2024
Ascensao J A; Denk J; Lok K; Yu Q; Wetmore K M; Hallatschek O
Rediversification following ecotype isolation reveals hidden adaptive potential Journal Article
Current Biology, 34 (4), pp. 855–867, 2024, ISSN: 0960-9822.
Abstract | Links | BibTeX | Altmetric | Tags: Demography and Ecology, Genotypes and Phenotypes, Historical Contingency
@article{Ascensao2024,
title = {Rediversification following ecotype isolation reveals hidden adaptive potential},
author = {Joao A. Ascensao and Jonas Denk and Kristen Lok and QinQin Yu and Kelly M. Wetmore and Oskar Hallatschek},
url = {https://www.sciencedirect.com/science/article/pii/S0960982224000290},
doi = {10.1016/j.cub.2024.01.029},
issn = {0960-9822},
year = {2024},
date = {2024-02-01},
urldate = {2024-02-01},
journal = {Current Biology},
volume = {34},
number = {4},
pages = {855--867},
abstract = {Microbial communities play a critical role in ecological processes, and their diversity is key to their functioning. However, little is known about whether communities can regenerate ecological diversity following ecotype removal or extinction and how the rediversified communities would compare to the original ones. Here, we show that simple two-ecotype communities from the \textit{E. coli} long-term evolution experiment (LTEE) consistently rediversified into two ecotypes following the isolation of one of the ecotypes, coexisting via negative frequency-dependent selection. Communities separated by more than 30,000 generations of evolutionary time rediversify in similar ways. The rediversified ecotype appears to share a number of growth traits with the ecotype it replaces. However, the rediversified community is also different from the original community in ways relevant to the mechanism of ecotype coexistence—for example, in stationary phase response and survival. We found substantial variation in the transcriptional states between the two original ecotypes, whereas the differences within the rediversified community were comparatively smaller, although the rediversified community showed unique patterns of differential expression. Our results suggest that evolution may leave room for alternative diversification processes even in a maximally reduced community of only two strains. We hypothesize that the presence of alternative evolutionary pathways may be even more pronounced in communities of many species where there are even more potential niches, highlighting an important role for perturbations, such as species removal, in evolving ecological communities.},
keywords = {Demography and Ecology, Genotypes and Phenotypes, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
2023
Turner C B; Blount Z D; Mitchell D H; Lenski R E
Evolution of a cross-feeding interaction following a key innovation in a long-term evolution experiment with Escherichia coli Journal Article
Microbiology (Reading), 169 (8), 2023.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Demography and Ecology, Historical Contingency
@article{Turner2023,
title = { Evolution of a cross-feeding interaction following a key innovation in a long-term evolution experiment with \textit{Escherichia coli} },
author = {Caroline B. Turner and Zachary D. Blount and Daniel H. Mitchell and Richard E. Lenski},
doi = {10.1099/mic.0.001390},
year = {2023},
date = {2023-08-01},
urldate = {2023-08-01},
journal = { Microbiology (Reading)},
volume = {169},
number = {8},
abstract = {The evolution of a novel trait can profoundly change an organism's effects on its environment, which can in turn affect the further evolution of that organism and any coexisting organisms. We examine these effects and feedbacks following the evolution of a novel function in the Long-Term Evolution Experiment (LTEE) with \textit{Escherichia coli}. A characteristic feature of \textit{E. coli} is its inability to grow aerobically on citrate (Cit^{−}). Nonetheless, a Cit^{+} variant with this capacity evolved in one LTEE population after 31 000 generations. The Cit^{+} clade then coexisted stably with another clade that retained the ancestral Cit^{−} phenotype. This coexistence was shaped by the evolution of a cross-feeding relationship based on C4-dicarboxylic acids, particularly succinate, fumarate, and malate, that the Cit^{+} variants release into the medium. Both the Cit^{−} and Cit^{+} cells evolved to grow on these excreted resources. The evolution of aerobic growth on citrate thus led to a transition from an ecosystem based on a single limiting resource, glucose, to one with at least five resources that were either shared or partitioned between the two coexisting clades. Our findings show that evolutionary novelties can change environmental conditions in ways that facilitate diversity by altering ecosystem structure and the evolutionary trajectories of coexisting lineages. },
keywords = {Citrate Evolution, Demography and Ecology, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
2022
Izutsu M; Lenski R E
Experimental test of the contributions of initial variation and new mutations to adaptive evolution in a novel environment Journal Article
Frontiers in Ecology and Evolution, 10 , pp. 958406, 2022, ISSN: 2296-701X.
Abstract | Links | BibTeX | Altmetric | Tags: Descendant Experiments, Historical Contingency, Parallelism and Divergence
@article{izutsu2022,
title = {Experimental test of the contributions of initial variation and new mutations to adaptive evolution in a novel environment},
author = {Minako Izutsu and Richard E. Lenski},
url = {https://www.frontiersin.org/articles/10.3389/fevo.2022.958406/full},
doi = {10.3389/fevo.2022.958406},
issn = {2296-701X},
year = {2022},
date = {2022-10-06},
urldate = {2022-10-01},
journal = {Frontiers in Ecology and Evolution},
volume = {10},
pages = {958406},
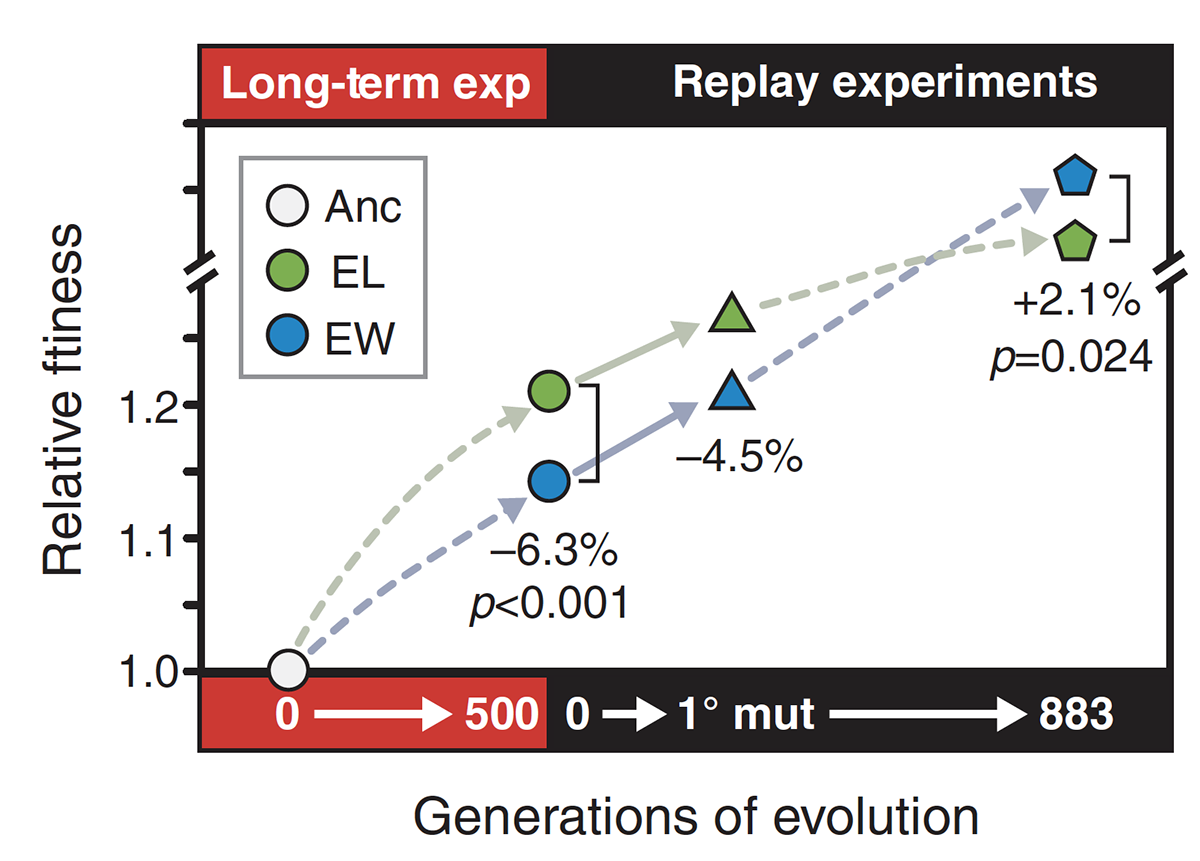
abstract = {Experimental evolution is an approach that allows researchers to study organisms as they evolve in controlled environments. Despite the growing popularity of this approach, there are conceptual gaps among projects that use different experimental designs. One such gap concerns the contributions to adaptation of genetic variation present at the start of an experiment and that of new mutations that arise during an experiment. The primary source of genetic variation has historically depended largely on the study organisms. In the long-term evolution experiment (LTEE) using \textit{Escherichia coli}, for example, each population started from a single haploid cell, and therefore, adaptation depended entirely on new mutations. Most other microbial evolution experiments have followed the same strategy. By contrast, evolution experiments using multicellular, sexually reproducing organisms typically start with preexisting variation that fuels the response to selection. New mutations may also come into play in later generations of these experiments, but it is generally difficult to quantify their contribution in these studies. Here, we performed an experiment using \textit{E. coli} to compare the contributions of initial genetic variation and new mutations to adaptation in a new environment. Our experiment had four treatments that varied in their starting diversity, with 18 populations in each treatment. One treatment depended entirely on new mutations, while the other three began with mixtures of clones, whole-population samples, or mixtures of whole-population samples from the LTEE. We tracked a genetic marker associated with different founders in two treatments. These data revealed significant variation in fitness among the founders, and that variation impacted evolution in the early generations of our experiment. However, there were no differences in fitness among the treatments after 500 or 2,000 generations in the new environment, despite the variation in fitness among the founders. These results indicate that new mutations quickly dominated, and eventually they contributed more to adaptation than did the initial variation. Our study thus shows that preexisting genetic variation can have a strong impact on early evolution in a new environment, but new beneficial mutations may contribute more to later evolution and can even drive some initially beneficial variants to extinction.},
keywords = {Descendant Experiments, Historical Contingency, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
Smith C E; Smith A N H; Cooper T F; Moore F B -G
Fitness of evolving bacterial populations is contingent on deep and shallow history but only shallow history creates predictable patterns Journal Article
Proceedings of the Royal Society B: Biological Sciences, 289 (1982), pp. 20221292, 2022.
Abstract | Links | BibTeX | Altmetric | Tags: Descendant Experiments, Fitness Trajectories, Genotypes and Phenotypes, Historical Contingency
@article{smith2022,
title = {Fitness of evolving bacterial populations is contingent on deep and shallow history but only shallow history creates predictable patterns},
author = {Chelsea E. Smith AND Adam N. H. Smith AND Tim F. Cooper AND Francisco B.-G. Moore},
url = {https://royalsocietypublishing.org/doi/10.1098/rspb.2022.1292},
doi = {10.1098/rspb.2022.1292},
year = {2022},
date = {2022-09-14},
journal = {Proceedings of the Royal Society B: Biological Sciences},
volume = {289},
number = {1982},
pages = {20221292},
abstract = {Long-term evolution experiments have tested the importance of genetic and environmental factors in influencing evolutionary outcomes. Differences in phylogenetic history, recent adaptation to distinct environments and chance events, all influence the fitness of a population. However, the interplay of these factors on a population's evolutionary potential remains relatively unexplored. We tracked the outcome of 2000 generations of evolution of four natural isolates of \textit{Escherichia coli} bacteria that were engineered to also create differences in shallow history by adding previously identified mutations selected in a separate long-term experiment. Replicate populations started from each progenitor evolved in four environments. We found that deep and shallow phylogenetic histories both contributed significantly to differences in evolved fitness, though by different amounts in different selection environments. With one exception, chance effects were not significant. Whereas the effect of deep history did not follow any detectable pattern, effects of shallow history followed a pattern of diminishing returns whereby fitter ancestors had smaller fitness increases. These results are consistent with adaptive evolution being contingent on the interaction of several evolutionary forces but demonstrate that the nature of these interactions is not fixed and may not be predictable even when the role of chance is small.},
keywords = {Descendant Experiments, Fitness Trajectories, Genotypes and Phenotypes, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
Jordan J A; Lenski R E; Card K J
Idiosyncratic fitness costs of ampicillin-resistant mutants derived from a long-term experiment with Escherichia coli Journal Article
Antibiotics, 11 (3), pp. 347, 2022, ISSN: 2079-6382.
Abstract | Links | BibTeX | Altmetric | Tags: Descendant Experiments, Genome Evolution, Historical Contingency
@article{Jordan2022.02.06.479266,
title = {Idiosyncratic fitness costs of ampicillin-resistant mutants derived from a long-term experiment with \textit{Escherichia coli}},
author = {Jalin A. Jordan and Richard E. Lenski and Kyle J. Card},
url = {https://www.biorxiv.org/content/10.1101/2022.02.06.479266v1},
doi = {10.3390/antibiotics11030347},
issn = {2079-6382},
year = {2022},
date = {2022-03-13},
urldate = {2022-03-13},
journal = {Antibiotics},
volume = {11},
number = {3},
pages = {347},
publisher = {Cold Spring Harbor Laboratory},
abstract = {Antibiotic resistance is a growing concern that has prompted a renewed focus on drug discovery, stewardship, and evolutionary studies of the patterns and processes that underlie this phenomenon. A resistant strain’s competitive fitness relative to its sensitive counterparts in the absence of drug can impact its spread and persistence in both clinical and community settings. In a prior study, we examined the fitness of tetracycline-resistant clones that evolved from five different Escherichia coli genotypes, which had diverged during a long-term evolution experiment. In this study, we build on that work to examine whether ampicillin-resistant mutants are also less fit in the absence of the drug than their sensitive parents, and whether the cost of resistance is constant or variable among independently derived lines. Like the tetracycline-resistant lines, the ampicillin-resistant mutants were often less fit than their sensitive parents, with significant variation in the fitness costs among the mutants. This variation was not associated with the level of resistance conferred by the mutations, nor did it vary across the different parental backgrounds. In our earlier study, some of the variation in fitness costs associated with tetracycline resistance was explained by the effects of different mutations affecting the same cellular pathway and even the same gene. In contrast, the variance among the ampicillin-resistant mutants was associated with different sets of target genes. About half of the resistant clones suffered large fitness deficits, and their mutations impacted major outer-membrane proteins or subunits of RNA polymerases. The other mutants experienced little or no fitness costs and with, one exception, they had mutations affecting other genes and functions. Our findings underscore the importance of comparative studies on the evolution of antibiotic resistance, and they highlight the nuanced processes that shape these phenotypes.},
keywords = {Descendant Experiments, Genome Evolution, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
2021
van Raay K; Stolyar S; Sevigny J; Draghi J; Lenski R E; Marx C J; Kerr B; Zaman L
Evolution with private resources reverses some changes from long-term evolution with public resources Unpublished
bioRxiv, 2021.
Abstract | Links | BibTeX | Altmetric | Tags: Cell Morphology, Correlated Responses, Demography and Ecology, Descendant Experiments, Genotypes and Phenotypes, Historical Contingency, Methods and Miscellaneous
@unpublished{Raay2021,
title = {Evolution with private resources reverses some changes from long-term evolution with public resources},
author = {Katrina {van Raay} and Sergey Stolyar and Jordana Sevigny and Jeremy Draghi and Richard E. Lenski and Christopher J. Marx and Benjamin Kerr and Luis Zaman},
url = {https://www.biorxiv.org/content/10.1101/2021.07.11.451942v1},
doi = {https://doi.org/10.1101/2021.07.11.451942},
year = {2021},
date = {2021-07-12},
urldate = {2021-07-12},
journal = {bioRxiv},
pages = {2021.07.11.451942},
abstract = {A population under selection to improve one trait may evolve a sub-optimal state for another trait due to tradeoffs and other evolutionary constraints. How this evolution affects the capacity of a population to adapt when conditions change to favor the second trait is an open question. We investigated this question using isolates from a lineage spanning 60,000 generations of the Long-Term Evolution Experiment (LTEE) with \textit{Escherichia coli}, where cells have access to a shared pool of resources, and have evolved increased competitive ability and a concomitant reduction in numerical yield. Using media-in oil emulsions we shifted the focus of selection to numerical yield, where cells grew in isolated patches with private resources. We found that the time spent evolving under shared resources did not affect the ability to re-evolve toward higher numerical yield. The evolution of numerical yield commonly occurred through mutations in the phosphoenolpyruvate phosphotransferase system. These mutants exhibit slower uptake of glucose, making them poorer competitors for public resources, and produce smaller cells that release less carbon as overflow metabolites. Our results demonstrate that mutations that were not part of adaptation under one selective regime may enable access to ancestral phenotypes when selection changes to favor evolutionary reversion. },
howpublished = {bioRxiv},
keywords = {Cell Morphology, Correlated Responses, Demography and Ecology, Descendant Experiments, Genotypes and Phenotypes, Historical Contingency, Methods and Miscellaneous},
pubstate = {published},
tppubtype = {unpublished}
}
Card K J; Thomas M D; Graves J L; Barrick J E; Lenski R E
Genomic evolution of antibiotic resistance is contingent on genetic background following a long-term experiment with Escherichia coli Journal Article
Proceedings of the National Academy of Sciences of the United States of America, 118 (5), pp. e2016886118, 2021, ISSN: 0027-8424.
Abstract | Links | BibTeX | Altmetric | Tags: Descendant Experiments, Genome Evolution, Historical Contingency
@article{Card2021,
title = {Genomic evolution of antibiotic resistance is contingent on genetic background following a long-term experiment with \textit{Escherichia coli}},
author = {Kyle J. Card and Misty D. Thomas and Joseph L. Graves and Jeffrey E. Barrick and Richard E. Lenski},
url = {http://www.pnas.org/lookup/doi/10.1073/pnas.2016886118},
doi = {10.1073/pnas.2016886118},
issn = {0027-8424},
year = {2021},
date = {2021-02-01},
urldate = {2021-02-01},
journal = {Proceedings of the National Academy of Sciences of the United States of America},
volume = {118},
number = {5},
pages = {e2016886118},
abstract = {Antibiotic resistance is a growing health concern. Efforts to control resistance would benefit from an improved ability to forecast when and how it will evolve. Epistatic interactions between mutations can promote divergent evolutionary trajectories, which complicates our ability to predict evolution. We recently showed that differences between genetic backgrounds can lead to idiosyncratic responses in the evolvability of phenotypic resistance, even among closely related \textit{Escherichia coli} strains. In this study, we examined whether a strain's genetic background also influences the genotypic evolution of resistance. Do lineages founded by different genotypes take parallel or divergent mutational paths to achieve their evolved resistance states? We addressed this question by sequencing the complete genomes of antibiotic-resistant clones that evolved from several different genetic starting points during our earlier experiments. We first validated our statistical approach by quantifying the specificity of genomic evolution with respect to antibiotic treatment. As expected, mutations in particular genes were strongly associated with each drug. Then, we determined that replicate lines evolved from the same founding genotypes had more parallel mutations at the gene level than lines evolved from different founding genotypes, although these effects were more subtle than those showing antibiotic specificity. Taken together with our previous work, we conclude that historical contingency can alter both genotypic and phenotypic pathways to antibiotic resistance.},
keywords = {Descendant Experiments, Genome Evolution, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
2019
Card K J; LaBar T; Gomez J B; Lenski R E
Historical contingency in the evolution of antibiotic resistance after decades of relaxed selection Journal Article
PLOS Biology, 17 (10), pp. e3000397, 2019, ISSN: 1545-7885.
Abstract | Links | BibTeX | Altmetric | Tags: Correlated Responses, Descendant Experiments, Historical Contingency
@article{nokey,
title = {Historical contingency in the evolution of antibiotic resistance after decades of relaxed selection},
author = {Kyle J. Card and Thomas LaBar and Jasper B. Gomez and Richard E. Lenski},
url = {https://dx.plos.org/10.1371/journal.pbio.3000397},
doi = {10.1371/journal.pbio.3000397},
issn = {1545-7885},
year = {2019},
date = {2019-10-23},
urldate = {2019-10-23},
journal = {PLOS Biology},
volume = {17},
number = {10},
pages = {e3000397},
abstract = {Populations often encounter changed environments that remove selection for the maintenance of particular phenotypic traits. The resulting genetic decay of those traits under relaxed selection reduces an organism’s fitness in its prior environment. However, whether and how such decay alters the subsequent evolvability of a population upon restoration of selection for a previously diminished trait is not well understood. We addressed this question using \textit{Escherichia coli} strains from the long-term evolution experiment (LTEE) that independently evolved for multiple decades in the absence of antibiotics. We first confirmed that these derived strains are typically more sensitive to various antibiotics than their common ancestor. We then subjected the ancestral and derived strains to various concentrations of these drugs to examine their potential to evolve increased resistance. We found that evolvability was idiosyncratic with respect to initial genotype; that is, the derived strains did not generally compensate for their greater susceptibility by “catching up” to the resistance level of the ancestor. Instead, the capacity to evolve increased resistance was constrained in some backgrounds, implying that evolvability depended upon prior mutations in a historically contingent fashion. We further subjected a time series of clones from one LTEE population to tetracycline and determined that an evolutionary constraint arose early in that population, corroborating the role of contingency. In summary, relaxed selection not only can drive populations to increased antibiotic susceptibility, but it can also affect the subsequent evolvability of antibiotic resistance in an unpredictable manner. This conclusion has potential implications for public health, and it underscores the need to consider the genetic context of pathogens when designing drug-treatment strategies.},
keywords = {Correlated Responses, Descendant Experiments, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
2018
Blount Z D; Lenski R E; Losos J B
Contingency and determinism in evolution: Replaying life's tape Journal Article
Science, 362 (6415), pp. eaam5979, 2018, ISSN: 0036-8075.
Abstract | Links | BibTeX | Altmetric | Tags: Historical Contingency, Parallelism and Divergence, Review Articles
@article{Blount2018,
title = {Contingency and determinism in evolution: Replaying life's tape},
author = {Zachary D. Blount and Richard E. Lenski and Jonathan B. Losos},
url = {https://www.sciencemag.org/lookup/doi/10.1126/science.aam5979},
doi = {10.1126/science.aam5979},
issn = {0036-8075},
year = {2018},
date = {2018-11-01},
urldate = {2018-11-01},
journal = {Science},
volume = {362},
number = {6415},
pages = {eaam5979},
abstract = {Historical processes display some degree of "contingency," meaning their outcomes are sensitive to seemingly inconsequential events that can fundamentally change the future. Contingency is what makes historical outcomes unpredictable. Unlike many other natural phenomena, evolution is a historical process. Evolutionary change is often driven by the deterministic force of natural selection, but natural selection works upon variation that arises unpredictably through time by random mutation, and even beneficial mutations can be lost by chance through genetic drift. Moreover, evolution has taken place within a planetary environment with a particular history of its own. This tension between determinism and contingency makes evolutionary biology a kind of hybrid between science and history. While philosophers of science examine the nuances of contingency, biologists have performed many empirical studies of evolutionary repeatability and contingency. Here, we review the experimental and comparative evidence from these studies. Replicate populations in evolutionary "replay" experiments often show parallel changes, especially in overall performance, although idiosyncratic outcomes show that the particulars of a lineage's history can affect which of several evolutionary paths is taken. Comparative biologists have found many notable examples of convergent adaptation to similar conditions, but quantification of how frequently such convergence occurs is difficult. On balance, the evidence indicates that evolution tends to be surprisingly repeatable among closely related lineages, but disparate outcomes become more likely as the footprint of history grows deeper. Ongoing research on the structure of adaptive landscapes is providing additional insight into the interplay of fate and chance in the evolutionary process.},
keywords = {Historical Contingency, Parallelism and Divergence, Review Articles},
pubstate = {published},
tppubtype = {article}
}
Leon D; D'Alton S; Quandt E M; Barrick J E
Innovation in an E. coli evolution experiment is contingent on maintaining adaptive potential until competition subsides Journal Article
PLOS Genetics, 14 (4), pp. e1007348, 2018, ISSN: 1553-7404.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Demography and Ecology, Historical Contingency
@article{nokey,
title = {Innovation in an \textit{E. coli} evolution experiment is contingent on maintaining adaptive potential until competition subsides},
author = {Dacia Leon and Simon D'Alton and Erik M. Quandt and Jeffrey E. Barrick},
url = {https://dx.plos.org/10.1371/journal.pgen.1007348},
doi = {10.1371/journal.pgen.1007348},
issn = {1553-7404},
year = {2018},
date = {2018-04-12},
urldate = {2018-04-12},
journal = {PLOS Genetics},
volume = {14},
number = {4},
pages = {e1007348},
abstract = {Key innovations are disruptive evolutionary events that enable a species to escape constraints and rapidly diversify. After 15 years of the Lenski long-term evolution experiment with \textit{Escherichia coli}, cells in one of the twelve populations evolved the ability to utilize citrate, an abundant but previously untapped carbon source in the environment. Descendants of these cells became dominant in the population and subsequently diversified as a consequence of invading this vacant niche. Mutations responsible for the appearance of rudimentary citrate utilization and for refining this ability have been characterized. However, the complete nature of the genetic and/or ecological events that set the stage for this key innovation is unknown. In particular, it is unclear why it took so long for citrate utilization to evolve and why it still has evolved in only one of the twelve \textit{E. coli} populations after 30 years of the Lenski experiment. In this study, we recapitulated the initial mutation needed to evolve citrate utilization in strains isolated from throughout the first 31,500 generations of the history of this population. We found that there was already a slight fitness benefit for this mutation in the original ancestor of the evolution experiment and in other early isolates. However, evolution of citrate utilization was blocked at this point due to competition with other mutations that improved fitness in the original niche. Subsequently, an anti-potentiated genetic background evolved in which it was deleterious to evolve rudimentary citrate utilization. Only later, after further mutations accumulated that restored the benefit of this first-step mutation and the overall rate of adaptation in the population slowed, was citrate utilization likely to evolve. Thus, intense competition and the types of mutations that it favors can lead to short-sighted evolutionary trajectories that hide a stepping stone needed to access a key innovation from many future generations.},
keywords = {Citrate Evolution, Demography and Ecology, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
Peng F; Widmann S; Wünsche A; Duan K; Donovan K A; Dobson R C J; Lenski R E; Cooper T F
Effects of Beneficial Mutations in pykF Gene Vary over Time and across Replicate Populations in a Long-Term Experiment with Bacteria Journal Article
Molecular Biology and Evolution, 35 (1), pp. 202–210, 2018, ISSN: 0737-4038.
Abstract | Links | BibTeX | Altmetric | Tags: Genotypes and Phenotypes, Historical Contingency
@article{Peng2018,
title = {Effects of Beneficial Mutations in \textit{pykF} Gene Vary over Time and across Replicate Populations in a Long-Term Experiment with Bacteria},
author = {Fen Peng and Scott Widmann and Andrea Wünsche and Kristina Duan and Katherine A. Donovan and Renwick C J Dobson and Richard E. Lenski and Tim F. Cooper},
url = {https://academic.oup.com/mbe/article/35/1/202/4562833},
doi = {10.1093/molbev/msx279},
issn = {0737-4038},
year = {2018},
date = {2018-01-01},
urldate = {2018-01-01},
journal = {Molecular Biology and Evolution},
volume = {35},
number = {1},
pages = {202--210},
abstract = {The fitness effects of mutations can depend on the genetic backgrounds in which they occur and thereby influence future opportunities for evolving populations. In particular, mutations that fix in a population might change the selective benefit of subsequent mutations, giving rise to historical contingency. We examine these effects by focusing on mutations in a key metabolic gene, \textit{pykF}, that arose independently early in the history of 12 \textit{Escherichia coli} populations during a long-Term evolution experiment. Eight different evolved nonsynonymous mutations conferred similar fitness benefits of ~10% when transferred into the ancestor, and these benefits were greater than the one conferred by a deletion mutation. In contrast, the same mutations had highly variable fitness effects, ranging from ~0% to 25%, in evolved clones isolated from the populations at 20,000 generations. Two mutations that were moved into these evolved clones conferred similar fitness effects in a given clone, but different effects between the clones, indicating epistatic interactions between the evolved \textit{pykF} alleles and the other mutations that had accumulated in each evolved clone. We also measured the fitness effects of six evolved \textit{pykF} alleles in the same populations in which they had fixed, but at seven time points between 0 and 50,000 generations. Variation in fitness effects was high at intermediate time points, and declined to a low level at 50,000 generations, when the mean fitness effect was lowest. Our results demonstrate the importance of genetic context in determining the fitness effects of different beneficial mutations even within the same gene.},
keywords = {Genotypes and Phenotypes, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
2017
Good B H; McDonald M J; Barrick J E; Lenski R E; Desai M M
The dynamics of molecular evolution over 60,000 generations Journal Article
Nature, 551 (7678), pp. 45–50, 2017, ISSN: 14764687.
Abstract | Links | BibTeX | Altmetric | Tags: Demography and Ecology, Genome Evolution, Historical Contingency, Mutation Rates, Parallelism and Divergence
@article{Good2017,
title = {The dynamics of molecular evolution over 60,000 generations},
author = {Benjamin H. Good and Michael J. McDonald and Jeffrey E. Barrick and Richard E. Lenski and Michael M. Desai},
url = {http://dx.doi.org/10.1038/nature24287},
doi = {10.1038/nature24287},
issn = {14764687},
year = {2017},
date = {2017-01-01},
urldate = {2017-01-01},
journal = {Nature},
volume = {551},
number = {7678},
pages = {45--50},
publisher = {Nature Publishing Group},
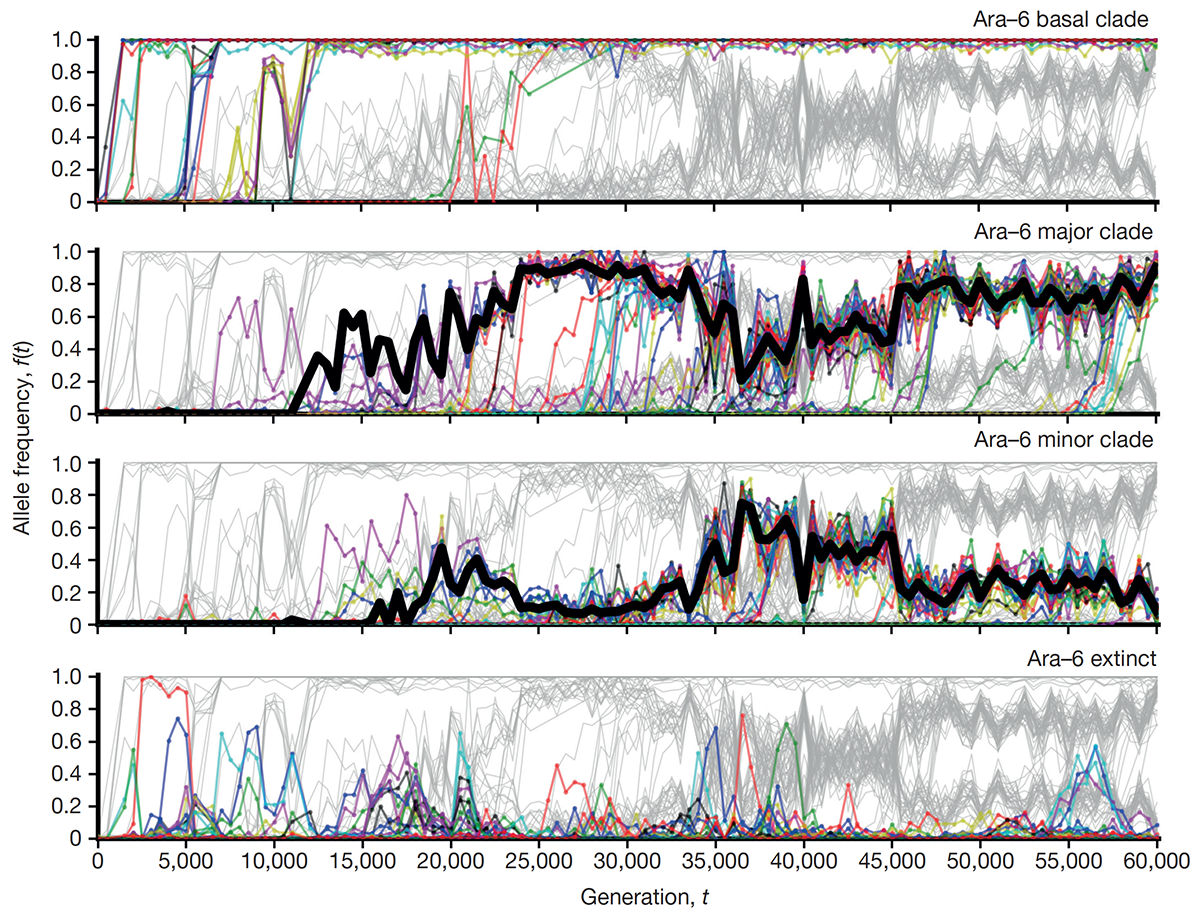
abstract = {The outcomes of evolution are determined by a stochastic dynamical process that governs how mutations arise and spread through a population. However, it is difficult to observe these dynamics directly over long periods and across entire genomes. Here we analyse the dynamics of molecular evolution in twelve experimental populations of \textit{Escherichia coli}, using whole-genome metagenomic sequencing at five hundred-generation intervals through sixty thousand generations. Although the rate of fitness gain declines over time, molecular evolution is characterized by signatures of rapid adaptation throughout the duration of the experiment, with multiple beneficial variants simultaneously competing for dominance in each population. Interactions between ecological and evolutionary processes play an important role, as long-term quasi-stable coexistence arises spontaneously in most populations, and evolution continues within each clade. We also present evidence that the targets of natural selection change over time, as epistasis and historical contingency alter the strength of selection on different genes. Together, these results show that long-term adaptation to a constant environment can be a more complex and dynamic process than is often assumed.},
keywords = {Demography and Ecology, Genome Evolution, Historical Contingency, Mutation Rates, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
2015
Turner C B; Blount Z D; Lenski R E
Replaying Evolution to Test the Cause of Extinction of One Ecotype in an Experimentally Evolved Population Journal Article
PLOS ONE, 10 (11), pp. e0142050, 2015, ISSN: 1932-6203.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Demography and Ecology, Historical Contingency
@article{Turner2015,
title = {Replaying Evolution to Test the Cause of Extinction of One Ecotype in an Experimentally Evolved Population},
author = {Caroline B. Turner and Zachary D. Blount and Richard E. Lenski},
editor = {Frederick M. Cohan},
url = {https://dx.plos.org/10.1371/journal.pone.0142050},
doi = {10.1371/journal.pone.0142050},
issn = {1932-6203},
year = {2015},
date = {2015-11-01},
urldate = {2015-11-01},
journal = {PLOS ONE},
volume = {10},
number = {11},
pages = {e0142050},
abstract = {In a long-term evolution experiment with \textit{Escherichia coli}, bacteria in one of twelve populations evolved the ability to consume citrate, a previously unexploited resource in a glucoselimited medium. This innovation led to the frequency-dependent coexistence of citrate-consuming (Cit+) and non-consuming (Cit-) ecotypes, with Cit-bacteria persisting on the exogenously supplied glucose as well as other carbon molecules released by the Cit+ bacteria. After more than 10,000 generations of coexistence, however, the Cit-lineage went extinct; cells with the Cit-phenotype dropped to levels below detection, and the Cit-clade could not be detected by molecular assays based on its unique genotype. We hypothesized that this extinction was a deterministic outcome of evolutionary change within the population, specifically the appearance of a more-fit Cit+ ecotype that competitively excluded the Cit-ecotype. We tested this hypothesis by re-evolving the population from a frozen population sample taken within 500 generations of the extinction and from another sample taken several thousand generations earlier, in each case for 500 generations and with 20-fold replication. To our surprise, the Cit-type did not go extinct in any of these replays, and Cit-cells also persisted in a single replicate that was propagated for 2,500 generations. Even more unexpectedly, we showed that the Cit-ecotype could reinvade the Cit+ population after its extinction. Taken together, these results indicate that the extinction of the Cit-ecotype was not a deterministic outcome driven by competitive exclusion by the Cit+ ecotype. The extinction also cannot be explained by demographic stochasticity alone, as the population size of the Cit-ecotype should have been many thousands of cells even during the daily transfer events. Instead, we infer that the extinction must have been caused by a rare chance event in which some aspect of the experimental conditions was inadvertently perturbed.},
keywords = {Citrate Evolution, Demography and Ecology, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
Quandt E M; Gollihar J; Blount Z D; Ellington A D; Georgiou G; Barrick J E
Fine-tuning citrate synthase flux potentiates and refines metabolic innovation in the Lenski evolution experiment. Journal Article
eLife, 4 (October), pp. e09696, 2015, ISSN: 2050-084X.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Genotypes and Phenotypes, Historical Contingency
@article{Quandt2015,
title = {Fine-tuning citrate synthase flux potentiates and refines metabolic innovation in the Lenski evolution experiment.},
author = {Erik M. Quandt and Jimmy Gollihar and Zachary D. Blount and Andrew D. Ellington and George Georgiou and Jeffrey E. Barrick},
url = {http://www.ncbi.nlm.nih.gov/pubmed/26465114
http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=PMC4718724},
doi = {10.7554/eLife.09696},
issn = {2050-084X},
year = {2015},
date = {2015-10-01},
urldate = {2015-10-01},
journal = {eLife},
volume = {4},
number = {October},
pages = {e09696},
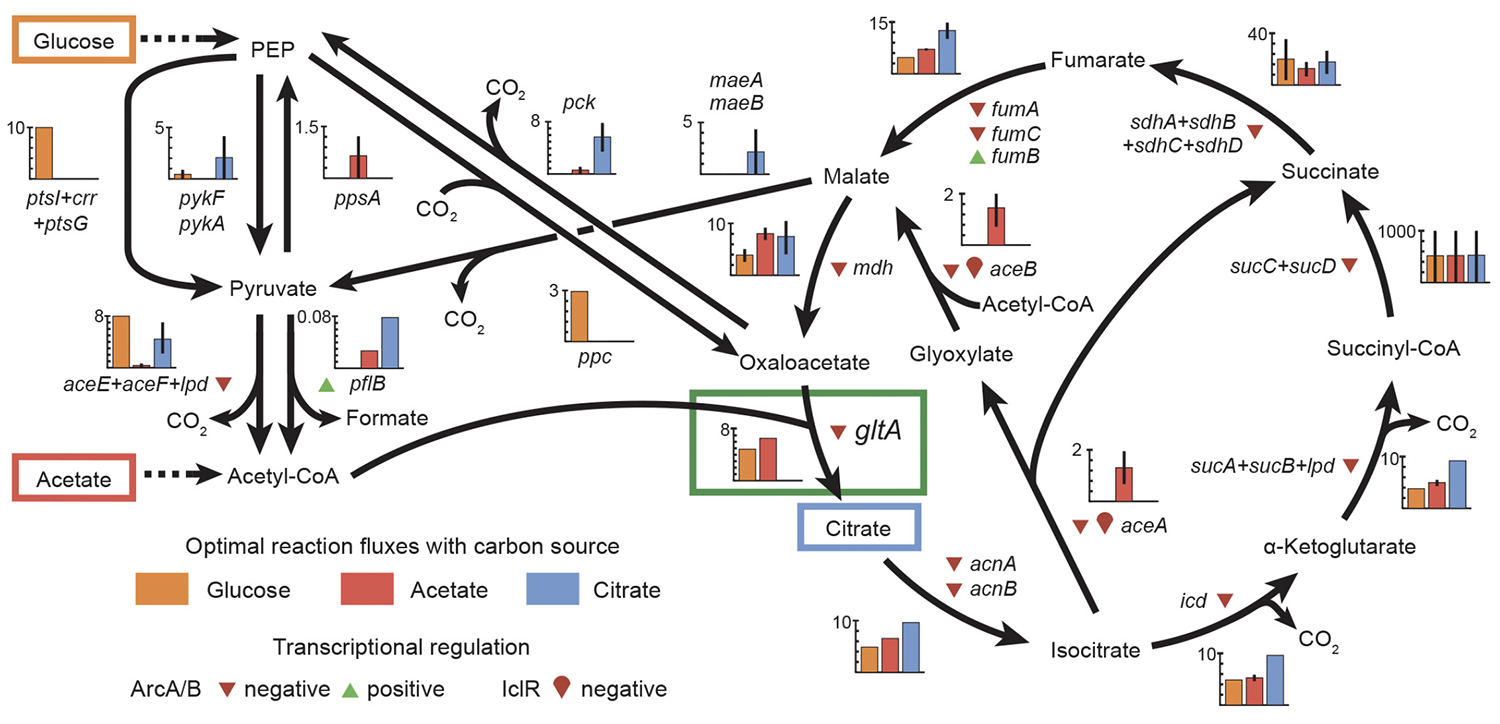
abstract = {Evolutionary innovations that enable organisms to colonize new ecological niches are rare compared to gradual evolutionary changes in existing traits. We discovered that key mutations in the \textit{gltA} gene, which encodes citrate synthase (CS), occurred both before and after \textit{Escherichia coli} gained the ability to grow aerobically on citrate (Cit(+) phenotype) during the Lenski long-term evolution experiment. The first \textit{gltA} mutation, which increases CS activity by disrupting NADH-inhibition of this enzyme, is beneficial for growth on the acetate and contributed to preserving the rudimentary Cit(+) trait from extinction when it first evolved. However, after Cit(+) was refined by further mutations, this potentiating \textit{gltA} mutation became deleterious to fitness. A second wave of beneficial \textit{gltA} mutations then evolved that reduced CS activity to below the ancestral level. Thus, dynamic reorganization of central metabolism made colonizing this new nutrient niche contingent on both co-opting and overcoming a history of prior adaptation.},
keywords = {Citrate Evolution, Genotypes and Phenotypes, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
2012
Blount Z D; Barrick J E; Davidson C J; Lenski R E
Genomic analysis of a key innovation in an experimental Escherichia coli population Journal Article
Nature, 489 (7417), pp. 513–518, 2012, ISSN: 0028-0836.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Genome Evolution, Genotypes and Phenotypes, Historical Contingency, Mutation Rates
@article{Blount2012,
title = {Genomic analysis of a key innovation in an experimental \emph{Escherichia coli} population},
author = {Zachary D. Blount and Jeffrey E. Barrick and Carla J. Davidson and Richard E. Lenski},
url = {http://www.nature.com/articles/nature11514},
doi = {10.1038/nature11514},
issn = {0028-0836},
year = {2012},
date = {2012-09-01},
urldate = {2012-09-01},
journal = {Nature},
volume = {489},
number = {7417},
pages = {513--518},
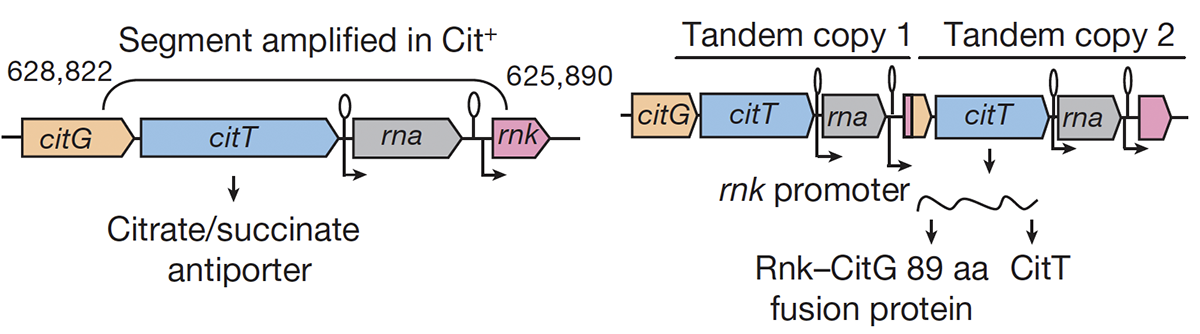
abstract = {Evolutionary novelties have been important in the history of life, but their origins are usually difficult to examine in detail. We previously described the evolution of a novel trait, aerobic citrate utilization (Cit +), in an experimental population of \textit{Escherichia coli}. Here we analyse genome sequences to investigate the history and genetic basis of this trait. At least three distinct clades coexisted for more than 10,000 generations before its emergence. The Cit + trait originated in one clade by a tandem duplication that captured an aerobically expressed promoter for the expression of a previously silent citrate transporter. The clades varied in their propensity to evolve this novel trait, although genotypes able to do so existed in all three clades, implying that multiple potentiating mutations arose during the population's history. Our findings illustrate the importance of promoter capture and altered gene regulation in mediating the exaptation events that often underlie evolutionary innovations.},
keywords = {Citrate Evolution, Genome Evolution, Genotypes and Phenotypes, Historical Contingency, Mutation Rates},
pubstate = {published},
tppubtype = {article}
}
2011
Woods R J; Barrick J E; Cooper T F; Shrestha U; Kauth M R; Lenski R E
Second-Order Selection for Evolvability in a Large Escherichia coli Population Journal Article
Science, 331 (6023), pp. 1433–1436, 2011, ISSN: 0036-8075.
Abstract | Links | BibTeX | Altmetric | Tags: Fitness Trajectories, Genome Evolution, Historical Contingency
@article{Woods2011,
title = {Second-Order Selection for Evolvability in a Large \emph{Escherichia coli} Population},
author = {Robert J. Woods and Jeffrey E. Barrick and Tim F. Cooper and Utpala Shrestha and Mark R. Kauth and Richard E. Lenski},
url = {https://www.sciencemag.org/lookup/doi/10.1126/science.1198914},
doi = {10.1126/science.1198914},
issn = {0036-8075},
year = {2011},
date = {2011-03-01},
urldate = {2011-03-01},
journal = {Science},
volume = {331},
number = {6023},
pages = {1433--1436},
abstract = {In theory, competition between asexual lineages can lead to second-order selection for greater evolutionary potential. To test this hypothesis, we revived a frozen population of \textit{Escherichia coli} from a long-term evolution experiment and compared the fitness and ultimate fates of four genetically distinct clones. Surprisingly, two clones with beneficial mutations that would eventually take over the population had significantly lower competitive fitness than two clones with mutations that later went extinct. By replaying evolution many times from these clones, we showed that the eventual winners likely prevailed because they had greater potential for further adaptation. Genetic interactions that reduce the benefit of certain regulatory mutations in the eventual losers appear to explain, at least in part, why they were outcompeted.},
keywords = {Fitness Trajectories, Genome Evolution, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
2008
Blount Z D; Borland C Z; Lenski R E
Historical contingency and the evolution of a key innovation in an experimental population of Escherichia coli Journal Article
Proceedings of the National Academy of Sciences of the United States of America, 105 (23), pp. 7899–7906, 2008, ISSN: 0027-8424.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Demography and Ecology, Historical Contingency
@article{Blount2008,
title = {Historical contingency and the evolution of a key innovation in an experimental population of \emph{Escherichia coli}},
author = {Zachary D. Blount and Christina Z. Borland and Richard E. Lenski},
url = {http://www.pnas.org/cgi/doi/10.1073/pnas.0803151105},
doi = {10.1073/pnas.0803151105},
issn = {0027-8424},
year = {2008},
date = {2008-06-01},
urldate = {2008-06-01},
journal = {Proceedings of the National Academy of Sciences of the United States of America},
volume = {105},
number = {23},
pages = {7899--7906},
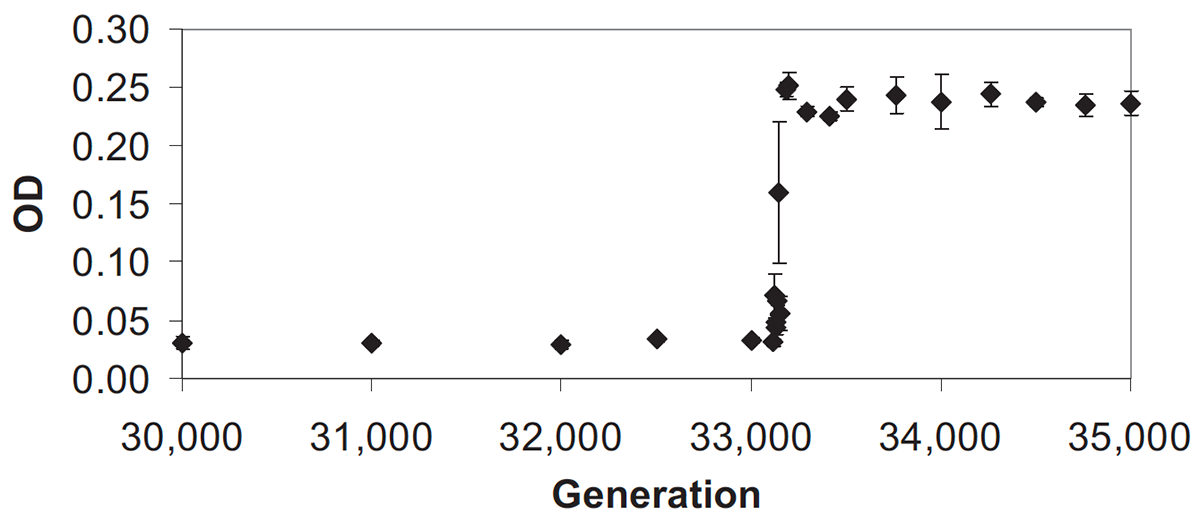
abstract = {The role of historical contingency in evolution has been much debated, but rarely tested. Twelve initially identical populations of \textit{Escherichia coli} were founded in 1988 to investigate this issue. They have since evolved in a glucose-limited medium that also contains citrate, which \textit{E. coli} cannot use as a carbon source under oxic conditions. No population evolved the capacity to exploit citrate for >30,000 generations, although each population tested billions of mutations. A citrate-using (Cit^{+}) variant finally evolved in one population by 31,500 generations, causing an increase in population size and diversity. The long-delayed and unique evolution of this function might indicate the involvement of some extremely rare mutation. Alternately, it may involve an ordinary mutation, but one whose physical occurrence or phenotypic expression is contingent on prior mutations in that population. We tested these hypotheses in experiments that “replayed” evolution from different points in that population's history. We observed no Cit^{+} mutants among 8.4 × 10^{12} ancestral cells, nor among 9 × 10^{12} cells from 60 clones sampled in the first 15,000 generations. However, we observed a significantly greater tendency for later clones to evolve Cit^{+}, indicating that some potentiating mutation arose by 20,000 generations. This potentiating change increased the mutation rate to Cit^{+} but did not cause generalized hypermutability. Thus, the evolution of this phenotype was contingent on the particular history of that population. More generally, we suggest that historical contingency is especially important when it facilitates the evolution of key innovations that are not easily evolved by gradual, cumulative selection.},
keywords = {Citrate Evolution, Demography and Ecology, Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
2001
Elena S F; Lenski R E
Epistasis between new mutations and genetic background and a test of genetic canalization Journal Article
Evolution, 55 (9), pp. 1746–1752, 2001, ISSN: 0014-3820.
Abstract | Links | BibTeX | Altmetric | Tags: Historical Contingency
@article{Elena2001,
title = {Epistasis between new mutations and genetic background and a test of genetic canalization},
author = {Santiago F. Elena and Richard E. Lenski},
url = {https://onlinelibrary.wiley.com/doi/10.1111/j.0014-3820.2001.tb00824.x},
doi = {10.1111/j.0014-3820.2001.tb00824.x},
issn = {0014-3820},
year = {2001},
date = {2001-09-01},
urldate = {2001-09-01},
journal = {Evolution},
volume = {55},
number = {9},
pages = {1746--1752},
abstract = {The importance for fitness of epistatic interactions among mutations is poorly known, yet epistasis can exert important effects on the dynamics of evolving populations. We showed previously that epistatic interactions are common between pairs of random insertion mutations in the bacterium \textit{Escherichia coli}. In this paper, we examine interactions between these mutations and other mutations by transducing each of twelve insertion mutations into two genetic backgrounds, one ancestral and the other having evolved in, and adapted to, a defined laboratory environment for 10,000 generations. To assess the effect of the mutation on fitness, we allowed each mutant to compete against its unmutated counterpart in that same environment. Overall, there was a strong positive correlation between the mutational effects on the two genetic backgrounds. Nonetheless, three of the twelve mutations had significantly different effects on the two backgrounds, indicating epistasis. There was no significant tendency for the mutations to be less harmful on the derived background. Thus, there is no evidence supporting the hypothesis that the derived bacteria had adapted, in part, by becoming buffered against the harmful effects of mutations.},
keywords = {Historical Contingency},
pubstate = {published},
tppubtype = {article}
}
1995
Travisano M; Mongold J A; Bennett A F; Lenski R E
Experimental Tests of the Roles of Adaptation, Chance, and History Journal Article
Science, 267 (January), 1995.
Abstract | Links | BibTeX | Altmetric | Tags: Cell Morphology, Correlated Responses, Descendant Experiments, Fitness Trajectories, Historical Contingency, Methods and Miscellaneous, Parallelism and Divergence
@article{Travisano1995,
title = {Experimental Tests of the Roles of Adaptation, Chance, and History},
author = {Michael Travisano and Judith A. Mongold and Albert F. Bennett and Richard E. Lenski},
url = {https://www.science.org/lookup/doi/10.1126/science.7809610},
doi = {https://doi.org/10.1126/science.7809610},
year = {1995},
date = {1995-01-01},
urldate = {1995-01-01},
journal = {Science},
volume = {267},
number = {January},
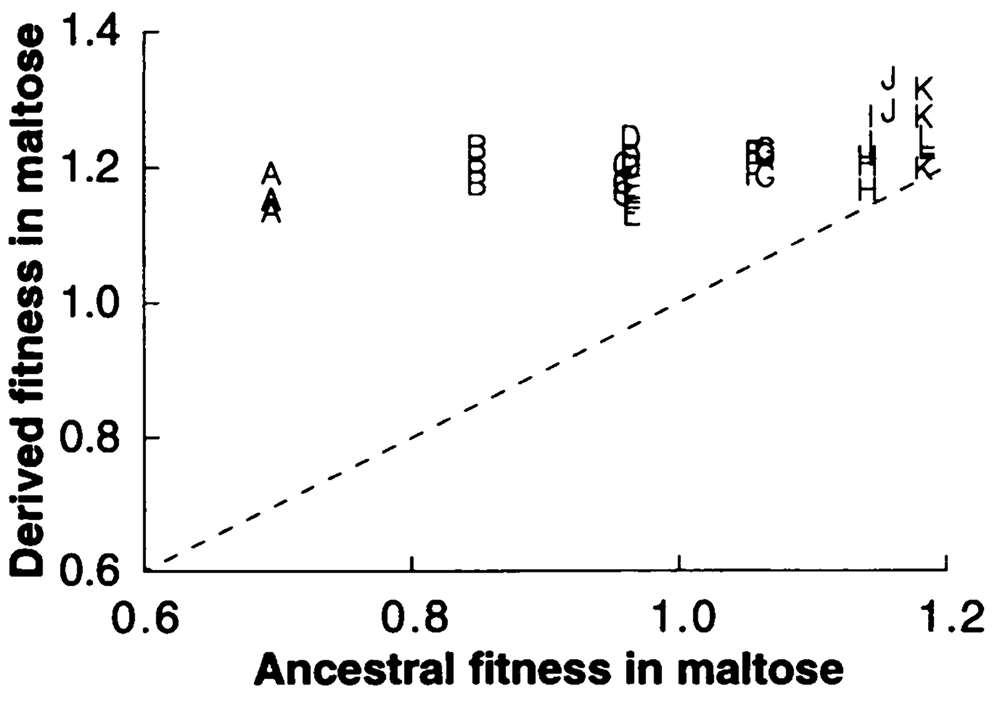
abstract = {The contributions of adaptation, chance, and history to the evolution of fitness and cell size were measured in two separate experiments using bacteria. In both experiments, populations propagated in identical environments achieved similar fitnesses, regardless of prior history or subsequent chance events. In contrast, the evolution of cell size, a trait weakly correlated with fitness, was more strongly influenced by history and chance.},
keywords = {Cell Morphology, Correlated Responses, Descendant Experiments, Fitness Trajectories, Historical Contingency, Methods and Miscellaneous, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}