2024
Ascensao J A; Denk J; Lok K; Yu Q; Wetmore K M; Hallatschek O
Rediversification following ecotype isolation reveals hidden adaptive potential Journal Article
Current Biology, 34 (4), pp. 855–867, 2024, ISSN: 0960-9822.
Abstract | Links | BibTeX | Altmetric | Tags: Demography and Ecology, Genotypes and Phenotypes, Historical Contingency
@article{Ascensao2024,
title = {Rediversification following ecotype isolation reveals hidden adaptive potential},
author = {Joao A. Ascensao and Jonas Denk and Kristen Lok and QinQin Yu and Kelly M. Wetmore and Oskar Hallatschek},
url = {https://www.sciencedirect.com/science/article/pii/S0960982224000290},
doi = {10.1016/j.cub.2024.01.029},
issn = {0960-9822},
year = {2024},
date = {2024-02-01},
urldate = {2024-02-01},
journal = {Current Biology},
volume = {34},
number = {4},
pages = {855--867},
abstract = {Microbial communities play a critical role in ecological processes, and their diversity is key to their functioning. However, little is known about whether communities can regenerate ecological diversity following ecotype removal or extinction and how the rediversified communities would compare to the original ones. Here, we show that simple two-ecotype communities from the \textit{E. coli} long-term evolution experiment (LTEE) consistently rediversified into two ecotypes following the isolation of one of the ecotypes, coexisting via negative frequency-dependent selection. Communities separated by more than 30,000 generations of evolutionary time rediversify in similar ways. The rediversified ecotype appears to share a number of growth traits with the ecotype it replaces. However, the rediversified community is also different from the original community in ways relevant to the mechanism of ecotype coexistence—for example, in stationary phase response and survival. We found substantial variation in the transcriptional states between the two original ecotypes, whereas the differences within the rediversified community were comparatively smaller, although the rediversified community showed unique patterns of differential expression. Our results suggest that evolution may leave room for alternative diversification processes even in a maximally reduced community of only two strains. We hypothesize that the presence of alternative evolutionary pathways may be even more pronounced in communities of many species where there are even more potential niches, highlighting an important role for perturbations, such as species removal, in evolving ecological communities.},
keywords = {Demography and Ecology, Genotypes and Phenotypes, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
2023
Turner C B; Blount Z D; Mitchell D H; Lenski R E
Evolution of a cross-feeding interaction following a key innovation in a long-term evolution experiment with Escherichia coli Journal Article
Microbiology (Reading), 169 (8), 2023.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Demography and Ecology, Historical Contingency
@article{Turner2023,
title = { Evolution of a cross-feeding interaction following a key innovation in a long-term evolution experiment with \textit{Escherichia coli} },
author = {Caroline B. Turner and Zachary D. Blount and Daniel H. Mitchell and Richard E. Lenski},
doi = {10.1099/mic.0.001390},
year = {2023},
date = {2023-08-01},
urldate = {2023-08-01},
journal = { Microbiology (Reading)},
volume = {169},
number = {8},
abstract = {The evolution of a novel trait can profoundly change an organism's effects on its environment, which can in turn affect the further evolution of that organism and any coexisting organisms. We examine these effects and feedbacks following the evolution of a novel function in the Long-Term Evolution Experiment (LTEE) with \textit{Escherichia coli}. A characteristic feature of \textit{E. coli} is its inability to grow aerobically on citrate (Cit^{−}). Nonetheless, a Cit^{+} variant with this capacity evolved in one LTEE population after 31 000 generations. The Cit^{+} clade then coexisted stably with another clade that retained the ancestral Cit^{−} phenotype. This coexistence was shaped by the evolution of a cross-feeding relationship based on C4-dicarboxylic acids, particularly succinate, fumarate, and malate, that the Cit^{+} variants release into the medium. Both the Cit^{−} and Cit^{+} cells evolved to grow on these excreted resources. The evolution of aerobic growth on citrate thus led to a transition from an ecosystem based on a single limiting resource, glucose, to one with at least five resources that were either shared or partitioned between the two coexisting clades. Our findings show that evolutionary novelties can change environmental conditions in ways that facilitate diversity by altering ecosystem structure and the evolutionary trajectories of coexisting lineages. },
keywords = {Citrate Evolution, Demography and Ecology, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
Mukherjee A; Ealy J; Huang Y; Benites N C; Polk M; Basan M
Coexisting ecotypes in long-term evolution emerged from interacting trade-offs Journal Article
Nature Communications, 14 (1), pp. 3805, 2023, ISSN: 2041-1723.
Abstract | Links | BibTeX | Altmetric | Tags: Demography and Ecology, Theory and Simulations
@article{Mukherjee2023,
title = {Coexisting ecotypes in long-term evolution emerged from interacting trade-offs},
author = {Avik Mukherjee and Jade Ealy and Yanqing Huang and Nina Catherine Benites and Mark Polk and Markus Basan},
url = {https://www.nature.com/articles/s41467-023-39471-9},
doi = {10.1038/s41467-023-39471-9},
issn = {2041-1723},
year = {2023},
date = {2023-06-26},
journal = {Nature Communications},
volume = {14},
number = {1},
pages = {3805},
abstract = {Evolution of complex communities of coexisting microbes remains poorly understood. The long-term evolution experiment on \textit{Escherichia coli} (LTEE) revealed the spontaneous emergence of stable coexistence of multiple ecotypes, which persisted for more than 14,000 generations of continuous evolution. Here, using a combination of experiments and computer simulations, we show that the emergence and persistence of this phenomenon can be explained by the combination of two interacting trade-offs, rooted in biochemical constraints: First, faster growth is enabled by higher fermentation and obligate acetate excretion. Second, faster growth results in longer lag times when utilizing acetate after glucose is depleted. This combination creates an ecological niche for a slower-growing ecotype, specialized in switching to acetate. These findings demonstrate that trade-offs can give rise to surprisingly complex communities with evolutionarily stable coexistence of multiple variants in even the simplest environments.},
keywords = {Demography and Ecology, Theory and Simulations},
pubstate = {published},
tppubtype = {article}
}
Jagdish T
The dynamics of fitness and pleiotropy in a long-term evolution experiment with Escherichia coli PhD Thesis
2023.
Abstract | Links | BibTeX | Tags: Correlated Responses, Demography and Ecology, Fitness Trajectories, Methods and Miscellaneous
@phdthesis{nokey,
title = {The dynamics of fitness and pleiotropy in a long-term evolution experiment with \textit{Escherichia coli}},
author = {Tanush Jagdish},
url = {https://dash.harvard.edu/handle/1/37375541},
year = {2023},
date = {2023-05-08},
urldate = {2023-05-08},
abstract = {Life on earth is shaped by a delicate balance between chance and necessity. A combination of natural selection and genetic drift has moulded random genetic variations over the course of billions of years to generate the complex and interconnected biosphere we see today. These evolutionary dynamics are ultimately the product of numerous genetic mechanisms operating in genomes with highly complex architectures. Unravelling the rules of genetic mechanisms responsible for evolutionary innovation is critical to developing a comprehensive and predictive evolutionary theory, but has evaded direct experimentation since the scale of resources and technology needed to study the patterns of genetic evolution has only recently become achievable. In this dissertation, I explore the use of microbial experimental evolution as a powerful tool for probing evolutionary questions, particularly by leveraging DNA-barcoding and high-throughput DNA sequencing. In chapter 1, I provide an overview of the field of microbial experimental evolution, dividing its history into two eras marked by qualitatively different methodological advancements. I make the case that we now stand at the dawn of a third era, where advances in genome-engineering coupled with low-cost, high-throughput DNA sequencing will allow experiments to finally probe evolution on a statistical scale. I then present two research studies that take advantage of microbial experimental evolution to investigate distinct genetic mechanisms key to the evolutionary process. In Chapter 2, I explore whether fitness can continue increasing in a population that has already adapted to its environment for over 30,000 generations, and whether fixations of beneficial mutations from population-wide DNA sequencing can predict jumps in fitness. My findings reveal that fitness continues to monotonically increase in step with the fixation of beneficial mutations, even though the rate of fixation has dramatically slowed down, highlighting the potential for ongoing adaptation even under constant environmental conditions. In Chapter 3, I develop a novel conjugation-based DNA-barcoding method for the Long-Term Evolution Experiment (LTEE) with \textit{Escherichia coli}, allowing me to examine the pleiotropic consequences of adaptation to glucose over 50,000 generations in 15 novel resource environments. My observations reveal broad patterns of both convergent and divergent evolution that correspond with mutations in key metabolic genes in clonal sequencing datasets, shedding light on the nature of pleiotropy and its evolution over extended timescales. Using microbial model systems in an evolutionary context has the unique advantage of being relevant both to fundamental evolutionary biology and human health. Since genetic drift and rare mutational events both play an outsized role in determining the evolutionary trajectories of populations, evolutionary questions in the modern age will increasingly be faced with issues of scale. Microbial experimental evolution offers both scale and tractability to solve this problem. Uniquely, this does not sacrifice on human relevance. Building a coarse-grained and comprehensive evolutionary theory is more significant to society today than ever before as the importance of clonal evolution in cancer, gut microbiomes and even pandemics becomes more clearly understood.},
keywords = {Correlated Responses, Demography and Ecology, Fitness Trajectories, Methods and Miscellaneous},
pubstate = {published},
tppubtype = {phdthesis}
}
Ascensao J A; Wetmore K M; Good B H; Arkin A P; Hallatschek O
Quantifying the adaptive potential of a nascent bacterial community Journal Article
Nature Communications, 14 (1), pp. 248, 2023.
Abstract | Links | BibTeX | Altmetric | Tags: Demography and Ecology, Descendant Experiments, Genome Evolution, Genotypes and Phenotypes
@article{ascensao2023,
title = {Quantifying the adaptive potential of a nascent bacterial community},
author = {Joao A. Ascensao and Kelly M. Wetmore and Benjamin H. Good and Adam P. Arkin and Oskar Hallatschek},
url = {https://www.nature.com/articles/s41467-022-35677-5},
doi = {10.1038/s41467-022-35677-5},
year = {2023},
date = {2023-01-16},
urldate = {2022-01-01},
journal = {Nature Communications},
volume = {14},
number = {1},
pages = {248},
publisher = {Cold Spring Harbor Laboratory},
abstract = {The fitness effects of all possible mutations available to an organism largely shape the dynamics of evolutionary adaptation. Yet, whether and how this adaptive landscape changes over evolutionary times, especially upon ecological diversification and changes in community composition, remains poorly understood. We sought to fill this gap by analyzing a stable community of two closely related ecotypes (“L” and “S”) shortly after they emerged within the \textit{E. coli} Long-Term Evolution Experiment (LTEE). We engineered genome-wide barcoded transposon libraries to measure the invasion fitness effects of all possible gene knockouts in the coexisting strains as well as their ancestor, for many different, ecologically relevant conditions. We find consistent statistical patterns of fitness effect variation across both genetic background and community composition, despite the idiosyncratic behavior of individual knockouts. Additionally, fitness effects are correlated with evolutionary outcomes for a number of conditions, possibly revealing shifting patterns of adaptation. Together, our results reveal how ecological and epistatic effects combine to shape the adaptive landscape in a nascent ecological community.},
keywords = {Demography and Ecology, Descendant Experiments, Genome Evolution, Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}
2022
Marshall D J; Malerba M; Lines T; Sezmis A L; Hasan C M; Lenski R E; McDonald M J
Long-term experimental evolution decouples size and production costs in Escherichia coli Journal Article
Proceedings of the National Academy of Sciences of the United States of America, 119 (21), pp. e2200713119, 2022.
Abstract | Links | BibTeX | Altmetric | Tags: Cell Morphology, Demography and Ecology
@article{marshall2021,
title = {Long-term experimental evolution decouples size and production costs in \emph{Escherichia coli}},
author = {Dustin J. Marshall and Martino Malerba and Thomas Lines and Aysha L. Sezmis and Chowdhury M. Hasan and Richard E. Lenski and Michael J. McDonald},
editor = {Ruth Shaw},
url = {https://pnas.org/doi/full/10.1073/pnas.2200713119},
doi = {10.1073/pnas.2200713119},
year = {2022},
date = {2022-05-20},
urldate = {2022-05-20},
journal = {Proceedings of the National Academy of Sciences of the United States of America},
volume = {119},
number = {21},
pages = {e2200713119},
abstract = {Body size covaries with population dynamics across life’s domains. Metabolism may impose fundamental constraints on the coevolution of size and demography, but experimental tests of the causal links remain elusive. We leverage a 60,000-generation experiment in which \textit{Escherichia coli} populations evolved larger cells to examine intraspecific metabolic scaling and correlations with demographic parameters. Over the course of their evolution, the cells have roughly doubled in size relative to their ancestors. These larger cells have metabolic rates that are absolutely higher, but relative to their size, they are lower. Metabolic theory successfully predicted the relations between size, metabolism, and maximum population density, including support for Damuth’s law of energy equivalence, such that populations of larger cells achieved lower maximum densities but higher maximum biomasses than populations of smaller cells. The scaling of metabolism with cell size thus predicted the scaling of size with maximum population density. In stark contrast to standard theory, however, populations of larger cells grew faster than those of smaller cells, contradicting the fundamental and intuitive assumption that the costs of building new individuals should scale directly with their size. The finding that the costs of production can be decoupled from size necessitates a reevaluation of the evolutionary drivers and ecological consequences of biological size more generally.},
howpublished = {bioRxiv},
keywords = {Cell Morphology, Demography and Ecology},
pubstate = {published},
tppubtype = {article}
}
2021
van Raay K; Stolyar S; Sevigny J; Draghi J; Lenski R E; Marx C J; Kerr B; Zaman L
Evolution with private resources reverses some changes from long-term evolution with public resources Unpublished
bioRxiv, 2021.
Abstract | Links | BibTeX | Altmetric | Tags: Cell Morphology, Correlated Responses, Demography and Ecology, Descendant Experiments, Genotypes and Phenotypes, Historical Contingency, Methods and Miscellaneous
@unpublished{Raay2021,
title = {Evolution with private resources reverses some changes from long-term evolution with public resources},
author = {Katrina {van Raay} and Sergey Stolyar and Jordana Sevigny and Jeremy Draghi and Richard E. Lenski and Christopher J. Marx and Benjamin Kerr and Luis Zaman},
url = {https://www.biorxiv.org/content/10.1101/2021.07.11.451942v1},
doi = {https://doi.org/10.1101/2021.07.11.451942},
year = {2021},
date = {2021-07-12},
urldate = {2021-07-12},
journal = {bioRxiv},
pages = {2021.07.11.451942},
abstract = {A population under selection to improve one trait may evolve a sub-optimal state for another trait due to tradeoffs and other evolutionary constraints. How this evolution affects the capacity of a population to adapt when conditions change to favor the second trait is an open question. We investigated this question using isolates from a lineage spanning 60,000 generations of the Long-Term Evolution Experiment (LTEE) with \textit{Escherichia coli}, where cells have access to a shared pool of resources, and have evolved increased competitive ability and a concomitant reduction in numerical yield. Using media-in oil emulsions we shifted the focus of selection to numerical yield, where cells grew in isolated patches with private resources. We found that the time spent evolving under shared resources did not affect the ability to re-evolve toward higher numerical yield. The evolution of numerical yield commonly occurred through mutations in the phosphoenolpyruvate phosphotransferase system. These mutants exhibit slower uptake of glucose, making them poorer competitors for public resources, and produce smaller cells that release less carbon as overflow metabolites. Our results demonstrate that mutations that were not part of adaptation under one selective regime may enable access to ancestral phenotypes when selection changes to favor evolutionary reversion. },
howpublished = {bioRxiv},
keywords = {Cell Morphology, Correlated Responses, Demography and Ecology, Descendant Experiments, Genotypes and Phenotypes, Historical Contingency, Methods and Miscellaneous},
pubstate = {published},
tppubtype = {unpublished}
}
2020
Atolia E; Cesar S; Arjes H A; Rajendram M; Shi H; Knapp B D; Khare S; Aranda-Díaz A; Lenski R E; Huang K C
Environmental and Physiological Factors Affecting High-Throughput Measurements of Bacterial Growth Journal Article
mBio, 11 (5), pp. 1–19, 2020, ISSN: 2161-2129.
Abstract | Links | BibTeX | Altmetric | Tags: Demography and Ecology
@article{Atolia2020,
title = {Environmental and Physiological Factors Affecting High-Throughput Measurements of Bacterial Growth},
author = {Esha Atolia and Spencer Cesar and Heidi A Arjes and Manohary Rajendram and Handuo Shi and Benjamin D Knapp and Somya Khare and Andrés Aranda-Díaz and Richard E. Lenski and Kerwyn Casey Huang},
editor = {Kelly T. Hughes},
url = {https://journals.asm.org/doi/10.1128/mBio.01378-20},
doi = {10.1128/mBio.01378-20},
issn = {2161-2129},
year = {2020},
date = {2020-10-01},
urldate = {2020-10-01},
journal = {mBio},
volume = {11},
number = {5},
pages = {1--19},
abstract = {How starved bacteria adapt and multiply under replete nutrient conditions is intimately linked to their history of previous growth, their physiological state, and the surrounding environment. While automated equipment has enabled high-throughput growth measurements, data interpretation and knowledge gaps regarding the determinants of growth kinetics complicate comparisons between strains. Here, we present a framework for growth measurements that improves accuracy and attenuates the effects of growth history. We determined that background absorbance quantification and multiple passaging cycles allow for accurate growth rate measurements even in carbon-poor media, which we used to reveal growth-rate increases during long-term laboratory evolution of \textit{Escherichia coli}. Using mathematical modeling, we showed that maximum growth rate depends on initial cell density. Finally, we demonstrated that growth of Bacillus subtilis with glycerol inhibits the future growth of most of the population, due to lipoteichoic acid synthesis. These studies highlight the challenges of accurate quantification of bacterial growth behaviors.},
keywords = {Demography and Ecology},
pubstate = {published},
tppubtype = {article}
}
Blount Z D; Maddamsetti R; Grant N A; Ahmed S T; Jagdish T; Baxter J A; Sommerfeld B A; Tillman A; Moore J; Slonczewski J L; Barrick J E; Lenski R E
Genomic and phenotypic evolution of Escherichia coli in a novel citrate-only resource environment Journal Article
eLife, 9 , pp. 1–64, 2020, ISSN: 2050-084X.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Demography and Ecology, Descendant Experiments, Genotypes and Phenotypes
@article{Blount2020,
title = {Genomic and phenotypic evolution of \textit{Escherichia coli} in a novel citrate-only resource environment},
author = {Zachary D. Blount and Rohan Maddamsetti and Nkrumah A. Grant and Sumaya T. Ahmed and Tanush Jagdish and Jessica A. Baxter and Brooke A. Sommerfeld and Alice Tillman and Jeremy Moore and Joan L. Slonczewski and Jeffrey E. Barrick and Richard E. Lenski},
url = {https://elifesciences.org/articles/55414},
doi = {10.7554/eLife.55414},
issn = {2050-084X},
year = {2020},
date = {2020-05-01},
urldate = {2020-05-01},
journal = {eLife},
volume = {9},
pages = {1--64},
abstract = {Evolutionary innovations allow populations to colonize new ecological niches. We previously reported that aerobic growth on citrate (Cit^{+}) evolved in an \textit{Escherichia coli} population during adaptation to a minimal glucose medium containing citrate (DM25). Cit^{+} variants can also grow in citrate-only medium (DM0), a novel environment for \textit{E. coli}. To study adaptation to this niche, we founded two sets of Cit^{+} populations and evolved them for 2500 generations in DM0 or DM25. The evolved lineages acquired numerous parallel mutations, many mediated by transposable elements. Several also evolved amplifications of regions containing the \textit{maeA} gene. Unexpectedly, some evolved populations and clones show apparent declines in fitness. We also found evidence of substantial cell death in Cit^{+} clones. Our results thus demonstrate rapid trait refinement and adaptation to the new citrate niche, while also suggesting a recalcitrant mismatch between \textit{E. coli} physiology and growth on citrate.},
keywords = {Citrate Evolution, Demography and Ecology, Descendant Experiments, Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}
2018
Bajić D; Vila J C C; Blount Z D; Sánchez A
On the deformability of an empirical fitness landscape by microbial evolution Journal Article
Proceedings of the National Academy of Sciences of the United States of America, 115 (44), pp. 11286-11291, 2018, ISSN: 0027-8424.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Demography and Ecology, Theory and Simulations
@article{nokey,
title = {On the deformability of an empirical fitness landscape by microbial evolution},
author = {Djordje Bajić and Jean C. C. Vila and Zachary D. Blount and Alvaro Sánchez},
url = {http://www.pnas.org/lookup/doi/10.1073/pnas.1808485115},
doi = {10.1073/pnas.1808485115},
issn = {0027-8424},
year = {2018},
date = {2018-10-30},
urldate = {2018-10-30},
journal = {Proceedings of the National Academy of Sciences of the United States of America},
volume = {115},
number = {44},
pages = {11286-11291},
abstract = {A fitness landscape is a map between the genotype and its reproductive success in a given environment. The topography of fitness landscapes largely governs adaptive dynamics, constraining evolutionary trajectories and the predictability of evolution. Theory suggests that this topography can be deformed by mutations that produce substantial changes to the environment. Despite its importance, the deformability of fitness landscapes has not been systematically studied beyond abstract models, and little is known about its reach and consequences in empirical systems. Here we have systematically characterized the deformability of the genome-wide metabolic fitness landscape of the bacterium \textit{Escherichia coli}. Deformability is quantified by the noncommutativity of epistatic interactions, which we experimentally demonstrate in mutant strains on the path to an evolutionary innovation. Our analysis shows that the deformation of fitness landscapes by metabolic mutations rarely affects evolutionary trajectories in the short range. However, mutations with large environmental effects produce long-range landscape deformations in distant regions of the genotype space that affect the fitness of later descendants. Our results therefore suggest that, even in situations in which mutations have strong environmental effects, fitness landscapes may retain their power to forecast evolution over small mutational distances despite the potential attenuation of that power over longer evolutionary trajectories. Our methods and results provide an avenue for integrating adaptive and eco-evolutionary dynamics with complex genetics and genomics. },
keywords = {Citrate Evolution, Demography and Ecology, Theory and Simulations},
pubstate = {published},
tppubtype = {article}
}
Leon D; D'Alton S; Quandt E M; Barrick J E
Innovation in an E. coli evolution experiment is contingent on maintaining adaptive potential until competition subsides Journal Article
PLOS Genetics, 14 (4), pp. e1007348, 2018, ISSN: 1553-7404.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Demography and Ecology, Historical Contingency
@article{nokey,
title = {Innovation in an \textit{E. coli} evolution experiment is contingent on maintaining adaptive potential until competition subsides},
author = {Dacia Leon and Simon D'Alton and Erik M. Quandt and Jeffrey E. Barrick},
url = {https://dx.plos.org/10.1371/journal.pgen.1007348},
doi = {10.1371/journal.pgen.1007348},
issn = {1553-7404},
year = {2018},
date = {2018-04-12},
urldate = {2018-04-12},
journal = {PLOS Genetics},
volume = {14},
number = {4},
pages = {e1007348},
abstract = {Key innovations are disruptive evolutionary events that enable a species to escape constraints and rapidly diversify. After 15 years of the Lenski long-term evolution experiment with \textit{Escherichia coli}, cells in one of the twelve populations evolved the ability to utilize citrate, an abundant but previously untapped carbon source in the environment. Descendants of these cells became dominant in the population and subsequently diversified as a consequence of invading this vacant niche. Mutations responsible for the appearance of rudimentary citrate utilization and for refining this ability have been characterized. However, the complete nature of the genetic and/or ecological events that set the stage for this key innovation is unknown. In particular, it is unclear why it took so long for citrate utilization to evolve and why it still has evolved in only one of the twelve \textit{E. coli} populations after 30 years of the Lenski experiment. In this study, we recapitulated the initial mutation needed to evolve citrate utilization in strains isolated from throughout the first 31,500 generations of the history of this population. We found that there was already a slight fitness benefit for this mutation in the original ancestor of the evolution experiment and in other early isolates. However, evolution of citrate utilization was blocked at this point due to competition with other mutations that improved fitness in the original niche. Subsequently, an anti-potentiated genetic background evolved in which it was deleterious to evolve rudimentary citrate utilization. Only later, after further mutations accumulated that restored the benefit of this first-step mutation and the overall rate of adaptation in the population slowed, was citrate utilization likely to evolve. Thus, intense competition and the types of mutations that it favors can lead to short-sighted evolutionary trajectories that hide a stepping stone needed to access a key innovation from many future generations.},
keywords = {Citrate Evolution, Demography and Ecology, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
2017
Good B H; McDonald M J; Barrick J E; Lenski R E; Desai M M
The dynamics of molecular evolution over 60,000 generations Journal Article
Nature, 551 (7678), pp. 45–50, 2017, ISSN: 14764687.
Abstract | Links | BibTeX | Altmetric | Tags: Demography and Ecology, Genome Evolution, Historical Contingency, Mutation Rates, Parallelism and Divergence
@article{Good2017,
title = {The dynamics of molecular evolution over 60,000 generations},
author = {Benjamin H. Good and Michael J. McDonald and Jeffrey E. Barrick and Richard E. Lenski and Michael M. Desai},
url = {http://dx.doi.org/10.1038/nature24287},
doi = {10.1038/nature24287},
issn = {14764687},
year = {2017},
date = {2017-01-01},
urldate = {2017-01-01},
journal = {Nature},
volume = {551},
number = {7678},
pages = {45--50},
publisher = {Nature Publishing Group},
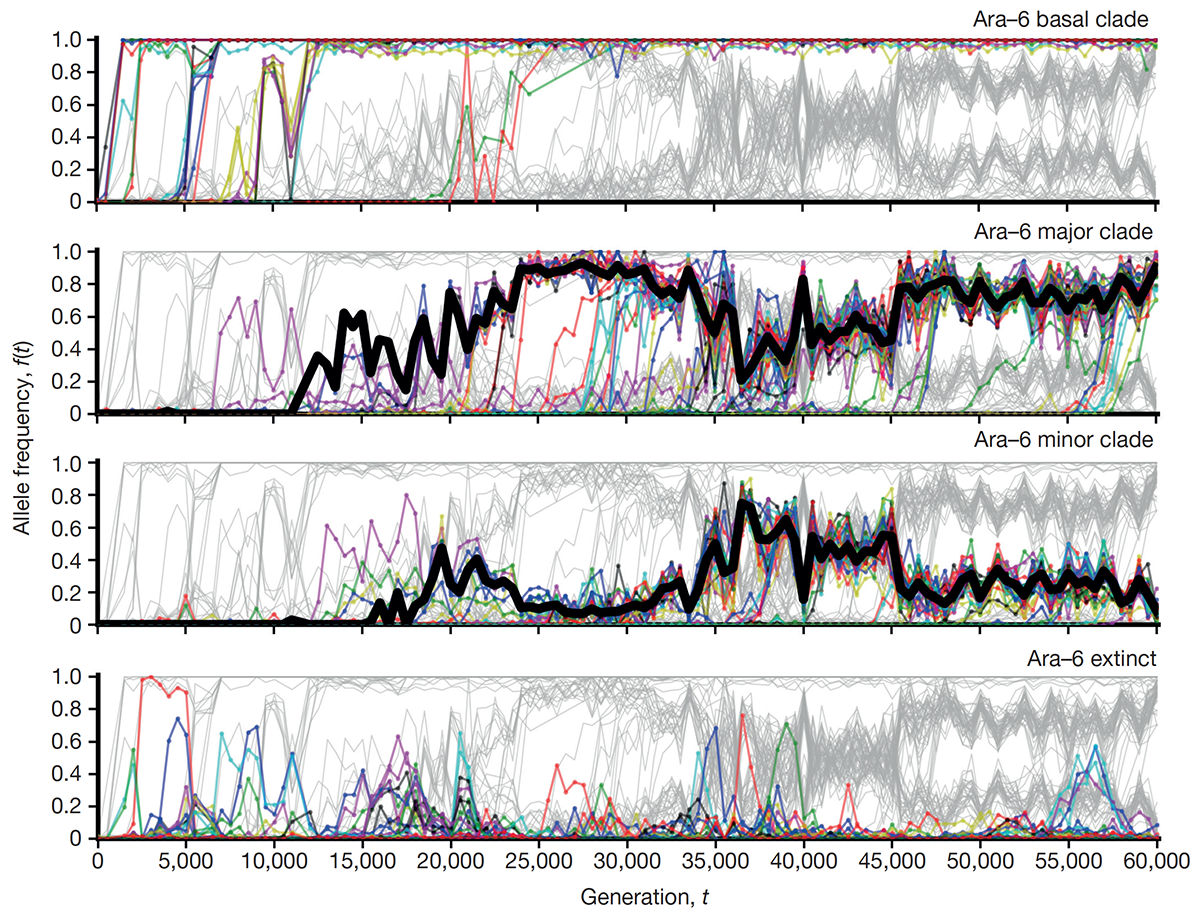
abstract = {The outcomes of evolution are determined by a stochastic dynamical process that governs how mutations arise and spread through a population. However, it is difficult to observe these dynamics directly over long periods and across entire genomes. Here we analyse the dynamics of molecular evolution in twelve experimental populations of \textit{Escherichia coli}, using whole-genome metagenomic sequencing at five hundred-generation intervals through sixty thousand generations. Although the rate of fitness gain declines over time, molecular evolution is characterized by signatures of rapid adaptation throughout the duration of the experiment, with multiple beneficial variants simultaneously competing for dominance in each population. Interactions between ecological and evolutionary processes play an important role, as long-term quasi-stable coexistence arises spontaneously in most populations, and evolution continues within each clade. We also present evidence that the targets of natural selection change over time, as epistasis and historical contingency alter the strength of selection on different genes. Together, these results show that long-term adaptation to a constant environment can be a more complex and dynamic process than is often assumed.},
keywords = {Demography and Ecology, Genome Evolution, Historical Contingency, Mutation Rates, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
2016
Großkopf T; Consuegra J; Gaffé J; Willison J C; Lenski R E; Soyer O S; Schneider D
Metabolic modelling in a dynamic evolutionary framework predicts adaptive diversification of bacteria in a long-term evolution experiment Journal Article
BMC Evolutionary Biology, 16 (1), pp. 163, 2016, ISSN: 1471-2148.
Abstract | Links | BibTeX | Altmetric | Tags: Demography and Ecology, Theory and Simulations
@article{Großkopf2016,
title = {Metabolic modelling in a dynamic evolutionary framework predicts adaptive diversification of bacteria in a long-term evolution experiment},
author = {Tobias Großkopf and Jessika Consuegra and Joël Gaffé and John C. Willison and Richard E. Lenski and Orkun S. Soyer and Dominique Schneider},
url = {https://bmcevolbiol.biomedcentral.com/articles/10.1186/s12862-016-0733-x},
doi = {10.1186/s12862-016-0733-x},
issn = {1471-2148},
year = {2016},
date = {2016-12-01},
urldate = {2016-12-01},
journal = {BMC Evolutionary Biology},
volume = {16},
number = {1},
pages = {163},
publisher = {BMC Evolutionary Biology},
abstract = {Background
Predicting adaptive trajectories is a major goal of evolutionary biology and useful for practical applications. Systems biology has enabled the development of genome-scale metabolic models. However, analysing these models via flux balance analysis (FBA) cannot predict many evolutionary outcomes including adaptive diversification, whereby an ancestral lineage diverges to fill multiple niches. Here we combine in silico evolution with FBA and apply this modelling framework, evoFBA, to a long-term evolution experiment with \textit{Escherichia coli}.
Results
Simulations predicted the adaptive diversification that occurred in one experimental population and generated hypotheses about the mechanisms that promoted coexistence of the diverged lineages. We experimentally tested and, on balance, verified these mechanisms, showing that diversification involved niche construction and character displacement through differential nutrient uptake and altered metabolic regulation.
Conclusion
The evoFBA framework represents a promising new way to model biochemical evolution, one that can generate testable predictions about evolutionary and ecosystem-level outcomes.},
keywords = {Demography and Ecology, Theory and Simulations},
pubstate = {published},
tppubtype = {article}
}
Predicting adaptive trajectories is a major goal of evolutionary biology and useful for practical applications. Systems biology has enabled the development of genome-scale metabolic models. However, analysing these models via flux balance analysis (FBA) cannot predict many evolutionary outcomes including adaptive diversification, whereby an ancestral lineage diverges to fill multiple niches. Here we combine in silico evolution with FBA and apply this modelling framework, evoFBA, to a long-term evolution experiment with Escherichia coli.
Results
Simulations predicted the adaptive diversification that occurred in one experimental population and generated hypotheses about the mechanisms that promoted coexistence of the diverged lineages. We experimentally tested and, on balance, verified these mechanisms, showing that diversification involved niche construction and character displacement through differential nutrient uptake and altered metabolic regulation.
Conclusion
The evoFBA framework represents a promising new way to model biochemical evolution, one that can generate testable predictions about evolutionary and ecosystem-level outcomes.
2015
Turner C B; Blount Z D; Lenski R E
Replaying Evolution to Test the Cause of Extinction of One Ecotype in an Experimentally Evolved Population Journal Article
PLOS ONE, 10 (11), pp. e0142050, 2015, ISSN: 1932-6203.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Demography and Ecology, Historical Contingency
@article{Turner2015,
title = {Replaying Evolution to Test the Cause of Extinction of One Ecotype in an Experimentally Evolved Population},
author = {Caroline B. Turner and Zachary D. Blount and Richard E. Lenski},
editor = {Frederick M. Cohan},
url = {https://dx.plos.org/10.1371/journal.pone.0142050},
doi = {10.1371/journal.pone.0142050},
issn = {1932-6203},
year = {2015},
date = {2015-11-01},
urldate = {2015-11-01},
journal = {PLOS ONE},
volume = {10},
number = {11},
pages = {e0142050},
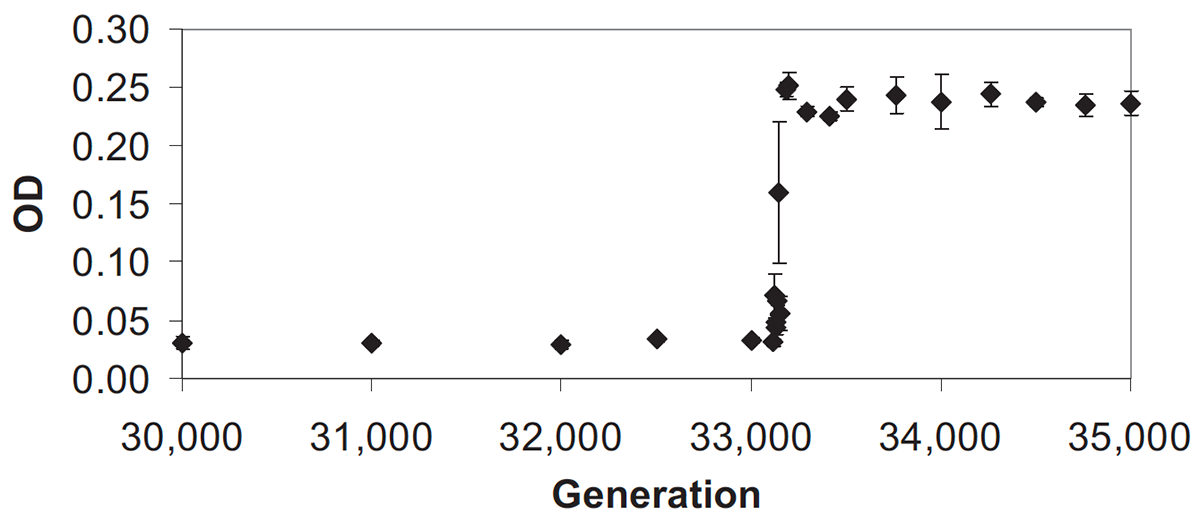
abstract = {In a long-term evolution experiment with \textit{Escherichia coli}, bacteria in one of twelve populations evolved the ability to consume citrate, a previously unexploited resource in a glucoselimited medium. This innovation led to the frequency-dependent coexistence of citrate-consuming (Cit+) and non-consuming (Cit-) ecotypes, with Cit-bacteria persisting on the exogenously supplied glucose as well as other carbon molecules released by the Cit+ bacteria. After more than 10,000 generations of coexistence, however, the Cit-lineage went extinct; cells with the Cit-phenotype dropped to levels below detection, and the Cit-clade could not be detected by molecular assays based on its unique genotype. We hypothesized that this extinction was a deterministic outcome of evolutionary change within the population, specifically the appearance of a more-fit Cit+ ecotype that competitively excluded the Cit-ecotype. We tested this hypothesis by re-evolving the population from a frozen population sample taken within 500 generations of the extinction and from another sample taken several thousand generations earlier, in each case for 500 generations and with 20-fold replication. To our surprise, the Cit-type did not go extinct in any of these replays, and Cit-cells also persisted in a single replicate that was propagated for 2,500 generations. Even more unexpectedly, we showed that the Cit-ecotype could reinvade the Cit+ population after its extinction. Taken together, these results indicate that the extinction of the Cit-ecotype was not a deterministic outcome driven by competitive exclusion by the Cit+ ecotype. The extinction also cannot be explained by demographic stochasticity alone, as the population size of the Cit-ecotype should have been many thousands of cells even during the daily transfer events. Instead, we infer that the extinction must have been caused by a rare chance event in which some aspect of the experimental conditions was inadvertently perturbed.},
keywords = {Citrate Evolution, Demography and Ecology, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
Ribeck N; Lenski R E
Modeling and quantifying frequency-dependent fitness in microbial populations with cross-feeding interactions Journal Article
Evolution, 69 (5), pp. 1313–1320, 2015, ISSN: 00143820.
Abstract | Links | BibTeX | Altmetric | Tags: Demography and Ecology, Theory and Simulations
@article{Ribeck2015,
title = {Modeling and quantifying frequency-dependent fitness in microbial populations with cross-feeding interactions},
author = {Noah Ribeck and Richard E. Lenski},
url = {https://onlinelibrary.wiley.com/doi/10.1111/evo.12645},
doi = {10.1111/evo.12645},
issn = {00143820},
year = {2015},
date = {2015-05-01},
urldate = {2015-05-01},
journal = {Evolution},
volume = {69},
number = {5},
pages = {1313--1320},
abstract = {Coexistence of two or more populations by frequency-dependent selection is common in nature, and it often arises even in well-mixed experiments with microbes. If ecology is to be incorporated into models of population genetics, then it is important to represent accurately the functional form of frequency-dependent interactions. However, measuring this functional form is problematic for traditional fitness assays, which assume a constant fitness difference between competitors over the course of an assay. Here, we present a theoretical framework for measuring the functional form of frequency-dependent fitness by accounting for changes in abundance and relative fitness during a competition assay. Using two examples of ecological coexistence that arose in a long-term evolution experiment with \textit{Escherichia coli}, we illustrate accurate quantification of the functional form of frequency-dependent relative fitness. Using a Monod-type model of growth dynamics, we show that two ecotypes in a typical cross-feeding interaction-such as when one bacterial population uses a byproduct generated by another-yields relative fitness that is linear with relative frequency.},
keywords = {Demography and Ecology, Theory and Simulations},
pubstate = {published},
tppubtype = {article}
}
Maddamsetti R; Lenski R E; Barrick J E
Adaptation, Clonal Interference, and Frequency-Dependent Interactions in a Long-Term Evolution Experiment with Escherichia coli Journal Article
Genetics, 200 (2), pp. 619-631, 2015, ISSN: 1943-2631.
Abstract | Links | BibTeX | Altmetric | Tags: Demography and Ecology, Fitness Trajectories, Genome Evolution
@article{nokey,
title = {Adaptation, Clonal Interference, and Frequency-Dependent Interactions in a Long-Term Evolution Experiment with \emph{Escherichia coli}},
author = {Rohan Maddamsetti and Richard E. Lenski and Jeffrey E. Barrick},
url = {https://academic.oup.com/genetics/article/200/2/619/5936186},
doi = {10.1534/genetics.115.176677},
issn = {1943-2631},
year = {2015},
date = {2015-04-24},
urldate = {2015-04-24},
journal = {Genetics},
volume = {200},
number = {2},
pages = {619-631},
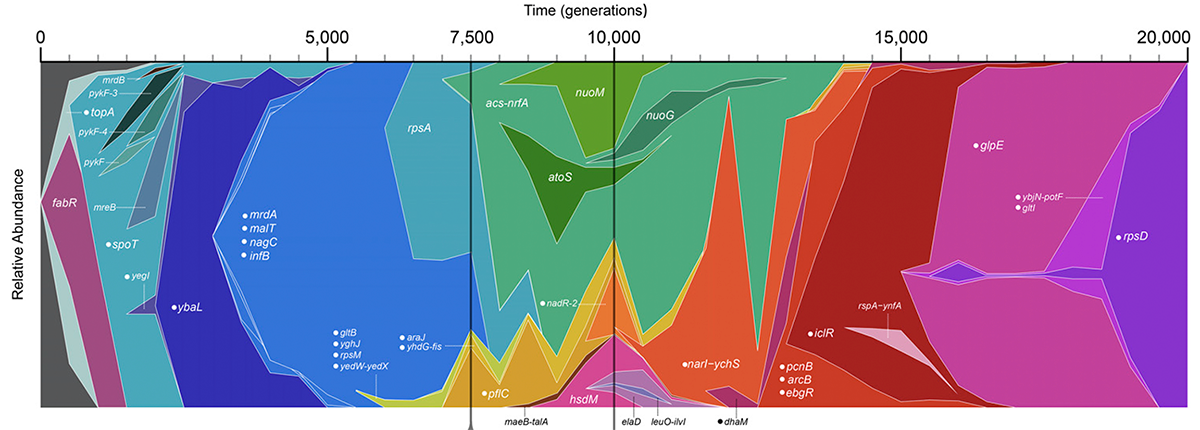
abstract = {Twelve replicate populations of \textit{Escherichia coli} have been evolving in the laboratory for >25 years and 60,000 generations. We analyzed bacteria from whole-population samples frozen every 500 generations through 20,000 generations for one well-studied population, called Ara−1. By tracking 42 known mutations in these samples, we reconstructed the history of this population’s genotypic evolution over this period. The evolutionary dynamics of Ara−1 show strong evidence of selective sweeps as well as clonal interference between competing lineages bearing different beneficial mutations. In some cases, sets of several mutations approached fixation simultaneously, often conveying no information about their order of origination; we present several possible explanations for the existence of these mutational cohorts. Against a backdrop of rapid selective sweeps both earlier and later, two genetically diverged clades coexisted for >6000 generations before one went extinct. In that time, many additional mutations arose in the clade that eventually prevailed. We show that the clades evolved a frequency-dependent interaction, which prevented the immediate competitive exclusion of either clade, but which collapsed as beneficial mutations accumulated in the clade that prevailed. Clonal interference and frequency dependence can occur even in the simplest microbial populations. Furthermore, frequency dependence may generate dynamics that extend the period of coexistence that would otherwise be sustained by clonal interference alone.},
keywords = {Demography and Ecology, Fitness Trajectories, Genome Evolution},
pubstate = {published},
tppubtype = {article}
}
2014
Plucain J; Hindré T; Gac M L; Tenaillon O; Cruveiller S; Medigue C; Leiby N; Harcombe W R; Marx C J; Lenski R E; Schneider D
Epistasis and Allele Specificity in the Emergence of a Stable Polymorphism in Escherichia coli Journal Article
Science, 343 (6177), pp. 1366–1369, 2014, ISSN: 0036-8075.
Abstract | Links | BibTeX | Altmetric | Tags: Demography and Ecology, Genotypes and Phenotypes
@article{Plucain2014,
title = {Epistasis and Allele Specificity in the Emergence of a Stable Polymorphism in \textit{Escherichia coli}},
author = {Jessica Plucain and Thomas Hindré and Mickaël Le Gac and Olivier Tenaillon and Stéphane Cruveiller and Claudine Medigue and Nicholas Leiby and William R. Harcombe and Christopher J. Marx and Richard E. Lenski and Dominique Schneider},
url = {https://www.sciencemag.org/lookup/doi/10.1126/science.1248688},
doi = {10.1126/science.1248688},
issn = {0036-8075},
year = {2014},
date = {2014-03-01},
urldate = {2014-03-01},
journal = {Science},
volume = {343},
number = {6177},
pages = {1366--1369},
abstract = {Ecological opportunities promote population divergence into coexisting lineages. However, the genetic mechanisms that enable new lineages to exploit these opportunities are poorly understood except in cases of single mutations. We examined how two \textit{Escherichia coli} lineages diverged from their common ancestor at the outset of a long-term coexistence. By sequencing genomes and reconstructing the genetic history of one lineage, we showed that three mutations together were sufficient to produce the frequency-dependent fitness effects that allowed this lineage to invade and stably coexist with the other. These mutations all affected regulatory genes and collectively caused substantial metabolic changes. Moreover, the particular derived alleles were critical for the initial divergence and invasion, indicating that the establishment of this polymorphism depended on specific epistatic interactions.},
keywords = {Demography and Ecology, Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}
2012
Gac M L; Plucain J; Hindré T; Lenski R E; Schneider D
Ecological and evolutionary dynamics of coexisting lineages during a long-term experiment with Escherichia coli Journal Article
Proceedings of the National Academy of Sciences of the United States of America, 109 (24), pp. 9487–9492, 2012, ISSN: 0027-8424.
Abstract | Links | BibTeX | Altmetric | Tags: Demography and Ecology, Genotypes and Phenotypes
@article{LeGac2012,
title = {Ecological and evolutionary dynamics of coexisting lineages during a long-term experiment with \textit{Escherichia coli}},
author = {Mickaël Le Gac and Jessica Plucain and Thomas Hindré and Richard E. Lenski and Dominique Schneider},
url = {http://www.pnas.org/cgi/doi/10.1073/pnas.1207091109},
doi = {10.1073/pnas.1207091109},
issn = {0027-8424},
year = {2012},
date = {2012-06-01},
urldate = {2012-06-01},
journal = {Proceedings of the National Academy of Sciences of the United States of America},
volume = {109},
number = {24},
pages = {9487--9492},
abstract = {Closely related organisms usually occupy similar ecological niches, leading to intense competition and even extinction. Such competition also can promote rapid phenotypic evolution and ecological divergence. This process may end with the stable occupation of distinct niches or, alternatively, may entail repeated bouts of evolution. Here we examine two \textit{Escherichia coli} lineages, called L and S, that coexisted for more than 30,000 generations after diverging from a common ancestor. Both lineages underwent sustained phenotypic evolution based on global transcription and resource utilization profiles, with L seeming to encroach over time on the catabolic profile of S. Reciprocal invasion experiments with L and S clones from the same or different generations revealed evolutionary changes in their interaction, including an asymmetry that confirmed the encroachment by L on the niche of the S lineage. In general, L and S clones from the same generation showed negative frequency-dependent effects, consistent with stable coexistence. However, L clones could invade S clones from both earlier and later generations, whereas S clones could invade only L clones from earlier generations. In this system, the long-term coexistence of competing lineages evidently depended on successive rounds of evolution, rather than on initial divergence followed by a static equilibrium.},
keywords = {Demography and Ecology, Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}
2009
Rozen D E; Philippe N; de Visser J A G M; Lenski R E; Schneider D
Death and cannibalism in a seasonal environment facilitate bacterial coexistence Journal Article
Ecology Letters, 12 (1), pp. 34–44, 2009, ISSN: 1461-023X.
Abstract | Links | BibTeX | Altmetric | Tags: Demography and Ecology
@article{de037636e2924110bf401bac283d1c0f,
title = {Death and cannibalism in a seasonal environment facilitate bacterial coexistence},
author = {Daniel E. Rozen and Nadège Philippe and J. Arjan G. M. {de Visser} and Richard E. Lenski and Dominique Schneider},
url = {https://onlinelibrary.wiley.com/doi/10.1111/j.1461-0248.2008.01257.x},
doi = {10.1111/j.1461-0248.2008.01257.x},
issn = {1461-023X},
year = {2009},
date = {2009-01-01},
urldate = {2009-01-01},
journal = {Ecology Letters},
volume = {12},
number = {1},
pages = {34--44},
publisher = {Wiley},
abstract = {Bacterial populations can evolve and adapt to become diverse niche specialists, even in seemingly homogeneous environments. One source of this diversity arises from newly 'constructed' niches that result from the activities of the bacteria themselves. Ecotypes specialized to exploit these distinct niches can subsequently coexist via frequency-dependent interactions. Here, we describe a novel form of niche construction that is based upon differential death and cannibalism, and which evolved during 20 000 generations of experimental evolution in \textit{Escherichia coli} in a seasonal environment with alternating growth and starvation. In one of 12 populations, two monophyletic ecotypes, S and L, evolved that stably coexist with one another. When grown and then starved in monoculture, the death rate of S exceeds that of L, whereas the reverse is observed in mixed cultures. As shown by experiments and numerical simulations, the competitive advantage of S cells is increased by extending the period of starvation, and this advantage results from their cannibalization of the debris of lysed L cells, which allows the S cells to increase both their growth rate and total cell density. At the molecular level, the polymorphism is associated with divergence in the activity of the alternative sigma factor RpoS, with S cells displaying no detectable activity, while L cells show increased activity relative to the ancestral genotype. Our results extend the repertoire of known cross-feeding mechanisms in microbes to include cannibalism during starvation, and confirm the central roles for niche construction and seasonality in the maintenance of microbial polymorphisms},
keywords = {Demography and Ecology},
pubstate = {published},
tppubtype = {article}
}
2008
Blount Z D; Borland C Z; Lenski R E
Historical contingency and the evolution of a key innovation in an experimental population of Escherichia coli Journal Article
Proceedings of the National Academy of Sciences of the United States of America, 105 (23), pp. 7899–7906, 2008, ISSN: 0027-8424.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Demography and Ecology, Historical Contingency
@article{Blount2008,
title = {Historical contingency and the evolution of a key innovation in an experimental population of \emph{Escherichia coli}},
author = {Zachary D. Blount and Christina Z. Borland and Richard E. Lenski},
url = {http://www.pnas.org/cgi/doi/10.1073/pnas.0803151105},
doi = {10.1073/pnas.0803151105},
issn = {0027-8424},
year = {2008},
date = {2008-06-01},
urldate = {2008-06-01},
journal = {Proceedings of the National Academy of Sciences of the United States of America},
volume = {105},
number = {23},
pages = {7899--7906},
abstract = {The role of historical contingency in evolution has been much debated, but rarely tested. Twelve initially identical populations of \textit{Escherichia coli} were founded in 1988 to investigate this issue. They have since evolved in a glucose-limited medium that also contains citrate, which \textit{E. coli} cannot use as a carbon source under oxic conditions. No population evolved the capacity to exploit citrate for >30,000 generations, although each population tested billions of mutations. A citrate-using (Cit^{+}) variant finally evolved in one population by 31,500 generations, causing an increase in population size and diversity. The long-delayed and unique evolution of this function might indicate the involvement of some extremely rare mutation. Alternately, it may involve an ordinary mutation, but one whose physical occurrence or phenotypic expression is contingent on prior mutations in that population. We tested these hypotheses in experiments that “replayed” evolution from different points in that population's history. We observed no Cit^{+} mutants among 8.4 × 10^{12} ancestral cells, nor among 9 × 10^{12} cells from 60 clones sampled in the first 15,000 generations. However, we observed a significantly greater tendency for later clones to evolve Cit^{+}, indicating that some potentiating mutation arose by 20,000 generations. This potentiating change increased the mutation rate to Cit^{+} but did not cause generalized hypermutability. Thus, the evolution of this phenotype was contingent on the particular history of that population. More generally, we suggest that historical contingency is especially important when it facilitates the evolution of key innovations that are not easily evolved by gradual, cumulative selection.},
keywords = {Citrate Evolution, Demography and Ecology, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}