from the populations at 20,000 generations. Two mutations that were moved into these evolved clones conferred similar fitness effects in a given clone, but different effects between the clones, indicating epistatic interactions between the evolved \textit{pykF} alleles and the other mutations that had accumulated in each evolved clone. We also measured the fitness effects of six evolved \textit{pykF} alleles in the same populations in which they had fixed, but at seven time points between 0 and 50,000 generations. Variation in fitness effects was high at intermediate time points, and declined to a low level at 50,000 generations, when the mean fitness effect was lowest. Our results demonstrate the importance of genetic context in determining the fitness effects of different beneficial mutations even within the same gene.},
keywords = {Genotypes and Phenotypes, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
2017
Maddamsetti R; Hatcher P J; Green A G; Williams B L; Marks D S; Lenski R E
Core genes evolve rapidly in the long-term evolution experiment with Escherichia coli Journal Article
Genome Biology and Evolution, 9 (4), pp. 1072–1083, 2017, ISSN: 17596653.
Abstract | Links | BibTeX | Altmetric | Tags: Genome Evolution, Genotypes and Phenotypes, Mutation Rates
@article{Maddamsetti2017,
title = {Core genes evolve rapidly in the long-term evolution experiment with \textit{Escherichia coli}},
author = {Rohan Maddamsetti and Philip J. Hatcher and Anna G. Green and Barry L. Williams and Debora S. Marks and Richard E. Lenski},
url = {https://academic.oup.com/gbe/article/9/4/1072/3100447},
doi = {10.1093/gbe/evx064},
issn = {17596653},
year = {2017},
date = {2017-01-01},
urldate = {2017-01-01},
journal = {Genome Biology and Evolution},
volume = {9},
number = {4},
pages = {1072--1083},
abstract = {Bacteria can evolve rapidly under positive selection owing to their vast numbers, allowing their genes to diversify by adapting to different environments. We asked whether the same genes that evolve rapidly in the long-term evolution experiment (LTEE) with \textit{Escherichia coli} have also diversified extensively in nature. To make this comparison, we identified ~2000 core genes shared among 60 \textit{E. coli} strains. During the LTEE, core genes accumulated significantly more nonsynonymous mutations than flexible (i.e., noncore) genes. Furthermore, core genes under positive selection in the LTEE are more conserved in nature than the average core gene. In some cases, adaptive mutations appear to modify protein functions, rather than merely knocking them out. The LTEE conditions are novel for \textit{E. coli}, at least in relation to its evolutionary history in nature. The constancy and simplicity of the environment likely favor the complete loss of some unused functions and the fine-tuning of others.},
keywords = {Genome Evolution, Genotypes and Phenotypes, Mutation Rates},
pubstate = {published},
tppubtype = {article}
}
2015
Quandt E M; Gollihar J; Blount Z D; Ellington A D; Georgiou G; Barrick J E
Fine-tuning citrate synthase flux potentiates and refines metabolic innovation in the Lenski evolution experiment. Journal Article
eLife, 4 (October), pp. e09696, 2015, ISSN: 2050-084X.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Genotypes and Phenotypes, Historical Contingency
@article{Quandt2015,
title = {Fine-tuning citrate synthase flux potentiates and refines metabolic innovation in the Lenski evolution experiment.},
author = {Erik M. Quandt and Jimmy Gollihar and Zachary D. Blount and Andrew D. Ellington and George Georgiou and Jeffrey E. Barrick},
url = {http://www.ncbi.nlm.nih.gov/pubmed/26465114
http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=PMC4718724},
doi = {10.7554/eLife.09696},
issn = {2050-084X},
year = {2015},
date = {2015-10-01},
urldate = {2015-10-01},
journal = {eLife},
volume = {4},
number = {October},
pages = {e09696},
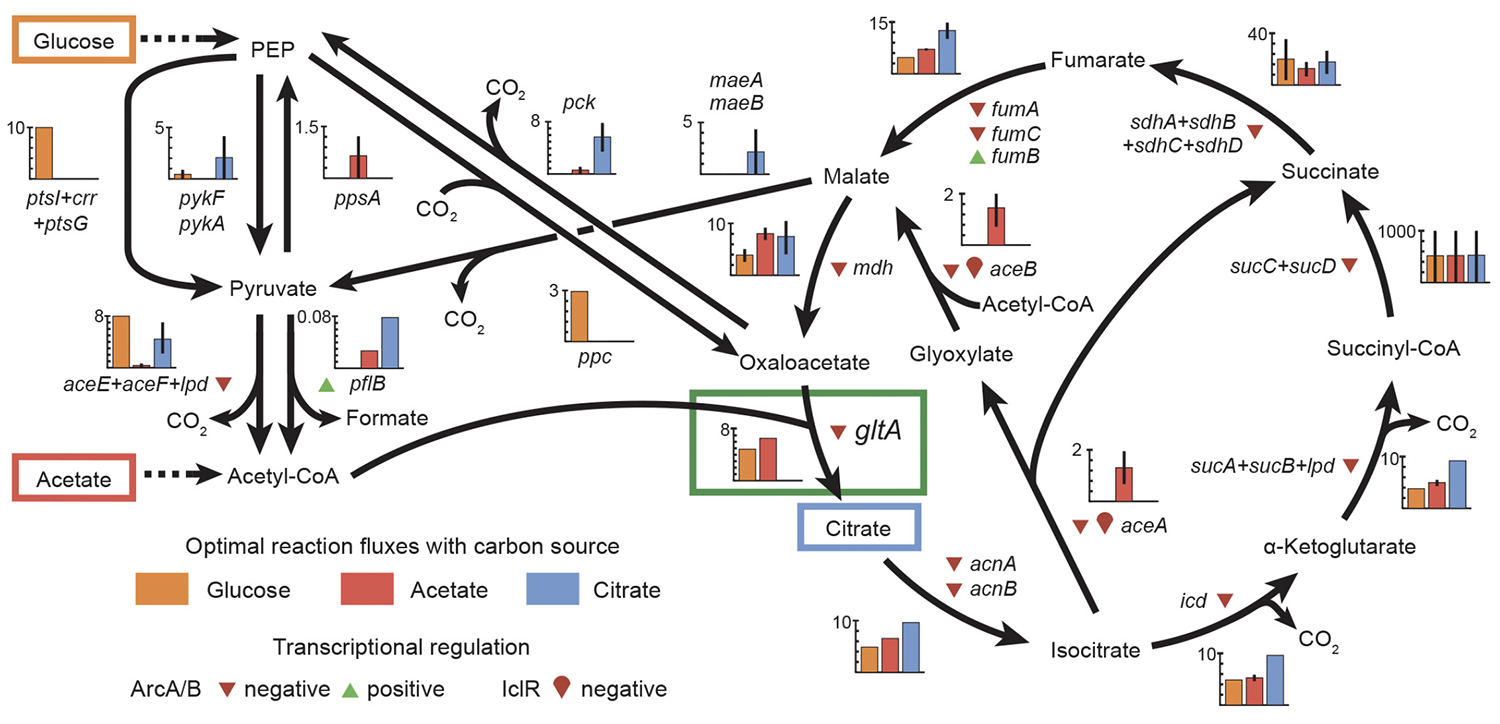
abstract = {Evolutionary innovations that enable organisms to colonize new ecological niches are rare compared to gradual evolutionary changes in existing traits. We discovered that key mutations in the \textit{gltA} gene, which encodes citrate synthase (CS), occurred both before and after \textit{Escherichia coli} gained the ability to grow aerobically on citrate (Cit(+) phenotype) during the Lenski long-term evolution experiment. The first \textit{gltA} mutation, which increases CS activity by disrupting NADH-inhibition of this enzyme, is beneficial for growth on the acetate and contributed to preserving the rudimentary Cit(+) trait from extinction when it first evolved. However, after Cit(+) was refined by further mutations, this potentiating \textit{gltA} mutation became deleterious to fitness. A second wave of beneficial \textit{gltA} mutations then evolved that reduced CS activity to below the ancestral level. Thus, dynamic reorganization of central metabolism made colonizing this new nutrient niche contingent on both co-opting and overcoming a history of prior adaptation.},
keywords = {Citrate Evolution, Genotypes and Phenotypes, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
Rudan M; Schneider D; Warnecke T; Krisko A
RNA chaperones buffer deleterious mutations in E. coli Journal Article
eLife, 4 , 2015, ISSN: 2050-084X.
Abstract | Links | BibTeX | Altmetric | Tags: Genotypes and Phenotypes
@article{Rudan2015,
title = {RNA chaperones buffer deleterious mutations in \textit{E. coli}},
author = {Marina Rudan and Dominique Schneider and Tobias Warnecke and Anita Krisko},
url = {https://elifesciences.org/articles/04745},
doi = {10.7554/eLife.04745},
issn = {2050-084X},
year = {2015},
date = {2015-03-01},
urldate = {2015-03-01},
journal = {eLife},
volume = {4},
abstract = {Both proteins and RNAs can misfold into non-functional conformations. Protein chaperones promote native folding of nascent polypeptides and refolding of misfolded species, thereby buffering mutations that compromise protein structure and function. Here, we show that RNA chaperones can also act as mutation buffers that enhance organismal fitness. Using competition assays, we demonstrate that overexpression of select RNA chaperones, including three DEAD box RNA helicases (DBRHs) (CsdA, SrmB, RhlB) and the cold shock protein CspA, improves fitness of two independently evolved \textit{Escherichia coli} mutator strains that have accumulated deleterious mutations during short- and long-term laboratory evolution. We identify strain-specific mutations that are deleterious and subject to buffering when introduced individually into the ancestral genotype. For DBRHs, we show that buffering requires helicase activity, implicating RNA structural remodelling in the buffering process. Our results suggest that RNA chaperones might play a fundamental role in RNA evolution and evolvability.},
keywords = {Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}