2025
Chihoub D; Pintard C; Lenski R E; Tenaillon O; Couce A
The evolution of robustness and fragility during long-term bacterial adaptation Unpublished
2025.
Abstract | Links | BibTeX | Altmetric | Tags: Correlated Responses, Fitness Trajectories
@unpublished{Chihoub2025,
title = {The evolution of robustness and fragility during long-term bacterial adaptation},
author = {Doha Chihoub and Coralie Pintard and Richard E. Lenski and Olivier Tenaillon and Alejandro Couce},
doi = {10.1101/2025.01.24.632760},
year = {2025},
date = {2025-01-27},
urldate = {2025-01-27},
abstract = {Most mutations affecting fitness are harmful, and their inevitable occurrence reduces mean population fitness. Theory predicts that well-adapted populations may evolve mechanisms to minimize this deleterious load. Direct selection to increase mutational robustness can be achieved in the laboratory. However, its spontaneous evolution during general adaptation remains uncertain, with mixed evidence across model systems. Here, we studied the effects of highly pleiotropic point mutations in \textit{Escherichia coli} over a 15,000-generation adaptive trajectory. The fitness effects of both beneficial and deleterious mutations were attenuated with increased adaptation over time. In contrast, pleiotropic effects in new environments became more severe and widespread with greater adaptation. These results show that trade-offs between robustness and fragility can rapidly evolve in regulatory networks, regardless of whether driven by adaptive or non-adaptive processes. More broadly, these results show that adaptation can generate a hidden potential for phenotypic diversity, unpredictably shaping evolutionary prospects in new environments.
},
keywords = {Correlated Responses, Fitness Trajectories},
pubstate = {published},
tppubtype = {unpublished}
}
2024
Izutsu M; Lake D M; Matson Z W D; Dodson J P; Lenski R E
Effects of periodic bottlenecks on the dynamics of adaptive evolution in microbial populations Journal Article
Microbiology (Reading, England), 170 (9), pp. 001494, 2024, ISSN: 1465-2080.
Abstract | Links | BibTeX | Altmetric | Tags: Descendant Experiments, Fitness Trajectories, Theory and Simulations
@article{Izutsu_2024,
title = {Effects of periodic bottlenecks on the dynamics of adaptive evolution in microbial populations},
author = {Minako Izutsu and Devin M. Lake and Zachary W. D. Matson and Jack P. Dodson and Richard E. Lenski},
doi = {10.1099/mic.0.001494},
issn = {1465-2080},
year = {2024},
date = {2024-09-18},
urldate = {2021-12-30},
journal = {Microbiology (Reading, England)},
volume = {170},
number = {9},
pages = {001494},
publisher = {LTEE},
abstract = {Population bottlenecks can impact the rate of adaptation in evolving populations. On the one hand, each bottleneck reduces the genetic variation that fuels adaptation. On the other hand, each founder that survives a bottleneck can undergo more generations and leave more descendants in a resource-limited environment, which allows surviving beneficial mutations to spread more quickly. A theoretical model predicted that the rate of fitness gains should be maximized using ~8-fold dilutions. Here we investigate the impact of repeated bottlenecks on the dynamics of adaptation using numerical simulations and experimental populations of Escherichia coli. Our simulations confirm the model's prediction when populations evolve in a regime where beneficial mutations are rare and waiting times between successful mutations are long. However, more extreme dilutions maximize fitness gains in simulations when beneficial mutations are common and clonal interference prevents most of them from fixing. To examine these predictions, we propagated 48 \textit{E. coli} populations with 2-, 8-, 100-, and 1000-fold dilutions for 150 days. Adaptation began earlier and fitness gains were greater with 100- and 1000-fold dilutions than with 8-fold dilutions, consistent with the simulations when beneficial mutations are common. However, the selection pressures in the 2-fold treatment were qualitatively different from the other treatments, violating a critical assumption of the model and simulations. Thus, varying the dilution factor during periodic bottlenecks can have multiple effects on the dynamics of adaptation caused by differential losses of diversity, different numbers of generations, and altered selection.},
keywords = {Descendant Experiments, Fitness Trajectories, Theory and Simulations},
pubstate = {published},
tppubtype = {article}
}
2023
Barrick J E; Blount Z D; Lake D M; Dwenger J H; Chavarria-Palma J E; Izutsu M; Wiser M J
Daily Transfers, Archiving Populations, and Measuring Fitness in the Long-Term Evolution Experiment with Escherichia coli Journal Article
Journal of Visualized Experiments, 198 , pp. e65342, 2023, ISSN: 1940-087X.
Abstract | Links | BibTeX | Altmetric | Tags: Fitness Trajectories, Methods and Miscellaneous
@article{Barrick2023,
title = {Daily Transfers, Archiving Populations, and Measuring Fitness in the Long-Term Evolution Experiment with Escherichia coli},
author = {Jeffrey E. Barrick and Zachary D. Blount and Devin M. Lake and Jack H. Dwenger and Jesus E. Chavarria-Palma and Minako Izutsu and Michael J. Wiser},
doi = {10.3791/65342},
issn = {1940-087X},
year = {2023},
date = {2023-08-18},
urldate = {2023-08-18},
journal = {Journal of Visualized Experiments},
volume = {198},
pages = {e65342},
abstract = {The Long-Term Evolution Experiment (LTEE) has followed twelve populations of \textit{Escherichia coli} as they have adapted to a simple laboratory environment for more than 35 years and 77,000 bacterial generations. The setup and procedures used in the LTEE epitomize reliable and reproducible methods for studying microbial evolution. In this protocol, we first describe how the LTEE populations are transferred to fresh medium and cultured each day. Then, we describe how the LTEE populations are regularly checked for possible signs of contamination and archived to provide a permanent frozen "fossil record" for later study. Multiple safeguards included in these procedures are designed to prevent contamination, detect various problems when they occur, and recover from disruptions without appreciably setting back the progress of the experiment. One way that the overall tempo and character of evolutionary changes are monitored in the LTEE is by measuring the competitive fitness of populations and strains from the experiment. We describe how co-culture competition assays are conducted and provide both a spreadsheet and an R package (fitnessR) for calculating relative fitness from the results. Over the course of the LTEE, the behaviors of some populations have changed in interesting ways, and new technologies like whole-genome sequencing have provided additional avenues for investigating how the populations have evolved. We end by discussing how the original LTEE procedures have been updated to accommodate or take advantage of these changes. This protocol will be useful for researchers who use the LTEE as a model system for studying connections between evolution and genetics, molecular biology, systems biology, and ecology. More broadly, the LTEE provides a tried-and-true template for those who are beginning their own evolution experiments with new microbes, environments, and questions. },
keywords = {Fitness Trajectories, Methods and Miscellaneous},
pubstate = {published},
tppubtype = {article}
}
Lenski R E
Revisiting the design of the long-term evolution experiment with Escherichia coli Journal Article
Journal of Molecular Evolution, 91 (3), pp. 241-253, 2023.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Descendant Experiments, Fitness Trajectories, Genome Evolution, Methods and Miscellaneous, Review Articles
@article{lenski2023,
title = {Revisiting the design of the long-term evolution experiment with \textit{Escherichia coli}},
author = {Richard E. Lenski},
url = {https://link.springer.com/epdf/10.1007/s00239-023-10095-3?sharing_token=zmDHuK0kbvnJBQq1k96fe_e4RwlQNchNByi7wbcMAY53KNkhv6F2YgRIeC8sZGNejxJrvlAGZWInruED5Dqdai5WeU2RAWL2PJNp0pL9QJO39B_ijCtRZcaW8jqM7PclDJfFwL_78U5zNlQYyCOsQwa1Yxha61uXUWhW-Buiq7o=},
doi = {10.1007/s00239-023-10095-3},
year = {2023},
date = {2023-06-01},
urldate = {2023-02-15},
journal = {Journal of Molecular Evolution},
volume = {91},
number = {3},
pages = {241-253},
abstract = {The long-term evolution experiment (LTEE) with \textit{Escherichia coli} began in 1988 and it continues to this day, with its 12 populations having recently reached 75,000 generations of evolution in a simple, well-controlled environment. The LTEE was designed to explore open-ended questions about the dynamics and repeatability of phenotypic and genetic evolution. Here I discuss various aspects of the LTEE’s experimental design that have enabled its stability and success, including the choices of the culture regime, growth medium, ancestral strain, and statistical replication. I also discuss some of the challenges associated with a long-running project, such as handling procedural errors (e.g., cross-contamination) and managing the expanding collection of frozen samples. The simplicity of the experimental design and procedures have supported the long-term stability of the LTEE. That stability—along with the inherent creativity of the evolutionary process and the emergence of new genomic technologies—provides a platform that has allowed talented students and collaborators to pose questions, collect data, and make discoveries that go far beyond anything I could have imagined at the start of the LTEE.},
keywords = {Citrate Evolution, Descendant Experiments, Fitness Trajectories, Genome Evolution, Methods and Miscellaneous, Review Articles},
pubstate = {published},
tppubtype = {article}
}
Jagdish T
The dynamics of fitness and pleiotropy in a long-term evolution experiment with Escherichia coli PhD Thesis
2023.
Abstract | Links | BibTeX | Tags: Correlated Responses, Demography and Ecology, Fitness Trajectories, Methods and Miscellaneous
@phdthesis{nokey,
title = {The dynamics of fitness and pleiotropy in a long-term evolution experiment with \textit{Escherichia coli}},
author = {Tanush Jagdish},
url = {https://dash.harvard.edu/handle/1/37375541},
year = {2023},
date = {2023-05-08},
urldate = {2023-05-08},
abstract = {Life on earth is shaped by a delicate balance between chance and necessity. A combination of natural selection and genetic drift has moulded random genetic variations over the course of billions of years to generate the complex and interconnected biosphere we see today. These evolutionary dynamics are ultimately the product of numerous genetic mechanisms operating in genomes with highly complex architectures. Unravelling the rules of genetic mechanisms responsible for evolutionary innovation is critical to developing a comprehensive and predictive evolutionary theory, but has evaded direct experimentation since the scale of resources and technology needed to study the patterns of genetic evolution has only recently become achievable. In this dissertation, I explore the use of microbial experimental evolution as a powerful tool for probing evolutionary questions, particularly by leveraging DNA-barcoding and high-throughput DNA sequencing. In chapter 1, I provide an overview of the field of microbial experimental evolution, dividing its history into two eras marked by qualitatively different methodological advancements. I make the case that we now stand at the dawn of a third era, where advances in genome-engineering coupled with low-cost, high-throughput DNA sequencing will allow experiments to finally probe evolution on a statistical scale. I then present two research studies that take advantage of microbial experimental evolution to investigate distinct genetic mechanisms key to the evolutionary process. In Chapter 2, I explore whether fitness can continue increasing in a population that has already adapted to its environment for over 30,000 generations, and whether fixations of beneficial mutations from population-wide DNA sequencing can predict jumps in fitness. My findings reveal that fitness continues to monotonically increase in step with the fixation of beneficial mutations, even though the rate of fixation has dramatically slowed down, highlighting the potential for ongoing adaptation even under constant environmental conditions. In Chapter 3, I develop a novel conjugation-based DNA-barcoding method for the Long-Term Evolution Experiment (LTEE) with \textit{Escherichia coli}, allowing me to examine the pleiotropic consequences of adaptation to glucose over 50,000 generations in 15 novel resource environments. My observations reveal broad patterns of both convergent and divergent evolution that correspond with mutations in key metabolic genes in clonal sequencing datasets, shedding light on the nature of pleiotropy and its evolution over extended timescales. Using microbial model systems in an evolutionary context has the unique advantage of being relevant both to fundamental evolutionary biology and human health. Since genetic drift and rare mutational events both play an outsized role in determining the evolutionary trajectories of populations, evolutionary questions in the modern age will increasingly be faced with issues of scale. Microbial experimental evolution offers both scale and tractability to solve this problem. Uniquely, this does not sacrifice on human relevance. Building a coarse-grained and comprehensive evolutionary theory is more significant to society today than ever before as the importance of clonal evolution in cancer, gut microbiomes and even pandemics becomes more clearly understood.},
keywords = {Correlated Responses, Demography and Ecology, Fitness Trajectories, Methods and Miscellaneous},
pubstate = {published},
tppubtype = {phdthesis}
}
2022
Smith C E; Smith A N H; Cooper T F; Moore F B -G
Fitness of evolving bacterial populations is contingent on deep and shallow history but only shallow history creates predictable patterns Journal Article
Proceedings of the Royal Society B: Biological Sciences, 289 (1982), pp. 20221292, 2022.
Abstract | Links | BibTeX | Altmetric | Tags: Descendant Experiments, Fitness Trajectories, Genotypes and Phenotypes, Historical Contingency
@article{smith2022,
title = {Fitness of evolving bacterial populations is contingent on deep and shallow history but only shallow history creates predictable patterns},
author = {Chelsea E. Smith AND Adam N. H. Smith AND Tim F. Cooper AND Francisco B.-G. Moore},
url = {https://royalsocietypublishing.org/doi/10.1098/rspb.2022.1292},
doi = {10.1098/rspb.2022.1292},
year = {2022},
date = {2022-09-14},
journal = {Proceedings of the Royal Society B: Biological Sciences},
volume = {289},
number = {1982},
pages = {20221292},
abstract = {Long-term evolution experiments have tested the importance of genetic and environmental factors in influencing evolutionary outcomes. Differences in phylogenetic history, recent adaptation to distinct environments and chance events, all influence the fitness of a population. However, the interplay of these factors on a population's evolutionary potential remains relatively unexplored. We tracked the outcome of 2000 generations of evolution of four natural isolates of \textit{Escherichia coli} bacteria that were engineered to also create differences in shallow history by adding previously identified mutations selected in a separate long-term experiment. Replicate populations started from each progenitor evolved in four environments. We found that deep and shallow phylogenetic histories both contributed significantly to differences in evolved fitness, though by different amounts in different selection environments. With one exception, chance effects were not significant. Whereas the effect of deep history did not follow any detectable pattern, effects of shallow history followed a pattern of diminishing returns whereby fitter ancestors had smaller fitness increases. These results are consistent with adaptive evolution being contingent on the interaction of several evolutionary forces but demonstrate that the nature of these interactions is not fixed and may not be predictable even when the role of chance is small.},
keywords = {Descendant Experiments, Fitness Trajectories, Genotypes and Phenotypes, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
2021
Consuegra J; Gaffé J; Lenski R E; Hindré T; Barrick J E; Tenaillon O; Schneider D
Insertion-sequence-mediated mutations both promote and constrain evolvability during a long-term experiment with bacteria Journal Article
Nature Communications, 12 (1), pp. 980, 2021, ISSN: 2041-1723.
Abstract | Links | BibTeX | Altmetric | Tags: Fitness Trajectories, Genome Evolution, Mutation Rates
@article{Consuegra2021,
title = {Insertion-sequence-mediated mutations both promote and constrain evolvability during a long-term experiment with bacteria},
author = {Jessika Consuegra and Joël Gaffé and Richard E. Lenski and Thomas Hindré and Jeffrey E. Barrick and Olivier Tenaillon and Dominique Schneider},
url = {http://www.nature.com/articles/s41467-021-21210-7},
doi = {https://doi.org/10.1038/s41467-021-21210-7},
issn = {2041-1723},
year = {2021},
date = {2021-12-01},
urldate = {2021-12-01},
journal = {Nature Communications},
volume = {12},
number = {1},
pages = {980},
publisher = {Springer US},
abstract = {Insertion sequences (IS) are ubiquitous bacterial mobile genetic elements, and the mutations they cause can be deleterious, neutral, or beneficial. The long-term dynamics of IS elements and their effects on bacteria are poorly understood, including whether they are primarily genomic parasites or important drivers of adaptation by natural selection. Here, we investigate the dynamics of IS elements and their contribution to genomic evolution and fitness during a long-term experiment with \textit{Escherichia coli}. IS elements account for ~35% of the mutations that reached high frequency through 50,000 generations in those populations that retained the ancestral point-mutation rate. In mutator populations, IS-mediated mutations are only half as frequent in absolute numbers. In one population, an exceptionally high ~8-fold increase in IS 150 copy number is associated with the beneficial effects of early insertion mutations; however, this expansion later slowed down owing to reduced IS 150 activity. This population also achieves the lowest fitness, suggesting that some avenues for further adaptation are precluded by the IS 150 -mediated mutations. More generally, across all populations, we find that higher IS activity becomes detrimental to adaptation over evolutionary time. Therefore, IS-mediated mutations can both promote and constrain evolvability.},
keywords = {Fitness Trajectories, Genome Evolution, Mutation Rates},
pubstate = {published},
tppubtype = {article}
}
Deatherage D E; Barrick J E
High-throughput characterization of mutations in genes that drive clonal evolution using multiplex adaptome capture sequencing Journal Article
Cell Systems, 12 (12), pp. 1187-1200, 2021, ISSN: 24054712.
Abstract | Links | BibTeX | Altmetric | Tags: Descendant Experiments, Fitness Trajectories, Methods and Miscellaneous
@article{DEATHERAGE2021,
title = {High-throughput characterization of mutations in genes that drive clonal evolution using multiplex adaptome capture sequencing},
author = {Daniel E. Deatherage and Jeffrey E. Barrick},
url = {https://www.ncbi.nlm.nih.gov/pmc/articles/PMC8678185/},
doi = {10.1016/j.cels.2021.08.011},
issn = {24054712},
year = {2021},
date = {2021-09-01},
urldate = {2021-09-01},
journal = {Cell Systems},
volume = {12},
number = {12},
pages = {1187-1200},
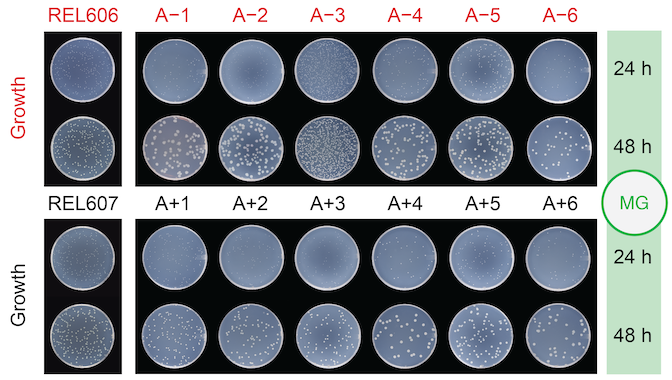
abstract = {Understanding how cells are likely to evolve can guide medical interventions and bioengineering efforts that must contend with unwanted mutations. The adaptome of a cell—the neighborhood of genetic changes that are most likely to drive adaptation in a given environment—can be mapped by tracking rare beneficial variants during the early stages of clonal evolution. We used multiplex adaptome capture sequencing (mAdCap-seq), a procedure that combines unique molecular identifiers and hybridization-based enrichment, to characterize mutations in eight \textit{Escherichia coli} genes known to be under selection in a laboratory environment. We tracked 301 mutations at frequencies as low as 0.01% and inferred the fitness effects of 240 of these mutations. There were distinct molecular signatures of selection on protein structure and function for the three genes with the most beneficial mutations. Our results demonstrate how mAdCap-seq can be used to deeply profile a targeted portion of a cell's adaptome.},
keywords = {Descendant Experiments, Fitness Trajectories, Methods and Miscellaneous},
pubstate = {published},
tppubtype = {article}
}
Grant N A; Maddamsetti R; Lenski R E
Maintenance of Metabolic Plasticity despite Relaxed Selection in a Long-Term Evolution Experiment with Escherichia coli Journal Article
The American Naturalist, 198 (1), pp. 93–112, 2021, ISSN: 0003-0147.
Abstract | Links | BibTeX | Altmetric | Tags: Correlated Responses, Fitness Trajectories, Genome Evolution, Genotypes and Phenotypes
@article{Grant2021b,
title = {Maintenance of Metabolic Plasticity despite Relaxed Selection in a Long-Term Evolution Experiment with \textit{Escherichia coli}},
author = {Nkrumah A. Grant and Rohan Maddamsetti and Richard E. Lenski},
url = {https://www.journals.uchicago.edu/doi/10.1086/714530},
doi = {10.1086/714530},
issn = {0003-0147},
year = {2021},
date = {2021-07-01},
urldate = {2021-07-01},
journal = {The American Naturalist},
volume = {198},
number = {1},
pages = {93--112},
abstract = {Traits that are unused in a given environment are subject to processes that tend to erode them, leading to reduced fitness in other environments. Although this general tendency is clear, we know much less about why some traits are lost while others are retained and about the roles of mutation and selection in generating different responses. We addressed these issues by examining populations of a facultative anaerobe, \textit{Escherichia coli}, that have evolved for 130 years in the presence of oxygen, with relaxed selection for anaerobic growth and the associated metabolic plasticity. We asked whether evolution led to the loss, improvement, or maintenance of anaerobic growth, and we analyzed gene expression and mutational data sets to understand the outcomes. We identified genomic signatures of both positive and purifying selection on aerobic-specific genes, while anaerobic-specific genes showed clear evidence of relaxed selection. We also found parallel evolution at two interacting loci that regulate anaerobic growth. We competed the ancestor and evolved clones from each population in an anoxic environment, and we found that anaerobic fitness had not decayed, despite relaxed selection. In summary, relaxed selection does not necessarily reduce an organism's fitness in other environments. Instead, the genetic architecture of the traits under relaxed selection and their correlations with traits under positive and purifying selection may sometimes determine evolutionary outcomes.},
keywords = {Correlated Responses, Fitness Trajectories, Genome Evolution, Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}
2020
Barrick J E; Deatherage D E; Lenski R E
Banzhaf, Wolfgang; Cheng, Betty H C; Deb, Kalyanmoy; Holekamp, Kay E; Lenski, Richard E; Ofria, Charles; Pennock, Robert T; Punch, William F; Whittaker, Danielle J (Ed.): Evolution in Action: Past, Present and Future: A Festschrift in Honor of Erik D. Goodman, pp. 77–89, Springer International Publishing, Cham, 2020, ISBN: 978-3-030-39831-6.
Abstract | Links | BibTeX | Altmetric | Tags: Fitness Trajectories
@inbook{Barrick2020,
title = {A Test of the Repeatability of Measurements of Relative Fitness in the Long-Term Evolution Experiment with \textit{Escherichia coli}},
author = {Jeffrey E. Barrick and Daniel E. Deatherage and Richard E. Lenski},
editor = {Wolfgang Banzhaf and Betty H C Cheng and Kalyanmoy Deb and Kay E Holekamp and Richard E Lenski and Charles Ofria and Robert T Pennock and William F Punch and Danielle J Whittaker},
url = {http://link.springer.com/10.1007/978-3-030-39831-6_8},
doi = {10.1007/978-3-030-39831-6_8},
isbn = {978-3-030-39831-6},
year = {2020},
date = {2020-01-01},
urldate = {2020-01-01},
booktitle = {Evolution in Action: Past, Present and Future: A Festschrift in Honor of Erik D. Goodman},
pages = {77--89},
publisher = {Springer International Publishing},
address = {Cham},
abstract = {Experimental studies of evolution using microbes have a long tradition, and these studies have increased greatly in number and scope in recent decades. Most such experiments have been short in duration, typically running for weeks or months. A venerable exception, the long-term evolution experiment (LTEE) with \textit{Escherichia coli} has continued for 30 years and 70,000 bacterial generations. The LTEE has become one of the cornerstones of the field of experimental evolution, in general, and the BEACON Center for the Study of Evolution in Action, in particular. Science laboratories and experiments usually have finite lifespans, but we hope that the LTEE can continue far into the future. There are practical issues associated with maintaining such a long-term experiment. One issue, which we address here, is whether key measurements made at one time and place are reproducible, within reasonable limits, at other times and places. This issue comes to the forefront when one considers moving an experiment like the LTEE from one lab to another. To that end, the Barrick lab at The University of Texas at Austin, measured the fitness values of samples from the 12 LTEE populations at 2,000, 10,000, and 50,000 generations and compared the new data to data previously obtained at Michigan State University. On balance, the datasets agree very well. More generally, this finding shows the value of simplicity in experimental design, such as using a chemically defined growth medium and appropriately storing samples from microbiological experiments. Even so, one must be vigilant in checking assumptions and procedures given the potential for uncontrolled factors (e.g., water quality) to affect outcomes. This vigilance is perhaps especially important for a trait like fitness, which integrates all aspects of organismal performance and may therefore be sensitive to any number of subtle environmental influences.},
keywords = {Fitness Trajectories},
pubstate = {published},
tppubtype = {inbook}
}
2015
Lenski R E; Wiser M J; Ribeck N; Blount Z D; Nahum J R; Morris J J; Zaman L; Turner C B; Wade B D; Maddamsetti R; Burmeister A R; Baird E J; Bundy J; Grant N A; Card K J; Rowles M; Weatherspoon K; Papoulis S E; Sullivan R; Clark C; Mulka J S; Hajela N
Sustained fitness gains and variability in fitness trajectories in the long-term evolution experiment with Escherichia coli Journal Article
Proceedings of the Royal Society B: Biological Sciences, 282 (1821), pp. 20152292, 2015, ISSN: 0962-8452.
Abstract | Links | BibTeX | Altmetric | Tags: Fitness Trajectories, Mutation Rates, Parallelism and Divergence
@article{nokey,
title = {Sustained fitness gains and variability in fitness trajectories in the long-term evolution experiment with \textit{Escherichia coli}},
author = {Richard E. Lenski and Michael J. Wiser and Noah Ribeck and Zachary D. Blount and Joshua R. Nahum and J. Jeffrey Morris and Luis Zaman and Caroline B. Turner and Brian D. Wade and Rohan Maddamsetti and Alita R. Burmeister and Elizabeth J. Baird and Jay Bundy and Nkrumah A. Grant and Kyle J. Card and Maia Rowles and Kiyana Weatherspoon and Spiridon E. Papoulis and Rachel Sullivan and Colleen Clark and Joseph S. Mulka and Neerja Hajela},
url = {https://royalsocietypublishing.org/doi/10.1098/rspb.2015.2292},
doi = {10.1098/rspb.2015.2292},
issn = {0962-8452},
year = {2015},
date = {2015-12-22},
urldate = {2015-12-22},
journal = {Proceedings of the Royal Society B: Biological Sciences},
volume = {282},
number = {1821},
pages = {20152292},
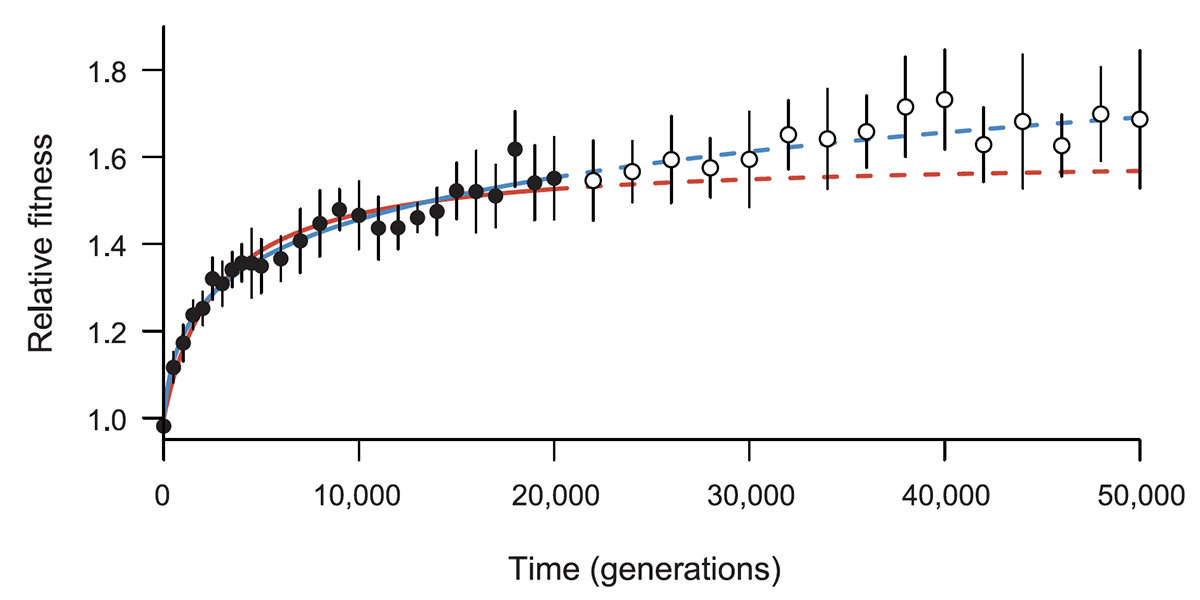
abstract = {Many populations live in environments subject to frequent biotic and abiotic changes. Nonetheless, it is interesting to ask whether an evolving population's mean fitness can increase indefinitely, and potentially without any limit, even in a constant environment. A recent study showed that fitness trajectories of \textit{Escherichia coli} populations over 50 000 generations were better described by a power-law model than by a hyperbolic model. According to the power-law model, the rate of fitness gain declines over time but fitness has no upper limit, whereas the hyperbolic model implies a hard limit. Here, we examine whether the previously estimated power-law model predicts the fitness trajectory for an additional 10 000 generations. To that end, we conducted more than 1100 new competitive fitness assays. Consistent with the previous study, the power-law model fits the new data better than the hyperbolic model. We also analysed the variability in fitness among populations, finding subtle, but significant, heterogeneity in mean fitness. Some, but not all, of this variation reflects differences in mutation rate that evolved over time. Taken together, our results imply that both adaptation and divergence can continue indefinitely—or at least for a long time—even in a constant environment.},
keywords = {Fitness Trajectories, Mutation Rates, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
Maddamsetti R; Lenski R E; Barrick J E
Adaptation, Clonal Interference, and Frequency-Dependent Interactions in a Long-Term Evolution Experiment with Escherichia coli Journal Article
Genetics, 200 (2), pp. 619-631, 2015, ISSN: 1943-2631.
Abstract | Links | BibTeX | Altmetric | Tags: Demography and Ecology, Fitness Trajectories, Genome Evolution
@article{nokey,
title = {Adaptation, Clonal Interference, and Frequency-Dependent Interactions in a Long-Term Evolution Experiment with \emph{Escherichia coli}},
author = {Rohan Maddamsetti and Richard E. Lenski and Jeffrey E. Barrick},
url = {https://academic.oup.com/genetics/article/200/2/619/5936186},
doi = {10.1534/genetics.115.176677},
issn = {1943-2631},
year = {2015},
date = {2015-04-24},
urldate = {2015-04-24},
journal = {Genetics},
volume = {200},
number = {2},
pages = {619-631},
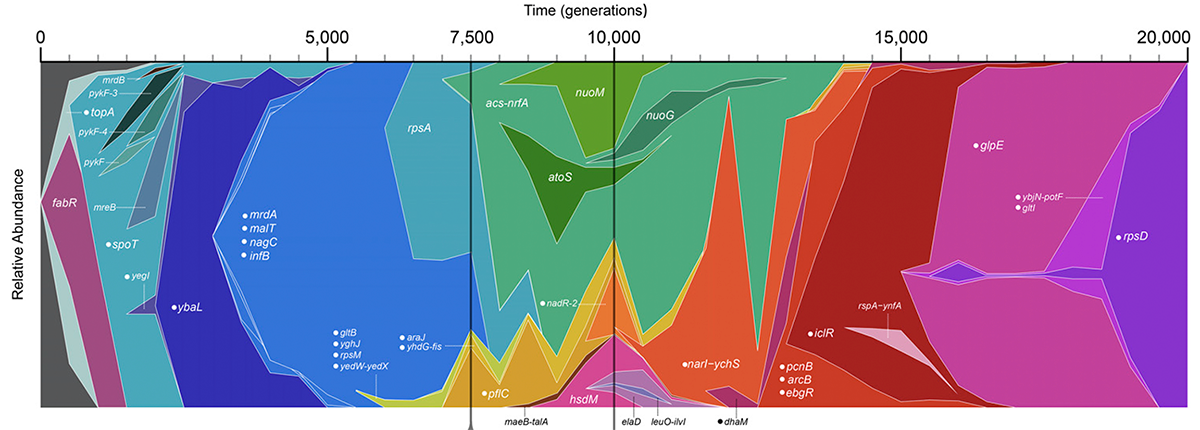
abstract = {Twelve replicate populations of \textit{Escherichia coli} have been evolving in the laboratory for >25 years and 60,000 generations. We analyzed bacteria from whole-population samples frozen every 500 generations through 20,000 generations for one well-studied population, called Ara−1. By tracking 42 known mutations in these samples, we reconstructed the history of this population’s genotypic evolution over this period. The evolutionary dynamics of Ara−1 show strong evidence of selective sweeps as well as clonal interference between competing lineages bearing different beneficial mutations. In some cases, sets of several mutations approached fixation simultaneously, often conveying no information about their order of origination; we present several possible explanations for the existence of these mutational cohorts. Against a backdrop of rapid selective sweeps both earlier and later, two genetically diverged clades coexisted for >6000 generations before one went extinct. In that time, many additional mutations arose in the clade that eventually prevailed. We show that the clades evolved a frequency-dependent interaction, which prevented the immediate competitive exclusion of either clade, but which collapsed as beneficial mutations accumulated in the clade that prevailed. Clonal interference and frequency dependence can occur even in the simplest microbial populations. Furthermore, frequency dependence may generate dynamics that extend the period of coexistence that would otherwise be sustained by clonal interference alone.},
keywords = {Demography and Ecology, Fitness Trajectories, Genome Evolution},
pubstate = {published},
tppubtype = {article}
}
2013
Wiser M J; Ribeck N; Lenski R E
Long-Term Dynamics of Adaptation in Asexual Populations Journal Article
Science, 342 (6164), pp. 1364–1367, 2013, ISSN: 0036-8075.
Abstract | Links | BibTeX | Altmetric | Tags: Fitness Trajectories, Mutation Rates, Theory and Simulations
@article{Wiser2013,
title = {Long-Term Dynamics of Adaptation in Asexual Populations},
author = {Michael J. Wiser and Noah Ribeck and Richard E. Lenski},
url = {https://www.sciencemag.org/lookup/doi/10.1126/science.1243357},
doi = {10.1126/science.1243357},
issn = {0036-8075},
year = {2013},
date = {2013-12-01},
urldate = {2013-12-01},
journal = {Science},
volume = {342},
number = {6164},
pages = {1364--1367},
abstract = {Experimental studies of evolution have increased greatly in number in recent years, stimulated by the growing power of genomic tools. However, organismal fitness remains the ultimate metric for interpreting these experiments, and the dynamics of fitness remain poorly understood over long time scales. Here, we examine fitness trajectories for 12 \textit{Escherichia coli} populations during 50,000 generations. Mean fitness appears to increase without bound, consistent with a power law. We also derive this power-law relation theoretically by incorporating clonal interference and diminishing-returns epistasis into a dynamical model of changes in mean fitness over time.},
keywords = {Fitness Trajectories, Mutation Rates, Theory and Simulations},
pubstate = {published},
tppubtype = {article}
}
Wielgoss S; Barrick J E; Tenaillon O; Wiser M J; Dittmar W J; Cruveiller S; Chane-Woon-Ming B; Médigue C; Lenski R E; Schneider D
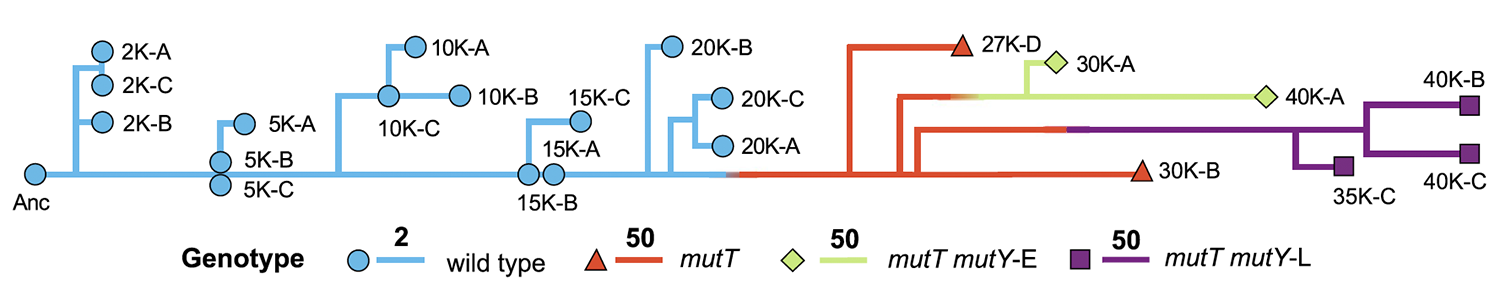
Mutation rate dynamics in a bacterial population reflect tension between adaptation and genetic load Journal Article
Proceedings of the National Academy of Sciences of the United States of America, 110 (1), pp. 222–227, 2013, ISSN: 0027-8424.
Abstract | Links | BibTeX | Altmetric | Tags: Fitness Trajectories, Genome Evolution, Genotypes and Phenotypes, Mutation Rates, Theory and Simulations
@article{Wielgoss2013,
title = {Mutation rate dynamics in a bacterial population reflect tension between adaptation and genetic load},
author = {Sébastien Wielgoss and Jeffrey E. Barrick and Olivier Tenaillon and Michael J. Wiser and W. James Dittmar and Stéphane Cruveiller and Béatrice Chane-Woon-Ming and Claudine Médigue and Richard E. Lenski and Dominique Schneider},
url = {http://www.pnas.org/cgi/doi/10.1073/pnas.1219574110},
doi = {10.1073/pnas.1219574110},
issn = {0027-8424},
year = {2013},
date = {2013-01-01},
urldate = {2013-01-01},
journal = {Proceedings of the National Academy of Sciences of the United States of America},
volume = {110},
number = {1},
pages = {222--227},
abstract = {Mutations are the ultimate source of heritable variation for evolution. Understanding how mutation rates themselves evolve is thus essential for quantitatively understanding many evolutionary processes. According to theory, mutation rates should be minimized for well-adapted populations living in stable environments, whereas hypermutators may evolve if conditions change. However, the long-term fate of hypermutators is unknown. Using a phylogenomic approach, we found that an adapting \textit{Escherichia coli} population that first evolved a \textit{mutT} hypermutator phenotype was later invaded by two independent lineages with \textit{mutY} mutations that reduced genome-wide mutation rates. Applying neutral theory to synonymous substitutions, we dated the emergence of these mutations and inferred that the \textit{mutT} mutation increased the point-mutation rate by ~150-fold, whereas the \textit{mutY} mutations reduced the rate by ~40-60%, with a corresponding decrease in the genetic load. Thus, the long-term fate of the hypermutators was governed by the selective advantage arising from a reduced mutation rate as the potential for further adaptation declined.},
keywords = {Fitness Trajectories, Genome Evolution, Genotypes and Phenotypes, Mutation Rates, Theory and Simulations},
pubstate = {published},
tppubtype = {article}
}
2011
Khan A I; Dinh D M; Schneider D; Lenski R E; Cooper T F
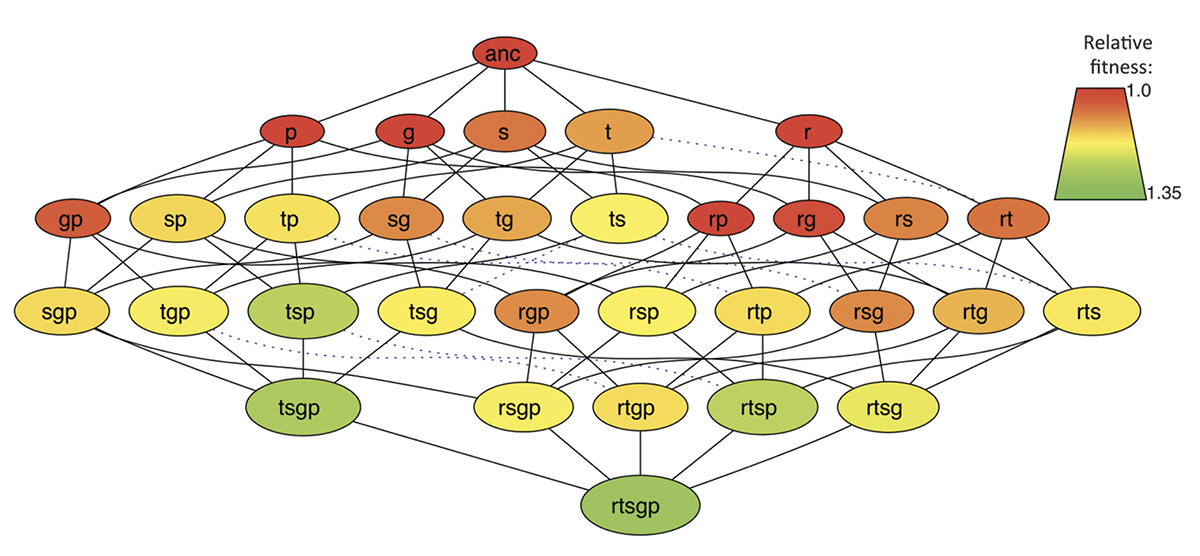
Negative Epistasis Between Beneficial Mutations in an Evolving Bacterial Population Journal Article
Science, 332 (6034), pp. 1193–1196, 2011, ISSN: 0036-8075.
Abstract | Links | BibTeX | Altmetric | Tags: Fitness Trajectories, Genome Evolution, Genotypes and Phenotypes
@article{Khan2011,
title = {Negative Epistasis Between Beneficial Mutations in an Evolving Bacterial Population},
author = {Aisha I. Khan and Duy M. Dinh and Dominique Schneider and Richard E. Lenski and Tim F. Cooper},
url = {https://www.sciencemag.org/lookup/doi/10.1126/science.1203801},
doi = {10.1126/science.1203801},
issn = {0036-8075},
year = {2011},
date = {2011-06-01},
urldate = {2011-06-01},
journal = {Science},
volume = {332},
number = {6034},
pages = {1193--1196},
abstract = {Epistatic interactions between mutations play a prominent role in evolutionary theories. Many studies have found that epistasis is widespread, but they have rarely considered beneficial mutations. We analyzed the effects of epistasis on fitness for the first five mutations to fix in an experimental population of \textit{Escherichia coli}. Epistasis depended on the effects of the combined mutations - the larger the expected benefit, the more negative the epistatic effect. Epistasis thus tended to produce diminishing returns with genotype fitness, although interactions involving one particular mutation had the opposite effect. These data support models in which negative epistasis contributes to declining rates of adaptation over time. Sign epistasis was rare in this genome-wide study, in contrast to its prevalence in an earlier study of mutations in a single gene.},
keywords = {Fitness Trajectories, Genome Evolution, Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}
Woods R J; Barrick J E; Cooper T F; Shrestha U; Kauth M R; Lenski R E
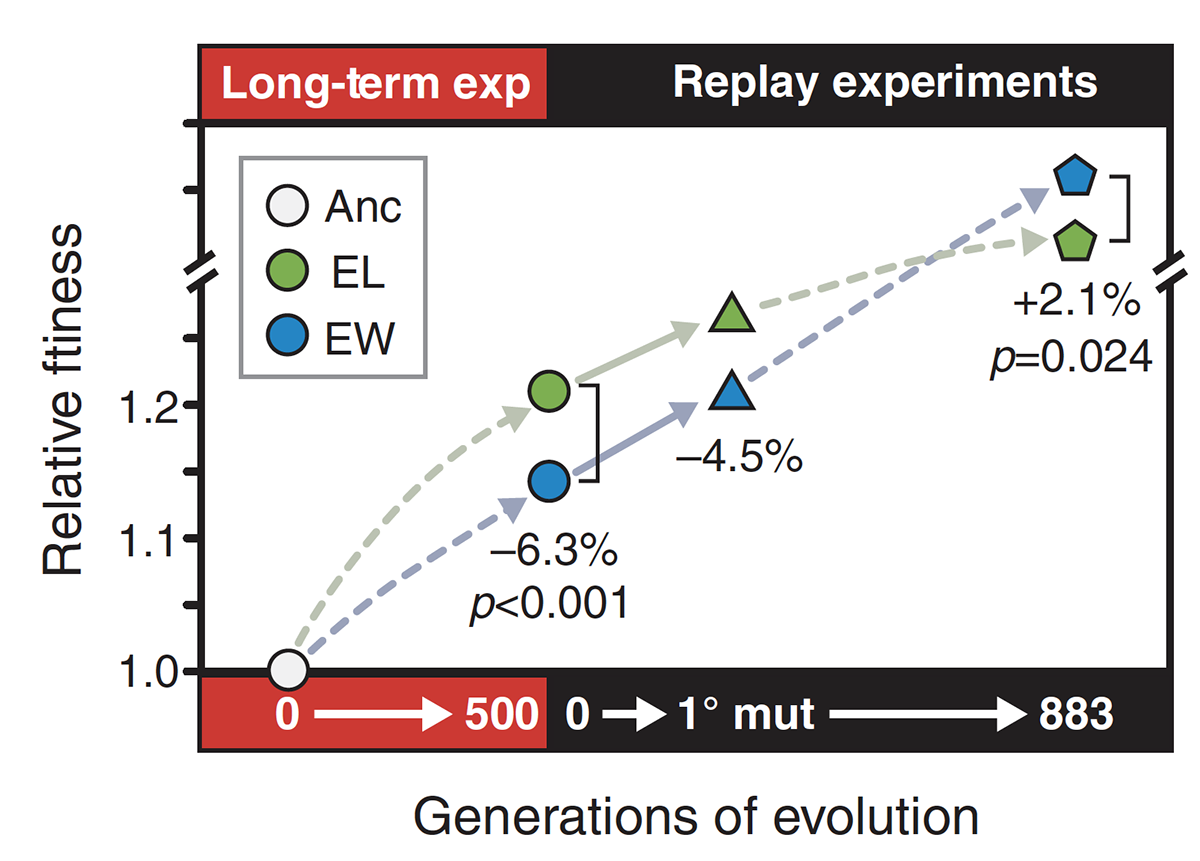
Second-Order Selection for Evolvability in a Large Escherichia coli Population Journal Article
Science, 331 (6023), pp. 1433–1436, 2011, ISSN: 0036-8075.
Abstract | Links | BibTeX | Altmetric | Tags: Fitness Trajectories, Genome Evolution, Historical Contingency
@article{Woods2011,
title = {Second-Order Selection for Evolvability in a Large \emph{Escherichia coli} Population},
author = {Robert J. Woods and Jeffrey E. Barrick and Tim F. Cooper and Utpala Shrestha and Mark R. Kauth and Richard E. Lenski},
url = {https://www.sciencemag.org/lookup/doi/10.1126/science.1198914},
doi = {10.1126/science.1198914},
issn = {0036-8075},
year = {2011},
date = {2011-03-01},
urldate = {2011-03-01},
journal = {Science},
volume = {331},
number = {6023},
pages = {1433--1436},
abstract = {In theory, competition between asexual lineages can lead to second-order selection for greater evolutionary potential. To test this hypothesis, we revived a frozen population of \textit{Escherichia coli} from a long-term evolution experiment and compared the fitness and ultimate fates of four genetically distinct clones. Surprisingly, two clones with beneficial mutations that would eventually take over the population had significantly lower competitive fitness than two clones with mutations that later went extinct. By replaying evolution many times from these clones, we showed that the eventual winners likely prevailed because they had greater potential for further adaptation. Genetic interactions that reduce the benefit of certain regulatory mutations in the eventual losers appear to explain, at least in part, why they were outcompeted.},
keywords = {Fitness Trajectories, Genome Evolution, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
2009
Barrick J E; Yu D S; Yoon S H; Jeong H; Oh T K; Schneider D; Lenski R E; Kim J F
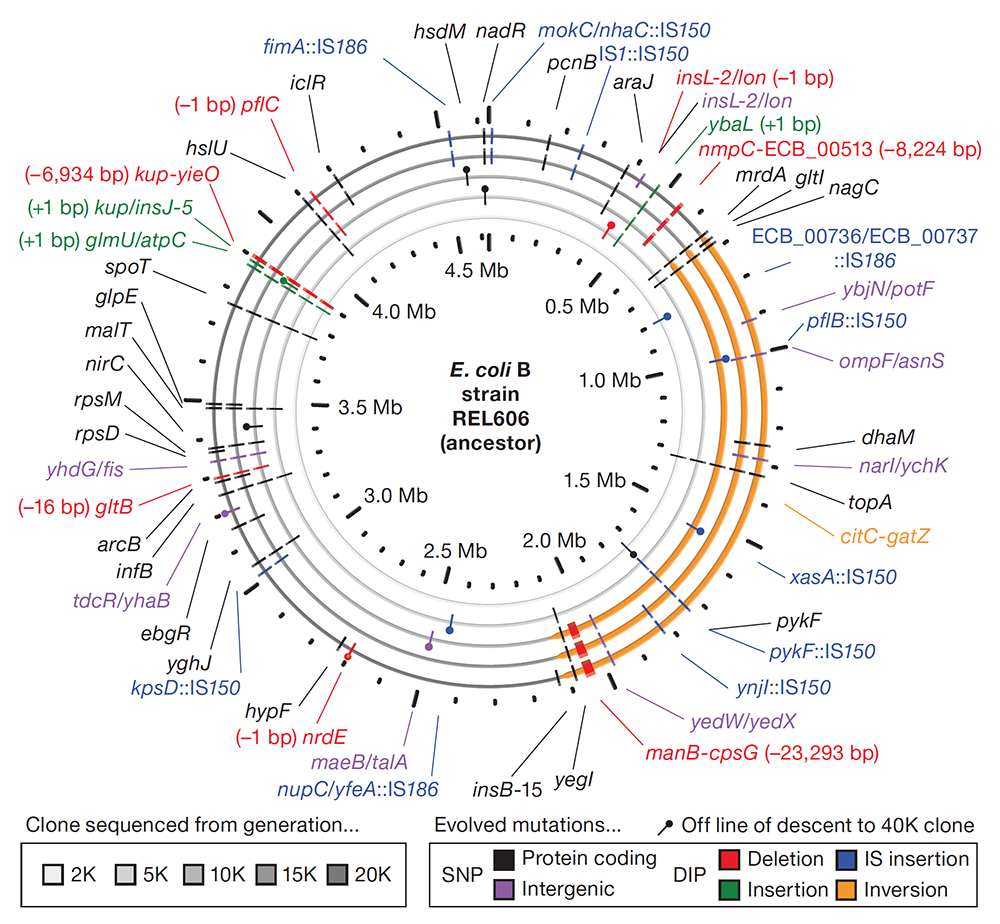
Genome evolution and adaptation in a long-term experiment with Escherichia coli. Journal Article
Nature, 461 (7268), pp. 1243–7, 2009, ISSN: 1476-4687.
Abstract | Links | BibTeX | Altmetric | Tags: Fitness Trajectories, Genome Evolution, Genotypes and Phenotypes, Mutation Rates, Parallelism and Divergence
@article{Barrick2009,
title = {Genome evolution and adaptation in a long-term experiment with \textit{Escherichia coli}.},
author = {Jeffrey E. Barrick and Dong Su Yu and Sung Ho Yoon and Haeyoung Jeong and Tae Kwang Oh and Dominique Schneider and Richard E. Lenski and Jihyun F. Kim},
url = {http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=19838166&retmode=ref&cmd=prlinks papers2://publication/doi/10.1038/nature08480 http://www.ncbi.nlm.nih.gov/pubmed/19838166},
doi = {10.1038/nature08480},
issn = {1476-4687},
year = {2009},
date = {2009-10-01},
urldate = {2009-10-01},
journal = {Nature},
volume = {461},
number = {7268},
pages = {1243--7},
publisher = {Nature Publishing Group},
address = {Department of Microbiology and Molecular Genetics, Michigan State University, East Lansing, Michigan 48824, USA.},
abstract = {The relationship between rates of genomic evolution and organismal adaptation remains uncertain, despite considerable interest. The feasibility of obtaining genome sequences from experimentally evolving populations offers the opportunity to investigate this relationship with new precision. Here we sequence genomes sampled through 40,000 generations from a laboratory population of \textit{Escherichia coli}. Although adaptation decelerated sharply, genomic evolution was nearly constant for 20,000 generations. Such clock-like regularity is usually viewed as the signature of neutral evolution, but several lines of evidence indicate that almost all of these mutations were beneficial. This same population later evolved an elevated mutation rate and accumulated hundreds of additional mutations dominated by a neutral signature. Thus, the coupling between genomic and adaptive evolution is complex and can be counterintuitive even in a constant environment. In particular, beneficial substitutions were surprisingly uniform over time, whereas neutral substitutions were highly variable.},
keywords = {Fitness Trajectories, Genome Evolution, Genotypes and Phenotypes, Mutation Rates, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
2007
Cooper T F
Recombination speeds adaptation by reducing competition between beneficial mutations in populations of Escherichia coli Journal Article
PLoS Biology, 5 (9), pp. e225, 2007.
Abstract | Links | BibTeX | Altmetric | Tags: Descendant Experiments, Fitness Trajectories
@article{Cooper2007,
title = {Recombination speeds adaptation by reducing competition between beneficial mutations in populations of \textit{Escherichia coli}},
author = {Tim F. Cooper},
doi = {10.1371/journal.pbio.0050225},
year = {2007},
date = {2007-01-01},
urldate = {2007-01-01},
journal = {PLoS Biology},
volume = {5},
number = {9},
pages = {e225},
abstract = {Identification of the selective forces contributing to the origin and maintenance of sex is a fundamental problem in biology. The Fisher-Muller model proposes that sex is advantageous because it allows beneficial mutations that arise in different lineages to recombine, thereby reducing clonal interference and speeding adaptation. I used the F plasmid to mediate recombination in the bacterium \textit{Escherichia coli} and measured its effect on adaptation at high and low mutation rates. Recombination increased the rate of adaptation approximately 3-fold more in the high mutation rate treatment, where beneficial mutations had to compete for fixation. Sequencing of candidate loci revealed the presence of a beneficial mutation in six high mutation rate lines. In the absence of recombination, this mutation took longer to fix and, over the course of its substitution, conferred a reduced competitive advantage, indicating interference between competing beneficial mutations. Together, these results provide experimental support for the Fisher-Muller model and demonstrate that plasmid-mediated gene transfer can accelerate bacterial adaptation.},
keywords = {Descendant Experiments, Fitness Trajectories},
pubstate = {published},
tppubtype = {article}
}
2002
de Visser J A G M; Lenski R E
Long-term experimental evolution in Escherichia coli. XI. Rejection of non-transitive interactions as cause of declining rate of adaptation. Journal Article
BMC evolutionary biology, 2 (May 2014), pp. 19, 2002, ISSN: 1471-2148.
Abstract | Links | BibTeX | Altmetric | Tags: Fitness Trajectories
@article{DeVisser2002,
title = {Long-term experimental evolution in \textit{Escherichia coli}. XI. Rejection of non-transitive interactions as cause of declining rate of adaptation.},
author = {J. Arjan G. M. {de Visser} and Richard E. Lenski},
url = {http://www.ncbi.nlm.nih.gov/pubmed/12443537
http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=PMC134600},
doi = {10.1186/1471-2148-2-19},
issn = {1471-2148},
year = {2002},
date = {2002-10-01},
urldate = {2002-10-01},
journal = {BMC evolutionary biology},
volume = {2},
number = {May 2014},
pages = {19},
abstract = {Background
Experimental populations of \textit{Escherichia coli} have evolved for 20,000 generations in a uniform environment. Their rate of improvement, as measured in competitions with the ancestor in that environment, has declined substantially over this period. This deceleration has been interpreted as the bacteria approaching a peak or plateau in a fitness landscape. Alternatively, this deceleration might be caused by non-transitive competitive interactions, in particular such that the measured advantage of later genotypes relative to earlier ones would be greater if they competed directly.
Results
To distinguish these two hypotheses, we performed a large set of competitions using one of the evolved lines. Twenty-one samples obtained at 1,000-generation intervals each competed against five genetically marked clones isolated at 5,000-generation intervals, with three-fold replication. The pattern of relative fitness values for these 315 pairwise competitions was compared with expectations under transitive and non-transitive models, the latter structured to produce the observed deceleration in fitness relative to the ancestor. In general, the relative fitness of later and earlier generations measured by direct competition agrees well with the fitness inferred from separately competing each against the ancestor. These data thus support the transitive model.
Conclusion
Non-transitive competitive interactions were not a major feature of evolution in this population. Instead, the pronounced deceleration in its rate of fitness improvement indicates that the population early on incorporated most of those mutations that provided the greatest gains, and subsequently relied on beneficial mutations that were fewer in number, smaller in effect, or both.},
keywords = {Fitness Trajectories},
pubstate = {published},
tppubtype = {article}
}
Experimental populations of Escherichia coli have evolved for 20,000 generations in a uniform environment. Their rate of improvement, as measured in competitions with the ancestor in that environment, has declined substantially over this period. This deceleration has been interpreted as the bacteria approaching a peak or plateau in a fitness landscape. Alternatively, this deceleration might be caused by non-transitive competitive interactions, in particular such that the measured advantage of later genotypes relative to earlier ones would be greater if they competed directly.
Results
To distinguish these two hypotheses, we performed a large set of competitions using one of the evolved lines. Twenty-one samples obtained at 1,000-generation intervals each competed against five genetically marked clones isolated at 5,000-generation intervals, with three-fold replication. The pattern of relative fitness values for these 315 pairwise competitions was compared with expectations under transitive and non-transitive models, the latter structured to produce the observed deceleration in fitness relative to the ancestor. In general, the relative fitness of later and earlier generations measured by direct competition agrees well with the fitness inferred from separately competing each against the ancestor. These data thus support the transitive model.
Conclusion
Non-transitive competitive interactions were not a major feature of evolution in this population. Instead, the pronounced deceleration in its rate of fitness improvement indicates that the population early on incorporated most of those mutations that provided the greatest gains, and subsequently relied on beneficial mutations that were fewer in number, smaller in effect, or both.
2000
Cooper V S; Lenski R E
The population genetics of ecological specialization in evolving Escherichia coli populations Journal Article
Nature, 407 (6805), pp. 736–739, 2000, ISSN: 0028-0836.
Abstract | Links | BibTeX | Altmetric | Tags: Correlated Responses, Fitness Trajectories, Mutation Rates
@article{Schneider2000b,
title = {The population genetics of ecological specialization in evolving \textit{Escherichia coli} populations},
author = {Vaughn S. Cooper and Richard E. Lenski},
url = {http://www.nature.com/articles/35037572},
doi = {10.1038/35037572},
issn = {0028-0836},
year = {2000},
date = {2000-10-01},
urldate = {2000-10-01},
journal = {Nature},
volume = {407},
number = {6805},
pages = {736--739},
abstract = {When organisms adapt genetically to one environment, they may lose fitness in other environments. Two distinct population genetic processes can produce ecological specialization—mutation accumulation and antagonistic pleiotropy. In mutation accumulation, mutations become fixed by genetic drift in genes that are not maintained by selection; adaptation to one environment and loss of adaptation to another are caused by different mutations. Antagonistic pleiotropy arises from trade-offs, such that the same mutations that are beneficial in one environment are detrimental in another. In general, it is difficult to distinguish between these processes. We analysed the decay of unused catabolic functions in 12 lines of \textit{Escherichia coli} propagated on glucose for 20,000 generations. During that time, several lines evolved high mutation rates. If mutation accumulation is important, their unused functions should decay more than the other lines, but no significant difference was observed. Moreover, most catabolic losses occurred early in the experiment when beneficial mutations were being rapidly fixed, a pattern predicted by antagonistic pleiotropy. Thus, antagonistic pleiotropy appears more important than mutation accumulation for the decay of unused catabolic functions in these populations.},
keywords = {Correlated Responses, Fitness Trajectories, Mutation Rates},
pubstate = {published},
tppubtype = {article}
}