2024
Izutsu M; Lake D M; Matson Z W D; Dodson J P; Lenski R E
Effects of periodic bottlenecks on the dynamics of adaptive evolution in microbial populations Journal Article
Microbiology (Reading, England), 170 (9), pp. 001494, 2024, ISSN: 1465-2080.
Abstract | Links | BibTeX | Altmetric | Tags: Descendant Experiments, Fitness Trajectories, Theory and Simulations
@article{Izutsu_2024,
title = {Effects of periodic bottlenecks on the dynamics of adaptive evolution in microbial populations},
author = {Minako Izutsu and Devin M. Lake and Zachary W. D. Matson and Jack P. Dodson and Richard E. Lenski},
doi = {10.1099/mic.0.001494},
issn = {1465-2080},
year = {2024},
date = {2024-09-18},
urldate = {2021-12-30},
journal = {Microbiology (Reading, England)},
volume = {170},
number = {9},
pages = {001494},
publisher = {LTEE},
abstract = {Population bottlenecks can impact the rate of adaptation in evolving populations. On the one hand, each bottleneck reduces the genetic variation that fuels adaptation. On the other hand, each founder that survives a bottleneck can undergo more generations and leave more descendants in a resource-limited environment, which allows surviving beneficial mutations to spread more quickly. A theoretical model predicted that the rate of fitness gains should be maximized using ~8-fold dilutions. Here we investigate the impact of repeated bottlenecks on the dynamics of adaptation using numerical simulations and experimental populations of Escherichia coli. Our simulations confirm the model's prediction when populations evolve in a regime where beneficial mutations are rare and waiting times between successful mutations are long. However, more extreme dilutions maximize fitness gains in simulations when beneficial mutations are common and clonal interference prevents most of them from fixing. To examine these predictions, we propagated 48 \textit{E. coli} populations with 2-, 8-, 100-, and 1000-fold dilutions for 150 days. Adaptation began earlier and fitness gains were greater with 100- and 1000-fold dilutions than with 8-fold dilutions, consistent with the simulations when beneficial mutations are common. However, the selection pressures in the 2-fold treatment were qualitatively different from the other treatments, violating a critical assumption of the model and simulations. Thus, varying the dilution factor during periodic bottlenecks can have multiple effects on the dynamics of adaptation caused by differential losses of diversity, different numbers of generations, and altered selection.},
keywords = {Descendant Experiments, Fitness Trajectories, Theory and Simulations},
pubstate = {published},
tppubtype = {article}
}
2023
Mukherjee A; Ealy J; Huang Y; Benites N C; Polk M; Basan M
Coexisting ecotypes in long-term evolution emerged from interacting trade-offs Journal Article
Nature Communications, 14 (1), pp. 3805, 2023, ISSN: 2041-1723.
Abstract | Links | BibTeX | Altmetric | Tags: Demography and Ecology, Theory and Simulations
@article{Mukherjee2023,
title = {Coexisting ecotypes in long-term evolution emerged from interacting trade-offs},
author = {Avik Mukherjee and Jade Ealy and Yanqing Huang and Nina Catherine Benites and Mark Polk and Markus Basan},
url = {https://www.nature.com/articles/s41467-023-39471-9},
doi = {10.1038/s41467-023-39471-9},
issn = {2041-1723},
year = {2023},
date = {2023-06-26},
journal = {Nature Communications},
volume = {14},
number = {1},
pages = {3805},
abstract = {Evolution of complex communities of coexisting microbes remains poorly understood. The long-term evolution experiment on \textit{Escherichia coli} (LTEE) revealed the spontaneous emergence of stable coexistence of multiple ecotypes, which persisted for more than 14,000 generations of continuous evolution. Here, using a combination of experiments and computer simulations, we show that the emergence and persistence of this phenomenon can be explained by the combination of two interacting trade-offs, rooted in biochemical constraints: First, faster growth is enabled by higher fermentation and obligate acetate excretion. Second, faster growth results in longer lag times when utilizing acetate after glucose is depleted. This combination creates an ecological niche for a slower-growing ecotype, specialized in switching to acetate. These findings demonstrate that trade-offs can give rise to surprisingly complex communities with evolutionarily stable coexistence of multiple variants in even the simplest environments.},
keywords = {Demography and Ecology, Theory and Simulations},
pubstate = {published},
tppubtype = {article}
}
2018
Bajić D; Vila J C C; Blount Z D; Sánchez A
On the deformability of an empirical fitness landscape by microbial evolution Journal Article
Proceedings of the National Academy of Sciences of the United States of America, 115 (44), pp. 11286-11291, 2018, ISSN: 0027-8424.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Demography and Ecology, Theory and Simulations
@article{nokey,
title = {On the deformability of an empirical fitness landscape by microbial evolution},
author = {Djordje Bajić and Jean C. C. Vila and Zachary D. Blount and Alvaro Sánchez},
url = {http://www.pnas.org/lookup/doi/10.1073/pnas.1808485115},
doi = {10.1073/pnas.1808485115},
issn = {0027-8424},
year = {2018},
date = {2018-10-30},
urldate = {2018-10-30},
journal = {Proceedings of the National Academy of Sciences of the United States of America},
volume = {115},
number = {44},
pages = {11286-11291},
abstract = {A fitness landscape is a map between the genotype and its reproductive success in a given environment. The topography of fitness landscapes largely governs adaptive dynamics, constraining evolutionary trajectories and the predictability of evolution. Theory suggests that this topography can be deformed by mutations that produce substantial changes to the environment. Despite its importance, the deformability of fitness landscapes has not been systematically studied beyond abstract models, and little is known about its reach and consequences in empirical systems. Here we have systematically characterized the deformability of the genome-wide metabolic fitness landscape of the bacterium \textit{Escherichia coli}. Deformability is quantified by the noncommutativity of epistatic interactions, which we experimentally demonstrate in mutant strains on the path to an evolutionary innovation. Our analysis shows that the deformation of fitness landscapes by metabolic mutations rarely affects evolutionary trajectories in the short range. However, mutations with large environmental effects produce long-range landscape deformations in distant regions of the genotype space that affect the fitness of later descendants. Our results therefore suggest that, even in situations in which mutations have strong environmental effects, fitness landscapes may retain their power to forecast evolution over small mutational distances despite the potential attenuation of that power over longer evolutionary trajectories. Our methods and results provide an avenue for integrating adaptive and eco-evolutionary dynamics with complex genetics and genomics. },
keywords = {Citrate Evolution, Demography and Ecology, Theory and Simulations},
pubstate = {published},
tppubtype = {article}
}
2016
Großkopf T; Consuegra J; Gaffé J; Willison J C; Lenski R E; Soyer O S; Schneider D
Metabolic modelling in a dynamic evolutionary framework predicts adaptive diversification of bacteria in a long-term evolution experiment Journal Article
BMC Evolutionary Biology, 16 (1), pp. 163, 2016, ISSN: 1471-2148.
Abstract | Links | BibTeX | Altmetric | Tags: Demography and Ecology, Theory and Simulations
@article{Großkopf2016,
title = {Metabolic modelling in a dynamic evolutionary framework predicts adaptive diversification of bacteria in a long-term evolution experiment},
author = {Tobias Großkopf and Jessika Consuegra and Joël Gaffé and John C. Willison and Richard E. Lenski and Orkun S. Soyer and Dominique Schneider},
url = {https://bmcevolbiol.biomedcentral.com/articles/10.1186/s12862-016-0733-x},
doi = {10.1186/s12862-016-0733-x},
issn = {1471-2148},
year = {2016},
date = {2016-12-01},
urldate = {2016-12-01},
journal = {BMC Evolutionary Biology},
volume = {16},
number = {1},
pages = {163},
publisher = {BMC Evolutionary Biology},
abstract = {Background
Predicting adaptive trajectories is a major goal of evolutionary biology and useful for practical applications. Systems biology has enabled the development of genome-scale metabolic models. However, analysing these models via flux balance analysis (FBA) cannot predict many evolutionary outcomes including adaptive diversification, whereby an ancestral lineage diverges to fill multiple niches. Here we combine in silico evolution with FBA and apply this modelling framework, evoFBA, to a long-term evolution experiment with \textit{Escherichia coli}.
Results
Simulations predicted the adaptive diversification that occurred in one experimental population and generated hypotheses about the mechanisms that promoted coexistence of the diverged lineages. We experimentally tested and, on balance, verified these mechanisms, showing that diversification involved niche construction and character displacement through differential nutrient uptake and altered metabolic regulation.
Conclusion
The evoFBA framework represents a promising new way to model biochemical evolution, one that can generate testable predictions about evolutionary and ecosystem-level outcomes.},
keywords = {Demography and Ecology, Theory and Simulations},
pubstate = {published},
tppubtype = {article}
}
Predicting adaptive trajectories is a major goal of evolutionary biology and useful for practical applications. Systems biology has enabled the development of genome-scale metabolic models. However, analysing these models via flux balance analysis (FBA) cannot predict many evolutionary outcomes including adaptive diversification, whereby an ancestral lineage diverges to fill multiple niches. Here we combine in silico evolution with FBA and apply this modelling framework, evoFBA, to a long-term evolution experiment with Escherichia coli.
Results
Simulations predicted the adaptive diversification that occurred in one experimental population and generated hypotheses about the mechanisms that promoted coexistence of the diverged lineages. We experimentally tested and, on balance, verified these mechanisms, showing that diversification involved niche construction and character displacement through differential nutrient uptake and altered metabolic regulation.
Conclusion
The evoFBA framework represents a promising new way to model biochemical evolution, one that can generate testable predictions about evolutionary and ecosystem-level outcomes.
Ribeck N; Mulka J S; Zaman L; Connelly B D; Lenski R E
Competition between continuously evolving lineages in asexual populations Journal Article
bioRxiv, 2016.
Abstract | Links | BibTeX | Altmetric | Tags: Theory and Simulations
@article{Ribeck062976,
title = {Competition between continuously evolving lineages in asexual populations},
author = {Noah Ribeck and Joseph S. Mulka and Luis Zaman and Brian D. Connelly and Richard E. Lenski},
url = {https://www.biorxiv.org/content/early/2016/07/10/062976},
doi = {10.1101/062976},
year = {2016},
date = {2016-01-01},
urldate = {2016-01-01},
journal = {bioRxiv},
publisher = {Cold Spring Harbor Laboratory},
abstract = {In an asexual population, the fate of a beneficial mutation depends on how its lineage competes against other mutant lineages in the population. With high beneficial mutation rates or large population sizes, competition between contending mutations is strong, and successful lineages can accumulate multiple mutations before any single one achieves fixation. Most current theory about asexual population dynamics either neglects this multiple-mutations regime or introduces simplifying assumptions that may not apply. Here, we develop a theoretical framework that describes the dynamics of adaptation and substitution over all mutation-rate regimes by conceptualizing the population as a collection of continuously adapting lineages. This model of textquotedblleftlineage interferencetextquotedblright shows that each new mutanttextquoterights advantage over the rest of the population must be above a critical threshold in order to likely achieve fixation, and we derive a simple expression for that threshold. We apply this framework to examine the role of beneficial mutations with different effect sizes across the transition to the multiple-mutations regime.},
keywords = {Theory and Simulations},
pubstate = {published},
tppubtype = {article}
}
2015
Ribeck N; Lenski R E
Modeling and quantifying frequency-dependent fitness in microbial populations with cross-feeding interactions Journal Article
Evolution, 69 (5), pp. 1313–1320, 2015, ISSN: 00143820.
Abstract | Links | BibTeX | Altmetric | Tags: Demography and Ecology, Theory and Simulations
@article{Ribeck2015,
title = {Modeling and quantifying frequency-dependent fitness in microbial populations with cross-feeding interactions},
author = {Noah Ribeck and Richard E. Lenski},
url = {https://onlinelibrary.wiley.com/doi/10.1111/evo.12645},
doi = {10.1111/evo.12645},
issn = {00143820},
year = {2015},
date = {2015-05-01},
urldate = {2015-05-01},
journal = {Evolution},
volume = {69},
number = {5},
pages = {1313--1320},
abstract = {Coexistence of two or more populations by frequency-dependent selection is common in nature, and it often arises even in well-mixed experiments with microbes. If ecology is to be incorporated into models of population genetics, then it is important to represent accurately the functional form of frequency-dependent interactions. However, measuring this functional form is problematic for traditional fitness assays, which assume a constant fitness difference between competitors over the course of an assay. Here, we present a theoretical framework for measuring the functional form of frequency-dependent fitness by accounting for changes in abundance and relative fitness during a competition assay. Using two examples of ecological coexistence that arose in a long-term evolution experiment with \textit{Escherichia coli}, we illustrate accurate quantification of the functional form of frequency-dependent relative fitness. Using a Monod-type model of growth dynamics, we show that two ecotypes in a typical cross-feeding interaction-such as when one bacterial population uses a byproduct generated by another-yields relative fitness that is linear with relative frequency.},
keywords = {Demography and Ecology, Theory and Simulations},
pubstate = {published},
tppubtype = {article}
}
2013
Wiser M J; Ribeck N; Lenski R E
Long-Term Dynamics of Adaptation in Asexual Populations Journal Article
Science, 342 (6164), pp. 1364–1367, 2013, ISSN: 0036-8075.
Abstract | Links | BibTeX | Altmetric | Tags: Fitness Trajectories, Mutation Rates, Theory and Simulations
@article{Wiser2013,
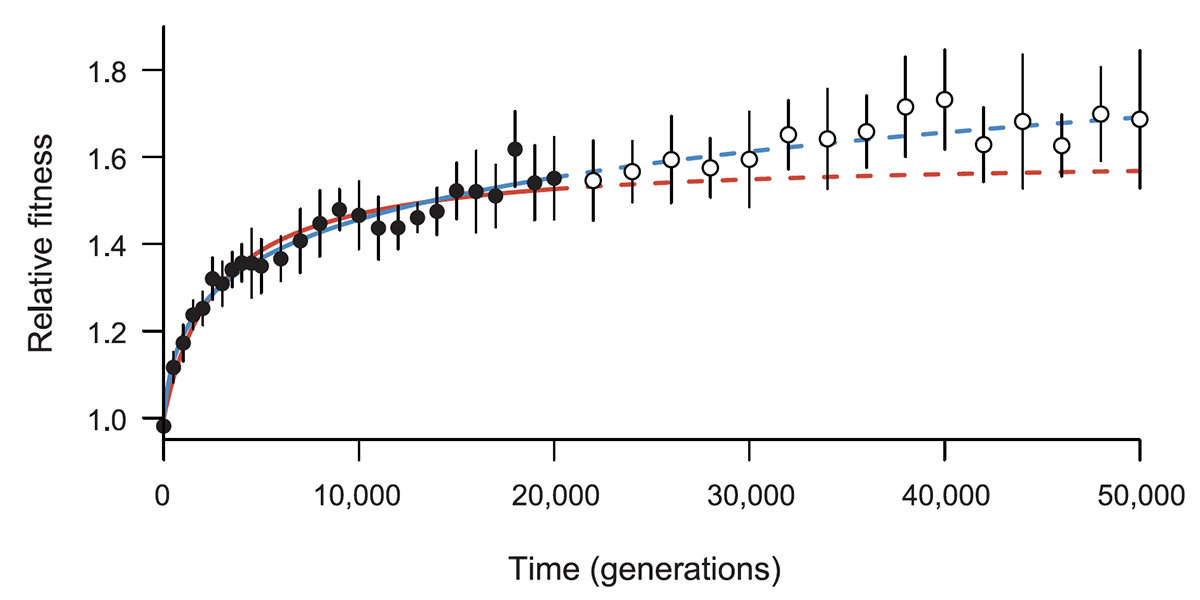
title = {Long-Term Dynamics of Adaptation in Asexual Populations},
author = {Michael J. Wiser and Noah Ribeck and Richard E. Lenski},
url = {https://www.sciencemag.org/lookup/doi/10.1126/science.1243357},
doi = {10.1126/science.1243357},
issn = {0036-8075},
year = {2013},
date = {2013-12-01},
urldate = {2013-12-01},
journal = {Science},
volume = {342},
number = {6164},
pages = {1364--1367},
abstract = {Experimental studies of evolution have increased greatly in number in recent years, stimulated by the growing power of genomic tools. However, organismal fitness remains the ultimate metric for interpreting these experiments, and the dynamics of fitness remain poorly understood over long time scales. Here, we examine fitness trajectories for 12 \textit{Escherichia coli} populations during 50,000 generations. Mean fitness appears to increase without bound, consistent with a power law. We also derive this power-law relation theoretically by incorporating clonal interference and diminishing-returns epistasis into a dynamical model of changes in mean fitness over time.},
keywords = {Fitness Trajectories, Mutation Rates, Theory and Simulations},
pubstate = {published},
tppubtype = {article}
}
Wielgoss S; Barrick J E; Tenaillon O; Wiser M J; Dittmar W J; Cruveiller S; Chane-Woon-Ming B; Médigue C; Lenski R E; Schneider D
Mutation rate dynamics in a bacterial population reflect tension between adaptation and genetic load Journal Article
Proceedings of the National Academy of Sciences of the United States of America, 110 (1), pp. 222–227, 2013, ISSN: 0027-8424.
Abstract | Links | BibTeX | Altmetric | Tags: Fitness Trajectories, Genome Evolution, Genotypes and Phenotypes, Mutation Rates, Theory and Simulations
@article{Wielgoss2013,
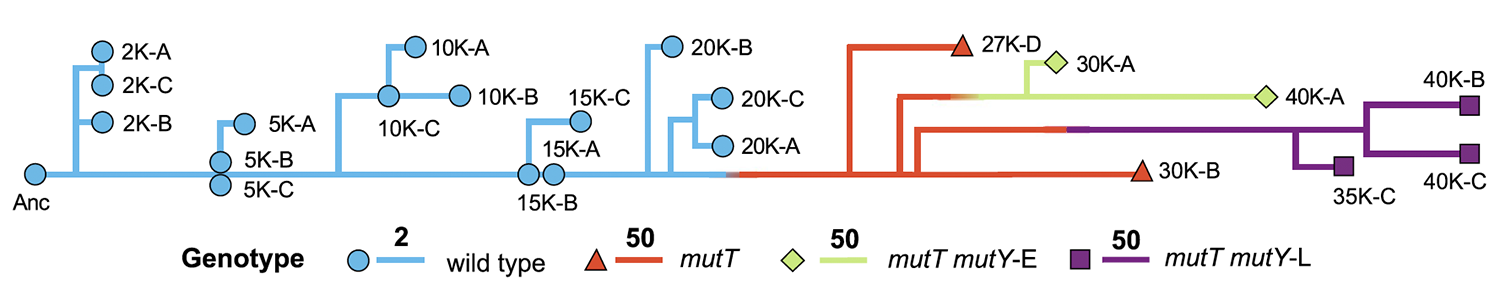
title = {Mutation rate dynamics in a bacterial population reflect tension between adaptation and genetic load},
author = {Sébastien Wielgoss and Jeffrey E. Barrick and Olivier Tenaillon and Michael J. Wiser and W. James Dittmar and Stéphane Cruveiller and Béatrice Chane-Woon-Ming and Claudine Médigue and Richard E. Lenski and Dominique Schneider},
url = {http://www.pnas.org/cgi/doi/10.1073/pnas.1219574110},
doi = {10.1073/pnas.1219574110},
issn = {0027-8424},
year = {2013},
date = {2013-01-01},
urldate = {2013-01-01},
journal = {Proceedings of the National Academy of Sciences of the United States of America},
volume = {110},
number = {1},
pages = {222--227},
abstract = {Mutations are the ultimate source of heritable variation for evolution. Understanding how mutation rates themselves evolve is thus essential for quantitatively understanding many evolutionary processes. According to theory, mutation rates should be minimized for well-adapted populations living in stable environments, whereas hypermutators may evolve if conditions change. However, the long-term fate of hypermutators is unknown. Using a phylogenomic approach, we found that an adapting \textit{Escherichia coli} population that first evolved a \textit{mutT} hypermutator phenotype was later invaded by two independent lineages with \textit{mutY} mutations that reduced genome-wide mutation rates. Applying neutral theory to synonymous substitutions, we dated the emergence of these mutations and inferred that the \textit{mutT} mutation increased the point-mutation rate by ~150-fold, whereas the \textit{mutY} mutations reduced the rate by ~40-60%, with a corresponding decrease in the genetic load. Thus, the long-term fate of the hypermutators was governed by the selective advantage arising from a reduced mutation rate as the potential for further adaptation declined.},
keywords = {Fitness Trajectories, Genome Evolution, Genotypes and Phenotypes, Mutation Rates, Theory and Simulations},
pubstate = {published},
tppubtype = {article}
}
1998
Gerrish P J; Lenski R E
The fate of competing beneficial mutations in an asexual population Journal Article
Genetica, 102-103 , pp. 127-44, 1998, ISBN: 978-94-010-6193-3.
Abstract | Links | BibTeX | Altmetric | Tags: Theory and Simulations
@article{article,
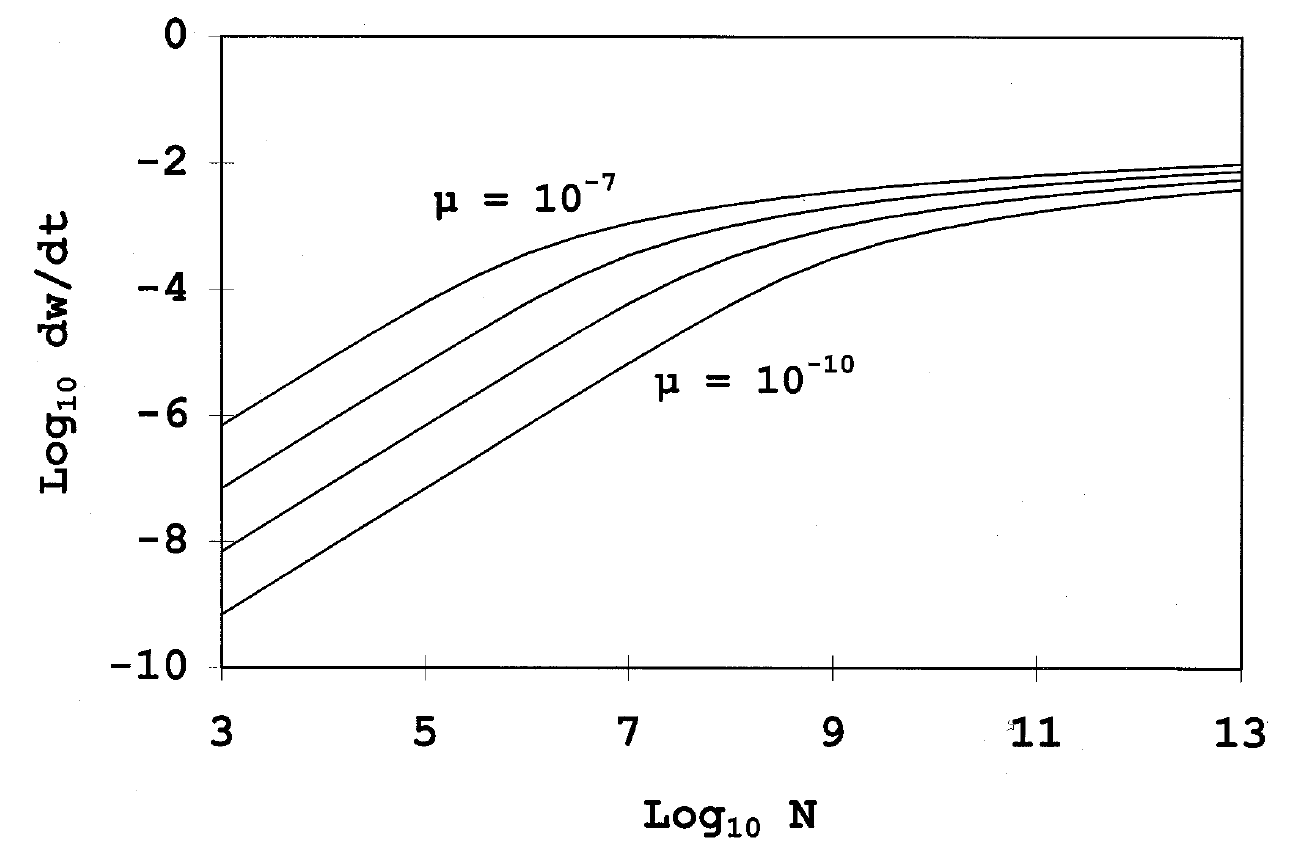
title = {The fate of competing beneficial mutations in an asexual population},
author = {Philip J. Gerrish and Richard E. Lenski},
url = {https://link.springer.com/article/10.1023%2FA%3A1017067816551},
doi = {10.1023/A:1017067816551},
isbn = {978-94-010-6193-3},
year = {1998},
date = {1998-01-01},
urldate = {1998-01-01},
journal = {Genetica},
volume = {102-103},
pages = {127-44},
abstract = {In sexual populations, beneficial mutations that occur in different lineages may be recombined into a single lineage. In asexual populations, however, clones that carry such alternative beneficial mutations compete with one another and, thereby, interfere with the expected progression of a given mutation to fixation. From theoretical exploration of such ‘clonal interference’, we have derived (1) a fixation probability for beneficial mutations, (2) an expected substitution rate, (3) an expected coefficient of selection for realized substitutions, (4) an expected rate of fitness increase, (5) the probability that a beneficial mutation transiently achieves polymorphic frequency (≥ 1%), and (6) the probability that a beneficial mutation transiently achieves majority status. Based on (2) and (3), we were able to estimate the beneficial mutation rate and the distribution of mutational effects from changes in mean fitness in an evolving \textit{E. coli} population.},
keywords = {Theory and Simulations},
pubstate = {published},
tppubtype = {article}
}
1997
Elena S F; Lenski R E
Long-term experimental evolution in Escherichia coli. VII. Mechanisms maintaining genetic variability within populations. Journal Article
Evolution, 51 (4), pp. 1058–1067, 1997, ISSN: 0014-3820.
Abstract | Links | BibTeX | Altmetric | Tags: Demography and Ecology, Fitness Trajectories, Mutation Rates, Theory and Simulations
@article{Elena1997,
title = {Long-term experimental evolution in \textit{Escherichia coli}. VII. Mechanisms maintaining genetic variability within populations.},
author = {Santiago F. Elena and Richard E. Lenski},
url = {https://onlinelibrary.wiley.com/doi/10.1111/j.1558-5646.1997.tb03953.x},
doi = {10.1111/j.1558-5646.1997.tb03953.x},
issn = {0014-3820},
year = {1997},
date = {1997-08-01},
urldate = {1997-08-01},
journal = {Evolution},
volume = {51},
number = {4},
pages = {1058--1067},
abstract = {Six replicate populations of the bacterium \textit{Escherichia coli} were propagated for more than 10,000 generations in a defined environment. We sought to quantify the variation among clones within these populations with respect to their relative fitness, and to evaluate the roles of three distinct population genetic processes in maintaining this variation. On average, a pair of clones from the same population differed from one another in their relative fitness by approximately 4%. This within-population variation was small compared with the average fitness gain relative to the common ancestor, but it was statistically significant. According to one hypothesis. The variation in fitness is transient and reflects the ongoing substitution of beneficial alleles. We used Fisher's fundamental theorem to compare the observed rate of each population's change in mean fitness with the extent of variation for fitness within that population, but we failed to discern any correspondence between these quantities. A second hypothesis supposes that the variation in fitness is maintained by recurrent deleterious mutations that give rise to a mutation-selection balance. To test this hypothesis, we made use of the fact that two of the six replicate populations had evolved mutator phenotypes, which gave them a genomic mutation rate approximately 100-fold higher than that of the other populations. There was a marginally significant correlation between a population's mutation rate and the extent of its within-population variance for fitness, but this correlation was driven by only one population (whereas two of the populations had elevated mutation rates). Under a third hypothesis, this variation is maintained by frequency-dependent selection, whereby genotypes have an advantage when they are rare relative to when they are common. In all six populations, clones were more fit, on average, when they were rare than when they were common, although the magnitude of the advantage when rare was usually small (~1% in five populations and ~45% in the other). These three hypotheses are not mutually exclusive, but frequency-dependent selection appears to be the primary force maintaining the fitness variation within these experimental populations.},
keywords = {Demography and Ecology, Fitness Trajectories, Mutation Rates, Theory and Simulations},
pubstate = {published},
tppubtype = {article}
}
1995
Johnson P A; Lenski R E; Hoppensteadt F C
Theoretical Analysis of Divergence in Mean Fitness between Initially Identical Populations Journal Article
Proceedings of the Royal Society, London, pp. 125–130, 1995.
Abstract | Links | BibTeX | Altmetric | Tags: Fitness Trajectories, Parallelism and Divergence, Theory and Simulations
@article{Johnson1995,
title = {Theoretical Analysis of Divergence in Mean Fitness between Initially Identical Populations},
author = {Paul A. Johnson and Richard E. Lenski and Frank C. Hoppensteadt},
url = {https://royalsocietypublishing.org/doi/10.1098/rspb.1995.0019},
doi = {10.1098/rspb.1995.0019},
year = {1995},
date = {1995-01-01},
urldate = {1995-01-01},
journal = {Proceedings of the Royal Society, London},
pages = {125--130},
abstract = {Initially identical populations in identical environments may subsequently diverge from one another not only via the effects of genetic drift on neutral alleles, but also by selection on beneficial alleles that arise stochastically by mutation. In the simple case of one locus with two alleles in a haploid organism, a full range of combinations of population sizes, selection pressures, mutation rates and fixation probabilities reveals two qualitatively distinct dynamics of divergence among such initially identical populations. We define a non-dimensional parameter \textit{k} that describes conditions for the occurrence of these different dynamics. One dynamic (\textit{k} > 1) occurs when beneficial mutations are sufficiently common that substitutions within the populations are essentially simultaneous; the other dynamic (\textit{k} < 1) occurs when beneficial mutations are so rare that substitutions are likely to occur as isolated events. If there are more than two alleles, or multiple loci, divergence among the populations can be sustained indefinitely if \textit{k} < 1. The parameter \textit{k} pertains to the nature of biological evolution and its tendency to be gradual or punctuated.},
keywords = {Fitness Trajectories, Parallelism and Divergence, Theory and Simulations},
pubstate = {published},
tppubtype = {article}
}