2023
Turner C B; Blount Z D; Mitchell D H; Lenski R E
Evolution of a cross-feeding interaction following a key innovation in a long-term evolution experiment with Escherichia coli Journal Article
Microbiology (Reading), 169 (8), 2023.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Demography and Ecology, Historical Contingency
@article{Turner2023,
title = { Evolution of a cross-feeding interaction following a key innovation in a long-term evolution experiment with \textit{Escherichia coli} },
author = {Caroline B. Turner and Zachary D. Blount and Daniel H. Mitchell and Richard E. Lenski},
doi = {10.1099/mic.0.001390},
year = {2023},
date = {2023-08-01},
urldate = {2023-08-01},
journal = { Microbiology (Reading)},
volume = {169},
number = {8},
abstract = {The evolution of a novel trait can profoundly change an organism's effects on its environment, which can in turn affect the further evolution of that organism and any coexisting organisms. We examine these effects and feedbacks following the evolution of a novel function in the Long-Term Evolution Experiment (LTEE) with \textit{Escherichia coli}. A characteristic feature of \textit{E. coli} is its inability to grow aerobically on citrate (Cit^{−}). Nonetheless, a Cit^{+} variant with this capacity evolved in one LTEE population after 31 000 generations. The Cit^{+} clade then coexisted stably with another clade that retained the ancestral Cit^{−} phenotype. This coexistence was shaped by the evolution of a cross-feeding relationship based on C4-dicarboxylic acids, particularly succinate, fumarate, and malate, that the Cit^{+} variants release into the medium. Both the Cit^{−} and Cit^{+} cells evolved to grow on these excreted resources. The evolution of aerobic growth on citrate thus led to a transition from an ecosystem based on a single limiting resource, glucose, to one with at least five resources that were either shared or partitioned between the two coexisting clades. Our findings show that evolutionary novelties can change environmental conditions in ways that facilitate diversity by altering ecosystem structure and the evolutionary trajectories of coexisting lineages. },
keywords = {Citrate Evolution, Demography and Ecology, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
Lenski R E
Revisiting the design of the long-term evolution experiment with Escherichia coli Journal Article
Journal of Molecular Evolution, 91 (3), pp. 241-253, 2023.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Descendant Experiments, Fitness Trajectories, Genome Evolution, Methods and Miscellaneous, Review Articles
@article{lenski2023,
title = {Revisiting the design of the long-term evolution experiment with \textit{Escherichia coli}},
author = {Richard E. Lenski},
url = {https://link.springer.com/epdf/10.1007/s00239-023-10095-3?sharing_token=zmDHuK0kbvnJBQq1k96fe_e4RwlQNchNByi7wbcMAY53KNkhv6F2YgRIeC8sZGNejxJrvlAGZWInruED5Dqdai5WeU2RAWL2PJNp0pL9QJO39B_ijCtRZcaW8jqM7PclDJfFwL_78U5zNlQYyCOsQwa1Yxha61uXUWhW-Buiq7o=},
doi = {10.1007/s00239-023-10095-3},
year = {2023},
date = {2023-06-01},
urldate = {2023-02-15},
journal = {Journal of Molecular Evolution},
volume = {91},
number = {3},
pages = {241-253},
abstract = {The long-term evolution experiment (LTEE) with \textit{Escherichia coli} began in 1988 and it continues to this day, with its 12 populations having recently reached 75,000 generations of evolution in a simple, well-controlled environment. The LTEE was designed to explore open-ended questions about the dynamics and repeatability of phenotypic and genetic evolution. Here I discuss various aspects of the LTEE’s experimental design that have enabled its stability and success, including the choices of the culture regime, growth medium, ancestral strain, and statistical replication. I also discuss some of the challenges associated with a long-running project, such as handling procedural errors (e.g., cross-contamination) and managing the expanding collection of frozen samples. The simplicity of the experimental design and procedures have supported the long-term stability of the LTEE. That stability—along with the inherent creativity of the evolutionary process and the emergence of new genomic technologies—provides a platform that has allowed talented students and collaborators to pose questions, collect data, and make discoveries that go far beyond anything I could have imagined at the start of the LTEE.},
keywords = {Citrate Evolution, Descendant Experiments, Fitness Trajectories, Genome Evolution, Methods and Miscellaneous, Review Articles},
pubstate = {published},
tppubtype = {article}
}
2020
Blount Z D; Maddamsetti R; Grant N A; Ahmed S T; Jagdish T; Baxter J A; Sommerfeld B A; Tillman A; Moore J; Slonczewski J L; Barrick J E; Lenski R E
Genomic and phenotypic evolution of Escherichia coli in a novel citrate-only resource environment Journal Article
eLife, 9 , pp. 1–64, 2020, ISSN: 2050-084X.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Demography and Ecology, Descendant Experiments, Genotypes and Phenotypes
@article{Blount2020,
title = {Genomic and phenotypic evolution of \textit{Escherichia coli} in a novel citrate-only resource environment},
author = {Zachary D. Blount and Rohan Maddamsetti and Nkrumah A. Grant and Sumaya T. Ahmed and Tanush Jagdish and Jessica A. Baxter and Brooke A. Sommerfeld and Alice Tillman and Jeremy Moore and Joan L. Slonczewski and Jeffrey E. Barrick and Richard E. Lenski},
url = {https://elifesciences.org/articles/55414},
doi = {10.7554/eLife.55414},
issn = {2050-084X},
year = {2020},
date = {2020-05-01},
urldate = {2020-05-01},
journal = {eLife},
volume = {9},
pages = {1--64},
abstract = {Evolutionary innovations allow populations to colonize new ecological niches. We previously reported that aerobic growth on citrate (Cit^{+}) evolved in an \textit{Escherichia coli} population during adaptation to a minimal glucose medium containing citrate (DM25). Cit^{+} variants can also grow in citrate-only medium (DM0), a novel environment for \textit{E. coli}. To study adaptation to this niche, we founded two sets of Cit^{+} populations and evolved them for 2500 generations in DM0 or DM25. The evolved lineages acquired numerous parallel mutations, many mediated by transposable elements. Several also evolved amplifications of regions containing the \textit{maeA} gene. Unexpectedly, some evolved populations and clones show apparent declines in fitness. We also found evidence of substantial cell death in Cit^{+} clones. Our results thus demonstrate rapid trait refinement and adaptation to the new citrate niche, while also suggesting a recalcitrant mismatch between \textit{E. coli} physiology and growth on citrate.},
keywords = {Citrate Evolution, Demography and Ecology, Descendant Experiments, Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}
2018
Bajić D; Vila J C C; Blount Z D; Sánchez A
On the deformability of an empirical fitness landscape by microbial evolution Journal Article
Proceedings of the National Academy of Sciences of the United States of America, 115 (44), pp. 11286-11291, 2018, ISSN: 0027-8424.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Demography and Ecology, Theory and Simulations
@article{nokey,
title = {On the deformability of an empirical fitness landscape by microbial evolution},
author = {Djordje Bajić and Jean C. C. Vila and Zachary D. Blount and Alvaro Sánchez},
url = {http://www.pnas.org/lookup/doi/10.1073/pnas.1808485115},
doi = {10.1073/pnas.1808485115},
issn = {0027-8424},
year = {2018},
date = {2018-10-30},
urldate = {2018-10-30},
journal = {Proceedings of the National Academy of Sciences of the United States of America},
volume = {115},
number = {44},
pages = {11286-11291},
abstract = {A fitness landscape is a map between the genotype and its reproductive success in a given environment. The topography of fitness landscapes largely governs adaptive dynamics, constraining evolutionary trajectories and the predictability of evolution. Theory suggests that this topography can be deformed by mutations that produce substantial changes to the environment. Despite its importance, the deformability of fitness landscapes has not been systematically studied beyond abstract models, and little is known about its reach and consequences in empirical systems. Here we have systematically characterized the deformability of the genome-wide metabolic fitness landscape of the bacterium \textit{Escherichia coli}. Deformability is quantified by the noncommutativity of epistatic interactions, which we experimentally demonstrate in mutant strains on the path to an evolutionary innovation. Our analysis shows that the deformation of fitness landscapes by metabolic mutations rarely affects evolutionary trajectories in the short range. However, mutations with large environmental effects produce long-range landscape deformations in distant regions of the genotype space that affect the fitness of later descendants. Our results therefore suggest that, even in situations in which mutations have strong environmental effects, fitness landscapes may retain their power to forecast evolution over small mutational distances despite the potential attenuation of that power over longer evolutionary trajectories. Our methods and results provide an avenue for integrating adaptive and eco-evolutionary dynamics with complex genetics and genomics. },
keywords = {Citrate Evolution, Demography and Ecology, Theory and Simulations},
pubstate = {published},
tppubtype = {article}
}
Leon D; D'Alton S; Quandt E M; Barrick J E
Innovation in an E. coli evolution experiment is contingent on maintaining adaptive potential until competition subsides Journal Article
PLOS Genetics, 14 (4), pp. e1007348, 2018, ISSN: 1553-7404.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Demography and Ecology, Historical Contingency
@article{nokey,
title = {Innovation in an \textit{E. coli} evolution experiment is contingent on maintaining adaptive potential until competition subsides},
author = {Dacia Leon and Simon D'Alton and Erik M. Quandt and Jeffrey E. Barrick},
url = {https://dx.plos.org/10.1371/journal.pgen.1007348},
doi = {10.1371/journal.pgen.1007348},
issn = {1553-7404},
year = {2018},
date = {2018-04-12},
urldate = {2018-04-12},
journal = {PLOS Genetics},
volume = {14},
number = {4},
pages = {e1007348},
abstract = {Key innovations are disruptive evolutionary events that enable a species to escape constraints and rapidly diversify. After 15 years of the Lenski long-term evolution experiment with \textit{Escherichia coli}, cells in one of the twelve populations evolved the ability to utilize citrate, an abundant but previously untapped carbon source in the environment. Descendants of these cells became dominant in the population and subsequently diversified as a consequence of invading this vacant niche. Mutations responsible for the appearance of rudimentary citrate utilization and for refining this ability have been characterized. However, the complete nature of the genetic and/or ecological events that set the stage for this key innovation is unknown. In particular, it is unclear why it took so long for citrate utilization to evolve and why it still has evolved in only one of the twelve \textit{E. coli} populations after 30 years of the Lenski experiment. In this study, we recapitulated the initial mutation needed to evolve citrate utilization in strains isolated from throughout the first 31,500 generations of the history of this population. We found that there was already a slight fitness benefit for this mutation in the original ancestor of the evolution experiment and in other early isolates. However, evolution of citrate utilization was blocked at this point due to competition with other mutations that improved fitness in the original niche. Subsequently, an anti-potentiated genetic background evolved in which it was deleterious to evolve rudimentary citrate utilization. Only later, after further mutations accumulated that restored the benefit of this first-step mutation and the overall rate of adaptation in the population slowed, was citrate utilization likely to evolve. Thus, intense competition and the types of mutations that it favors can lead to short-sighted evolutionary trajectories that hide a stepping stone needed to access a key innovation from many future generations.},
keywords = {Citrate Evolution, Demography and Ecology, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
2015
Turner C B; Blount Z D; Lenski R E
Replaying Evolution to Test the Cause of Extinction of One Ecotype in an Experimentally Evolved Population Journal Article
PLOS ONE, 10 (11), pp. e0142050, 2015, ISSN: 1932-6203.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Demography and Ecology, Historical Contingency
@article{Turner2015,
title = {Replaying Evolution to Test the Cause of Extinction of One Ecotype in an Experimentally Evolved Population},
author = {Caroline B. Turner and Zachary D. Blount and Richard E. Lenski},
editor = {Frederick M. Cohan},
url = {https://dx.plos.org/10.1371/journal.pone.0142050},
doi = {10.1371/journal.pone.0142050},
issn = {1932-6203},
year = {2015},
date = {2015-11-01},
urldate = {2015-11-01},
journal = {PLOS ONE},
volume = {10},
number = {11},
pages = {e0142050},
abstract = {In a long-term evolution experiment with \textit{Escherichia coli}, bacteria in one of twelve populations evolved the ability to consume citrate, a previously unexploited resource in a glucoselimited medium. This innovation led to the frequency-dependent coexistence of citrate-consuming (Cit+) and non-consuming (Cit-) ecotypes, with Cit-bacteria persisting on the exogenously supplied glucose as well as other carbon molecules released by the Cit+ bacteria. After more than 10,000 generations of coexistence, however, the Cit-lineage went extinct; cells with the Cit-phenotype dropped to levels below detection, and the Cit-clade could not be detected by molecular assays based on its unique genotype. We hypothesized that this extinction was a deterministic outcome of evolutionary change within the population, specifically the appearance of a more-fit Cit+ ecotype that competitively excluded the Cit-ecotype. We tested this hypothesis by re-evolving the population from a frozen population sample taken within 500 generations of the extinction and from another sample taken several thousand generations earlier, in each case for 500 generations and with 20-fold replication. To our surprise, the Cit-type did not go extinct in any of these replays, and Cit-cells also persisted in a single replicate that was propagated for 2,500 generations. Even more unexpectedly, we showed that the Cit-ecotype could reinvade the Cit+ population after its extinction. Taken together, these results indicate that the extinction of the Cit-ecotype was not a deterministic outcome driven by competitive exclusion by the Cit+ ecotype. The extinction also cannot be explained by demographic stochasticity alone, as the population size of the Cit-ecotype should have been many thousands of cells even during the daily transfer events. Instead, we infer that the extinction must have been caused by a rare chance event in which some aspect of the experimental conditions was inadvertently perturbed.},
keywords = {Citrate Evolution, Demography and Ecology, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
Quandt E M; Gollihar J; Blount Z D; Ellington A D; Georgiou G; Barrick J E
Fine-tuning citrate synthase flux potentiates and refines metabolic innovation in the Lenski evolution experiment. Journal Article
eLife, 4 (October), pp. e09696, 2015, ISSN: 2050-084X.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Genotypes and Phenotypes, Historical Contingency
@article{Quandt2015,
title = {Fine-tuning citrate synthase flux potentiates and refines metabolic innovation in the Lenski evolution experiment.},
author = {Erik M. Quandt and Jimmy Gollihar and Zachary D. Blount and Andrew D. Ellington and George Georgiou and Jeffrey E. Barrick},
url = {http://www.ncbi.nlm.nih.gov/pubmed/26465114
http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=PMC4718724},
doi = {10.7554/eLife.09696},
issn = {2050-084X},
year = {2015},
date = {2015-10-01},
urldate = {2015-10-01},
journal = {eLife},
volume = {4},
number = {October},
pages = {e09696},
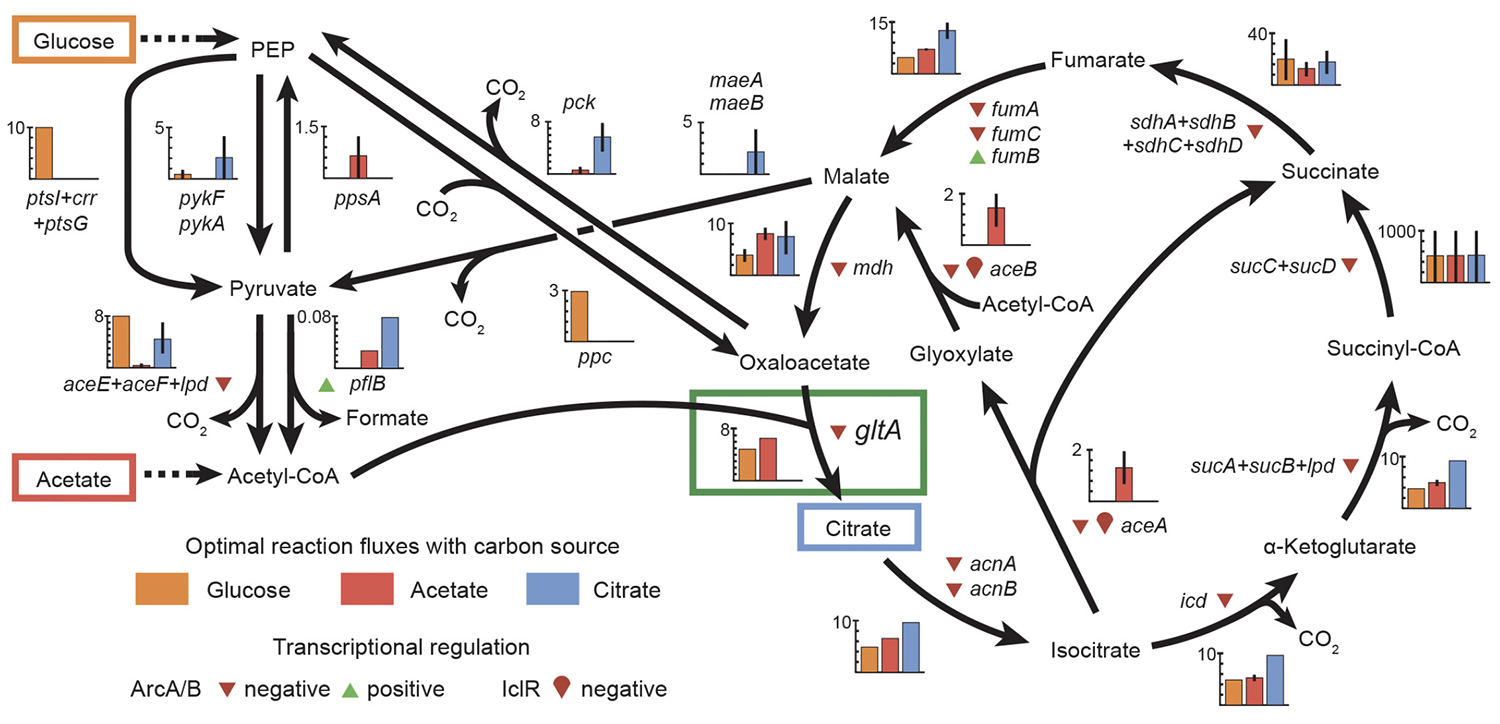
abstract = {Evolutionary innovations that enable organisms to colonize new ecological niches are rare compared to gradual evolutionary changes in existing traits. We discovered that key mutations in the \textit{gltA} gene, which encodes citrate synthase (CS), occurred both before and after \textit{Escherichia coli} gained the ability to grow aerobically on citrate (Cit(+) phenotype) during the Lenski long-term evolution experiment. The first \textit{gltA} mutation, which increases CS activity by disrupting NADH-inhibition of this enzyme, is beneficial for growth on the acetate and contributed to preserving the rudimentary Cit(+) trait from extinction when it first evolved. However, after Cit(+) was refined by further mutations, this potentiating \textit{gltA} mutation became deleterious to fitness. A second wave of beneficial \textit{gltA} mutations then evolved that reduced CS activity to below the ancestral level. Thus, dynamic reorganization of central metabolism made colonizing this new nutrient niche contingent on both co-opting and overcoming a history of prior adaptation.},
keywords = {Citrate Evolution, Genotypes and Phenotypes, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
2014
Quandt E M; Deatherage D E; Ellington A D; Georgiou G; Barrick J E
Recursive genomewide recombination and sequencing reveals a key refinement step in the evolution of a metabolic innovation in Escherichia coli Journal Article
Proceedings of the National Academy of Sciences of the United States of America, 111 (6), pp. 2217–2222, 2014, ISSN: 0027-8424.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Genotypes and Phenotypes, Methods and Miscellaneous
@article{Quandt2014,
title = {Recursive genomewide recombination and sequencing reveals a key refinement step in the evolution of a metabolic innovation in \textit{Escherichia coli}},
author = {Erik M. Quandt and Daniel E. Deatherage and Andrew D. Ellington and George Georgiou and Jeffrey E. Barrick},
url = {http://www.pnas.org/lookup/doi/10.1073/pnas.1314561111},
doi = {10.1073/pnas.1314561111},
issn = {0027-8424},
year = {2014},
date = {2014-02-01},
urldate = {2014-02-01},
journal = {Proceedings of the National Academy of Sciences of the United States of America},
volume = {111},
number = {6},
pages = {2217--2222},
abstract = {Evolutionary innovations often arise from complex genetic and ecological interactions, which can make it challenging to understand retrospectively how a novel trait arose. In a long-term experiment, \textit{Escherichia coli} gained the ability to use abundant citrate (Cit+) in the growth medium after ~31,500 generations of evolution. Exploiting this previously untapped resource was highly beneficial: later Cit+ variants achieve a much higher population density in this environment. All Cit+ individuals share a mutation that activates aerobic expression of the \textit{citT} citrate transporter, but this mutation confers only an extremely weak Cit+ phenotype on its own. To determine which of the other >70 mutations in early Cit+ clones were needed to take full advantage of citrate, we developed a recursive genomewide recombination and sequencing method (REGRES) and performed genetic backcrosses to purge mutations not required for Cit+ from an evolved strain. We discovered a mutation that increased expression of the \textit{dctA} C4-dicarboxylate transporter greatly enhanced the Cit+ phenotype after it evolved. Surprisingly, strains containing just the \textit{citT} and \textit{dctA} mutations fully use citrate, indicating that earlier mutations thought to have potentiated the initial evolution of Cit+ are not required for expression of the refined version of this trait. Instead, this metabolic innovation may be contingent on a genetic background, and possibly ecological context, that enabled \textit{citT} mutants to persist among competitors long enough to obtain \textit{dctA} or equivalent mutations that conferred an overwhelming advantage. More generally, refinement of an emergent trait from a rudimentary form may be crucial to its evolutionary success.},
keywords = {Citrate Evolution, Genotypes and Phenotypes, Methods and Miscellaneous},
pubstate = {published},
tppubtype = {article}
}
2012
Leiby N; Harcombe W R; Marx C J
BMC Evolutionary Biology, 12 (1), pp. 151, 2012, ISSN: 1471-2148.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Correlated Responses
@article{Leiby2012,
title = {Multiple long-term, experimentally-evolved populations of \textit{Escherichia coli} acquire dependence upon citrate as an iron chelator for optimal growth on glucose},
author = {Nicholas Leiby and William R. Harcombe and Christopher J. Marx},
url = {https://bmcevolbiol.biomedcentral.com/articles/10.1186/1471-2148-12-151},
doi = {10.1186/1471-2148-12-151},
issn = {1471-2148},
year = {2012},
date = {2012-12-01},
urldate = {2012-12-01},
journal = {BMC Evolutionary Biology},
volume = {12},
number = {1},
pages = {151},
abstract = {Background
Specialization for ecological niches is a balance of evolutionary adaptation and its accompanying tradeoffs. Here we focus on the Lenski Long-Term Evolution Experiment, which has maintained cultures of \textit{Escherichia coli} in the same defined seasonal environment for 50,000 generations. Over this time, much adaptation and specialization to the environment has occurred. The presence of citrate in the growth media selected one lineage to gain the novel ability to utilize citrate as a carbon source after 31,000 generations. Here we test whether other strains have specialized to rely on citrate after 50,000 generations.
Results
We show that in addition to the citrate-catabolizing strain, three other lineages evolving in parallel have acquired a dependence on citrate for optimal growth on glucose. None of these strains were stimulated indirectly by the sodium present in disodium citrate, nor exhibited even partial utilization of citrate as a carbon source. Instead, all three of these citrate-stimulated populations appear to rely on it as a chelator of iron.
Conclusions
The strains we examine here have evolved specialization to their environment through apparent loss of function. Our results are most consistent with the accumulation of mutations in iron transport genes that were obviated by abundant citrate. The results present another example where a subtle decision in the design of an evolution experiment led to unexpected evolutionary outcomes.},
keywords = {Citrate Evolution, Correlated Responses},
pubstate = {published},
tppubtype = {article}
}
Specialization for ecological niches is a balance of evolutionary adaptation and its accompanying tradeoffs. Here we focus on the Lenski Long-Term Evolution Experiment, which has maintained cultures of Escherichia coli in the same defined seasonal environment for 50,000 generations. Over this time, much adaptation and specialization to the environment has occurred. The presence of citrate in the growth media selected one lineage to gain the novel ability to utilize citrate as a carbon source after 31,000 generations. Here we test whether other strains have specialized to rely on citrate after 50,000 generations.
Results
We show that in addition to the citrate-catabolizing strain, three other lineages evolving in parallel have acquired a dependence on citrate for optimal growth on glucose. None of these strains were stimulated indirectly by the sodium present in disodium citrate, nor exhibited even partial utilization of citrate as a carbon source. Instead, all three of these citrate-stimulated populations appear to rely on it as a chelator of iron.
Conclusions
The strains we examine here have evolved specialization to their environment through apparent loss of function. Our results are most consistent with the accumulation of mutations in iron transport genes that were obviated by abundant citrate. The results present another example where a subtle decision in the design of an evolution experiment led to unexpected evolutionary outcomes.
Blount Z D; Barrick J E; Davidson C J; Lenski R E
Genomic analysis of a key innovation in an experimental Escherichia coli population Journal Article
Nature, 489 (7417), pp. 513–518, 2012, ISSN: 0028-0836.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Genome Evolution, Genotypes and Phenotypes, Historical Contingency, Mutation Rates
@article{Blount2012,
title = {Genomic analysis of a key innovation in an experimental \emph{Escherichia coli} population},
author = {Zachary D. Blount and Jeffrey E. Barrick and Carla J. Davidson and Richard E. Lenski},
url = {http://www.nature.com/articles/nature11514},
doi = {10.1038/nature11514},
issn = {0028-0836},
year = {2012},
date = {2012-09-01},
urldate = {2012-09-01},
journal = {Nature},
volume = {489},
number = {7417},
pages = {513--518},
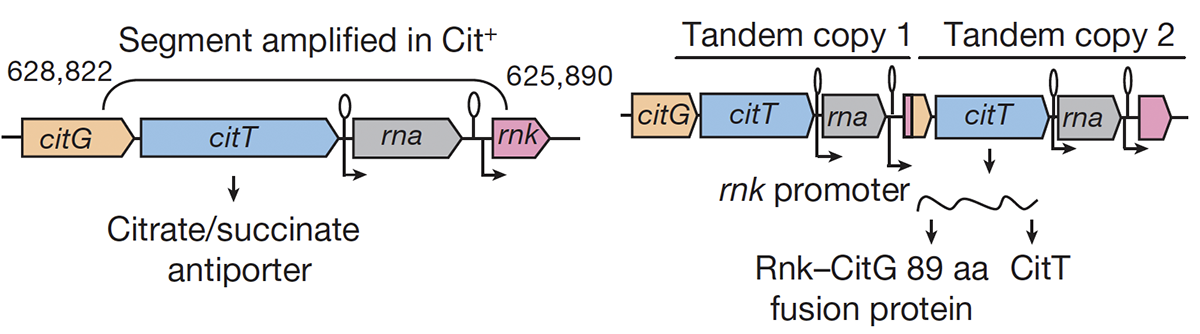
abstract = {Evolutionary novelties have been important in the history of life, but their origins are usually difficult to examine in detail. We previously described the evolution of a novel trait, aerobic citrate utilization (Cit +), in an experimental population of \textit{Escherichia coli}. Here we analyse genome sequences to investigate the history and genetic basis of this trait. At least three distinct clades coexisted for more than 10,000 generations before its emergence. The Cit + trait originated in one clade by a tandem duplication that captured an aerobically expressed promoter for the expression of a previously silent citrate transporter. The clades varied in their propensity to evolve this novel trait, although genotypes able to do so existed in all three clades, implying that multiple potentiating mutations arose during the population's history. Our findings illustrate the importance of promoter capture and altered gene regulation in mediating the exaptation events that often underlie evolutionary innovations.},
keywords = {Citrate Evolution, Genome Evolution, Genotypes and Phenotypes, Historical Contingency, Mutation Rates},
pubstate = {published},
tppubtype = {article}
}
2008
Blount Z D; Borland C Z; Lenski R E
Historical contingency and the evolution of a key innovation in an experimental population of Escherichia coli Journal Article
Proceedings of the National Academy of Sciences of the United States of America, 105 (23), pp. 7899–7906, 2008, ISSN: 0027-8424.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Demography and Ecology, Historical Contingency
@article{Blount2008,
title = {Historical contingency and the evolution of a key innovation in an experimental population of \emph{Escherichia coli}},
author = {Zachary D. Blount and Christina Z. Borland and Richard E. Lenski},
url = {http://www.pnas.org/cgi/doi/10.1073/pnas.0803151105},
doi = {10.1073/pnas.0803151105},
issn = {0027-8424},
year = {2008},
date = {2008-06-01},
urldate = {2008-06-01},
journal = {Proceedings of the National Academy of Sciences of the United States of America},
volume = {105},
number = {23},
pages = {7899--7906},
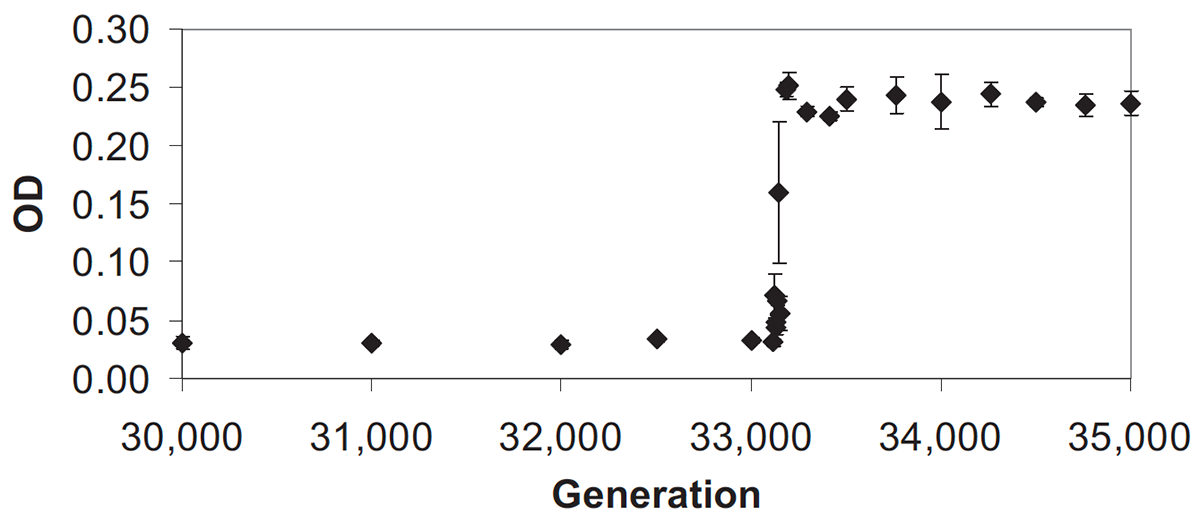
abstract = {The role of historical contingency in evolution has been much debated, but rarely tested. Twelve initially identical populations of \textit{Escherichia coli} were founded in 1988 to investigate this issue. They have since evolved in a glucose-limited medium that also contains citrate, which \textit{E. coli} cannot use as a carbon source under oxic conditions. No population evolved the capacity to exploit citrate for >30,000 generations, although each population tested billions of mutations. A citrate-using (Cit^{+}) variant finally evolved in one population by 31,500 generations, causing an increase in population size and diversity. The long-delayed and unique evolution of this function might indicate the involvement of some extremely rare mutation. Alternately, it may involve an ordinary mutation, but one whose physical occurrence or phenotypic expression is contingent on prior mutations in that population. We tested these hypotheses in experiments that “replayed” evolution from different points in that population's history. We observed no Cit^{+} mutants among 8.4 × 10^{12} ancestral cells, nor among 9 × 10^{12} cells from 60 clones sampled in the first 15,000 generations. However, we observed a significantly greater tendency for later clones to evolve Cit^{+}, indicating that some potentiating mutation arose by 20,000 generations. This potentiating change increased the mutation rate to Cit^{+} but did not cause generalized hypermutability. Thus, the evolution of this phenotype was contingent on the particular history of that population. More generally, we suggest that historical contingency is especially important when it facilitates the evolution of key innovations that are not easily evolved by gradual, cumulative selection.},
keywords = {Citrate Evolution, Demography and Ecology, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}