2013
Wielgoss S; Barrick J E; Tenaillon O; Wiser M J; Dittmar W J; Cruveiller S; Chane-Woon-Ming B; Médigue C; Lenski R E; Schneider D
Mutation rate dynamics in a bacterial population reflect tension between adaptation and genetic load Journal Article
Proceedings of the National Academy of Sciences of the United States of America, 110 (1), pp. 222–227, 2013, ISSN: 0027-8424.
Abstract | Links | BibTeX | Altmetric | Tags: Fitness Trajectories, Genome Evolution, Genotypes and Phenotypes, Mutation Rates, Theory and Simulations
@article{Wielgoss2013,
title = {Mutation rate dynamics in a bacterial population reflect tension between adaptation and genetic load},
author = {Sébastien Wielgoss and Jeffrey E. Barrick and Olivier Tenaillon and Michael J. Wiser and W. James Dittmar and Stéphane Cruveiller and Béatrice Chane-Woon-Ming and Claudine Médigue and Richard E. Lenski and Dominique Schneider},
url = {http://www.pnas.org/cgi/doi/10.1073/pnas.1219574110},
doi = {10.1073/pnas.1219574110},
issn = {0027-8424},
year = {2013},
date = {2013-01-01},
urldate = {2013-01-01},
journal = {Proceedings of the National Academy of Sciences of the United States of America},
volume = {110},
number = {1},
pages = {222--227},
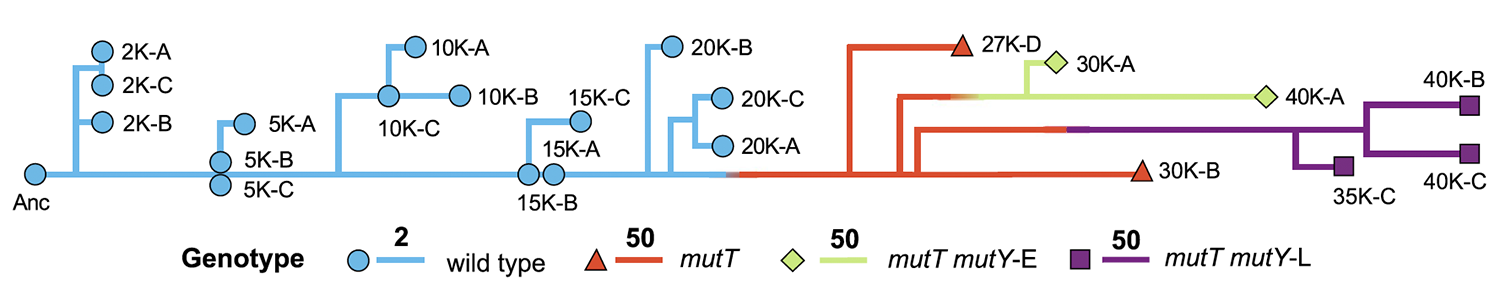
abstract = {Mutations are the ultimate source of heritable variation for evolution. Understanding how mutation rates themselves evolve is thus essential for quantitatively understanding many evolutionary processes. According to theory, mutation rates should be minimized for well-adapted populations living in stable environments, whereas hypermutators may evolve if conditions change. However, the long-term fate of hypermutators is unknown. Using a phylogenomic approach, we found that an adapting \textit{Escherichia coli} population that first evolved a \textit{mutT} hypermutator phenotype was later invaded by two independent lineages with \textit{mutY} mutations that reduced genome-wide mutation rates. Applying neutral theory to synonymous substitutions, we dated the emergence of these mutations and inferred that the \textit{mutT} mutation increased the point-mutation rate by ~150-fold, whereas the \textit{mutY} mutations reduced the rate by ~40-60%, with a corresponding decrease in the genetic load. Thus, the long-term fate of the hypermutators was governed by the selective advantage arising from a reduced mutation rate as the potential for further adaptation declined.},
keywords = {Fitness Trajectories, Genome Evolution, Genotypes and Phenotypes, Mutation Rates, Theory and Simulations},
pubstate = {published},
tppubtype = {article}
}
2012
Blount Z D; Barrick J E; Davidson C J; Lenski R E
Genomic analysis of a key innovation in an experimental Escherichia coli population Journal Article
Nature, 489 (7417), pp. 513–518, 2012, ISSN: 0028-0836.
Abstract | Links | BibTeX | Altmetric | Tags: Citrate Evolution, Genome Evolution, Genotypes and Phenotypes, Historical Contingency, Mutation Rates
@article{Blount2012,
title = {Genomic analysis of a key innovation in an experimental \emph{Escherichia coli} population},
author = {Zachary D. Blount and Jeffrey E. Barrick and Carla J. Davidson and Richard E. Lenski},
url = {http://www.nature.com/articles/nature11514},
doi = {10.1038/nature11514},
issn = {0028-0836},
year = {2012},
date = {2012-09-01},
urldate = {2012-09-01},
journal = {Nature},
volume = {489},
number = {7417},
pages = {513--518},
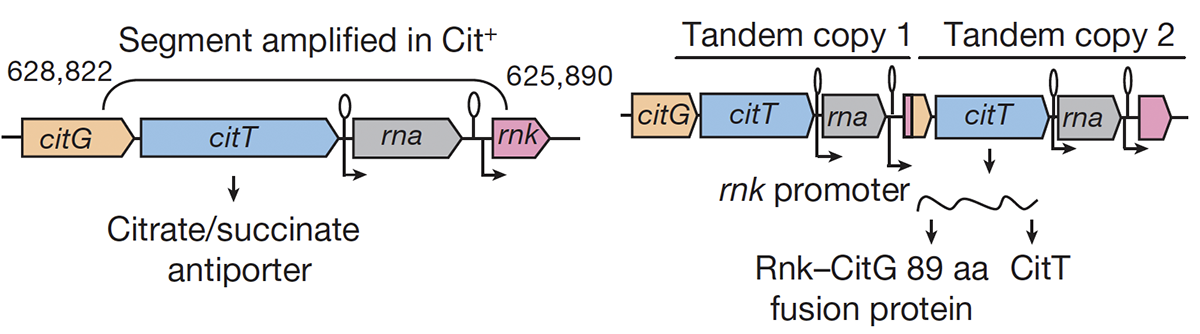
abstract = {Evolutionary novelties have been important in the history of life, but their origins are usually difficult to examine in detail. We previously described the evolution of a novel trait, aerobic citrate utilization (Cit +), in an experimental population of \textit{Escherichia coli}. Here we analyse genome sequences to investigate the history and genetic basis of this trait. At least three distinct clades coexisted for more than 10,000 generations before its emergence. The Cit + trait originated in one clade by a tandem duplication that captured an aerobically expressed promoter for the expression of a previously silent citrate transporter. The clades varied in their propensity to evolve this novel trait, although genotypes able to do so existed in all three clades, implying that multiple potentiating mutations arose during the population's history. Our findings illustrate the importance of promoter capture and altered gene regulation in mediating the exaptation events that often underlie evolutionary innovations.},
keywords = {Citrate Evolution, Genome Evolution, Genotypes and Phenotypes, Historical Contingency, Mutation Rates},
pubstate = {published},
tppubtype = {article}
}
Gac M L; Plucain J; Hindré T; Lenski R E; Schneider D
Ecological and evolutionary dynamics of coexisting lineages during a long-term experiment with Escherichia coli Journal Article
Proceedings of the National Academy of Sciences of the United States of America, 109 (24), pp. 9487–9492, 2012, ISSN: 0027-8424.
Abstract | Links | BibTeX | Altmetric | Tags: Demography and Ecology, Genotypes and Phenotypes
@article{LeGac2012,
title = {Ecological and evolutionary dynamics of coexisting lineages during a long-term experiment with \textit{Escherichia coli}},
author = {Mickaël Le Gac and Jessica Plucain and Thomas Hindré and Richard E. Lenski and Dominique Schneider},
url = {http://www.pnas.org/cgi/doi/10.1073/pnas.1207091109},
doi = {10.1073/pnas.1207091109},
issn = {0027-8424},
year = {2012},
date = {2012-06-01},
urldate = {2012-06-01},
journal = {Proceedings of the National Academy of Sciences of the United States of America},
volume = {109},
number = {24},
pages = {9487--9492},
abstract = {Closely related organisms usually occupy similar ecological niches, leading to intense competition and even extinction. Such competition also can promote rapid phenotypic evolution and ecological divergence. This process may end with the stable occupation of distinct niches or, alternatively, may entail repeated bouts of evolution. Here we examine two \textit{Escherichia coli} lineages, called L and S, that coexisted for more than 30,000 generations after diverging from a common ancestor. Both lineages underwent sustained phenotypic evolution based on global transcription and resource utilization profiles, with L seeming to encroach over time on the catabolic profile of S. Reciprocal invasion experiments with L and S clones from the same or different generations revealed evolutionary changes in their interaction, including an asymmetry that confirmed the encroachment by L on the niche of the S lineage. In general, L and S clones from the same generation showed negative frequency-dependent effects, consistent with stable coexistence. However, L clones could invade S clones from both earlier and later generations, whereas S clones could invade only L clones from earlier generations. In this system, the long-term coexistence of competing lineages evidently depended on successive rounds of evolution, rather than on initial divergence followed by a static equilibrium.},
keywords = {Demography and Ecology, Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}
2011
Khan A I; Dinh D M; Schneider D; Lenski R E; Cooper T F
Negative Epistasis Between Beneficial Mutations in an Evolving Bacterial Population Journal Article
Science, 332 (6034), pp. 1193–1196, 2011, ISSN: 0036-8075.
Abstract | Links | BibTeX | Altmetric | Tags: Fitness Trajectories, Genome Evolution, Genotypes and Phenotypes
@article{Khan2011,
title = {Negative Epistasis Between Beneficial Mutations in an Evolving Bacterial Population},
author = {Aisha I. Khan and Duy M. Dinh and Dominique Schneider and Richard E. Lenski and Tim F. Cooper},
url = {https://www.sciencemag.org/lookup/doi/10.1126/science.1203801},
doi = {10.1126/science.1203801},
issn = {0036-8075},
year = {2011},
date = {2011-06-01},
urldate = {2011-06-01},
journal = {Science},
volume = {332},
number = {6034},
pages = {1193--1196},
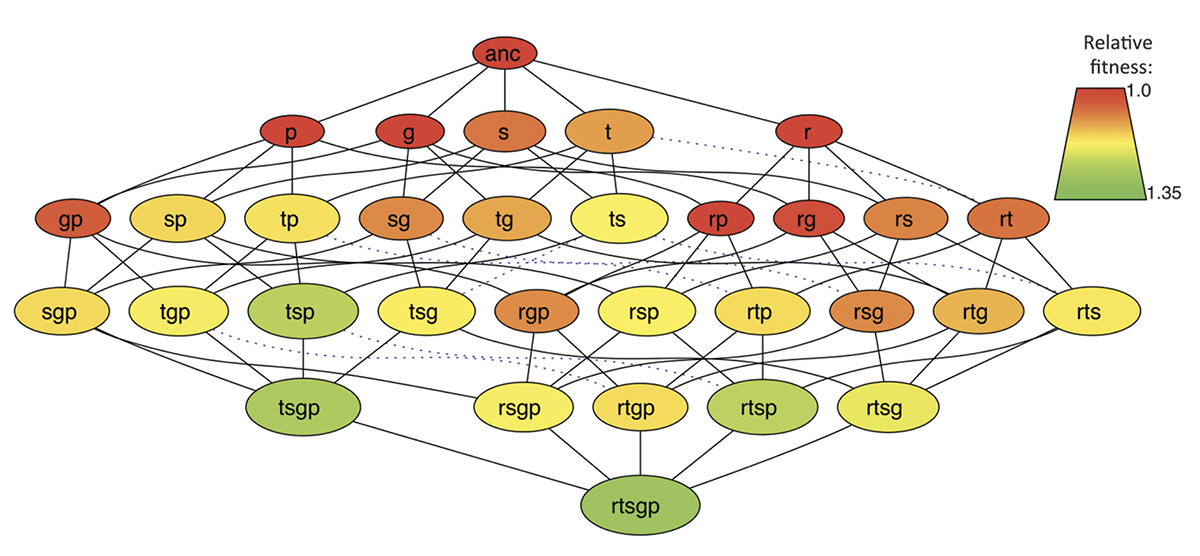
abstract = {Epistatic interactions between mutations play a prominent role in evolutionary theories. Many studies have found that epistasis is widespread, but they have rarely considered beneficial mutations. We analyzed the effects of epistasis on fitness for the first five mutations to fix in an experimental population of \textit{Escherichia coli}. Epistasis depended on the effects of the combined mutations - the larger the expected benefit, the more negative the epistatic effect. Epistasis thus tended to produce diminishing returns with genotype fitness, although interactions involving one particular mutation had the opposite effect. These data support models in which negative epistasis contributes to declining rates of adaptation over time. Sign epistasis was rare in this genome-wide study, in contrast to its prevalence in an earlier study of mutations in a single gene.},
keywords = {Fitness Trajectories, Genome Evolution, Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}
Crozat E; Hindré T; Kühn L; Garin J; Lenski R E; Schneider D
Altered Regulation of the OmpF Porin by Fis in Escherichia coli during an Evolution Experiment and between B and K-12 Strains Journal Article
Journal of Bacteriology, 193 (2), pp. 429–440, 2011, ISSN: 0021-9193.
Abstract | Links | BibTeX | Altmetric | Tags: Genotypes and Phenotypes
@article{Crozat2011,
title = {Altered Regulation of the OmpF Porin by Fis in \textit{Escherichia coli} during an Evolution Experiment and between B and K-12 Strains},
author = {Estelle Crozat and Thomas Hindré and Lauriane Kühn and Jérome Garin and Richard E. Lenski and Dominique Schneider},
url = {https://journals.asm.org/doi/10.1128/JB.01341-10},
doi = {10.1128/JB.01341-10},
issn = {0021-9193},
year = {2011},
date = {2011-01-01},
urldate = {2011-01-01},
journal = {Journal of Bacteriology},
volume = {193},
number = {2},
pages = {429--440},
abstract = {The phenotypic plasticity of global regulatory networks provides bacteria with rapid acclimation to a wide range of environmental conditions, while genetic changes in those networks provide additional flexibility as bacteria evolve across long time scales. We previously identified mutations in the global regulator-encoding gene \textit{fis} that enhanced organismal fitness during a long-term evolution experiment with \textit{Escherichia coli}. To gain insight into the effects of these mutations, we produced two-dimensional protein gels with strains carrying different \textit{fis} alleles, including a beneficial evolved allele and one with an in-frame deletion. We found that Fis controls the expression of the major porin-encoding gene \textit{ompF} in the \textit{E. coli} B-derived ancestral strain used in the evolution experiment, a relationship that has not been described before. We further showed that this regulatory connection evolved over two different time scales, perhaps explaining why it was not observed before. On the longer time scale, we showed that this regulation of \textit{ompF} by Fis is absent from the more widely studied K-12 strain and thus is specific to the B strain. On a shorter time scale, this regulatory linkage was lost during 20,000 generations of experimental evolution of the B strain. Finally, we mapped the Fis binding sites in the \textit{ompF} regulatory region, and we present a hypothetical model of \textit{ompF} expression that includes its other known regulators.},
keywords = {Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}
2010
Meyer J R; Agrawal A A; Quick R T; Dobias D T; Schneider D; Lenski R E
Parallel changes in host resistance to viral infection during 45,000 generations of relaxed selection Journal Article
Evolution, 64 (10), pp. no–no, 2010, ISSN: 00143820.
Abstract | Links | BibTeX | Altmetric | Tags: Correlated Responses, Genotypes and Phenotypes, Parallelism and Divergence
@article{Meyer2010,
title = {Parallel changes in host resistance to viral infection during 45,000 generations of relaxed selection},
author = {Justin R. Meyer and Anurag A. Agrawal and Ryan T. Quick and Devin T. Dobias and Dominique Schneider and Richard E. Lenski},
url = {https://onlinelibrary.wiley.com/doi/10.1111/j.1558-5646.2010.01049.x},
doi = {10.1111/j.1558-5646.2010.01049.x},
issn = {00143820},
year = {2010},
date = {2010-08-01},
urldate = {2010-08-01},
journal = {Evolution},
volume = {64},
number = {10},
pages = {no--no},
abstract = {The dynamics of host susceptibility to parasites are often influenced by trade-offs between the costs and benefits of resistance. We assayed changes in the resistance to three viruses in six lines of \textit{Escherichia coli} that had been evolving for almost 45,000 generations in their absence. The common ancestor of these lines was completely resistant to T6, partially resistant to T6* (a mutant of T6 with altered host range), and sensitive to lambda. None of the populations changed with respect to resistance to T6, whereas all six evolved increased susceptibility to T6*, probably ameliorating a cost of resistance. More surprisingly, however, the majority of lines evolved complete resistance to lambda, despite not encountering that virus during this period. By coupling our results with previous work, we infer that resistance to lambda evolved as a pleiotropic effect of a beneficial mutation that downregulated an unused metabolic pathway. The strong parallelism between the lines implies that selection had almost deterministic effects on the evolution of these patterns of host resistance. The opposite outcomes for resistance to T6* and lambda demonstrate that the evolution of host resistance under relaxed selection cannot be fully predicted by simple trade-off models.},
keywords = {Correlated Responses, Genotypes and Phenotypes, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
2009
Barrick J E; Yu D S; Yoon S H; Jeong H; Oh T K; Schneider D; Lenski R E; Kim J F
Genome evolution and adaptation in a long-term experiment with Escherichia coli. Journal Article
Nature, 461 (7268), pp. 1243–7, 2009, ISSN: 1476-4687.
Abstract | Links | BibTeX | Altmetric | Tags: Fitness Trajectories, Genome Evolution, Genotypes and Phenotypes, Mutation Rates, Parallelism and Divergence
@article{Barrick2009,
title = {Genome evolution and adaptation in a long-term experiment with \textit{Escherichia coli}.},
author = {Jeffrey E. Barrick and Dong Su Yu and Sung Ho Yoon and Haeyoung Jeong and Tae Kwang Oh and Dominique Schneider and Richard E. Lenski and Jihyun F. Kim},
url = {http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=19838166&retmode=ref&cmd=prlinks papers2://publication/doi/10.1038/nature08480 http://www.ncbi.nlm.nih.gov/pubmed/19838166},
doi = {10.1038/nature08480},
issn = {1476-4687},
year = {2009},
date = {2009-10-01},
urldate = {2009-10-01},
journal = {Nature},
volume = {461},
number = {7268},
pages = {1243--7},
publisher = {Nature Publishing Group},
address = {Department of Microbiology and Molecular Genetics, Michigan State University, East Lansing, Michigan 48824, USA.},
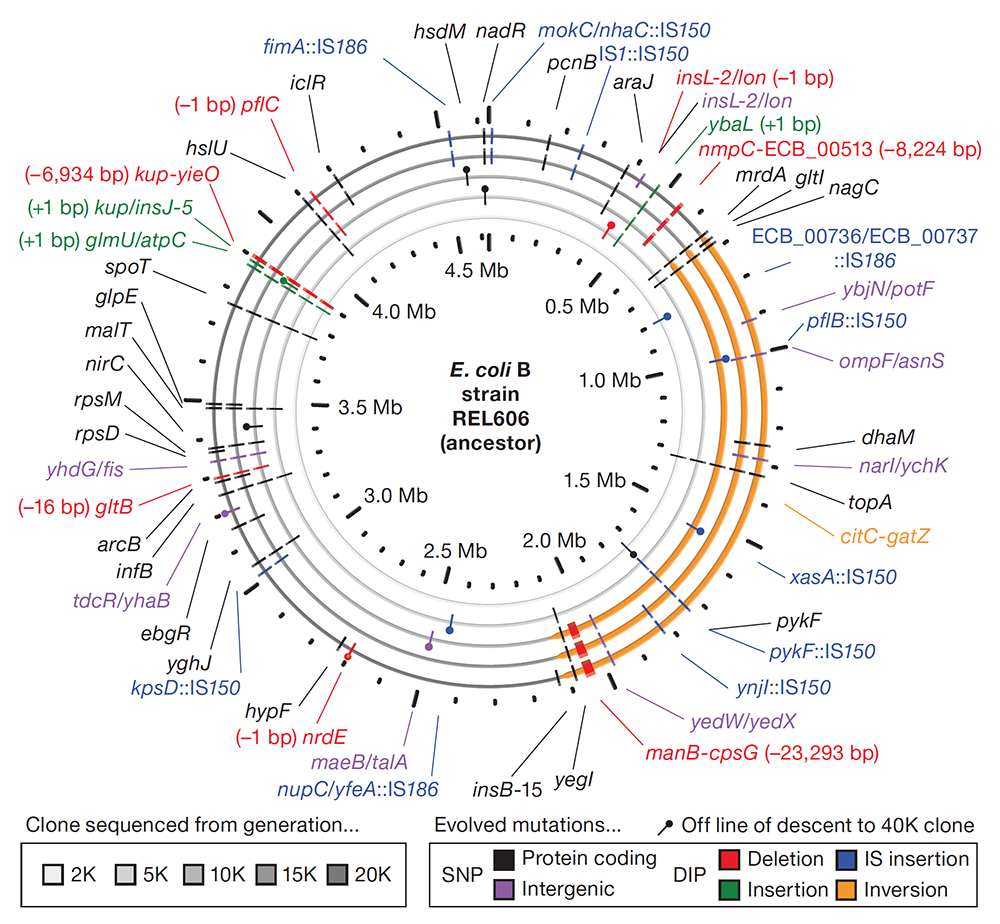
abstract = {The relationship between rates of genomic evolution and organismal adaptation remains uncertain, despite considerable interest. The feasibility of obtaining genome sequences from experimentally evolving populations offers the opportunity to investigate this relationship with new precision. Here we sequence genomes sampled through 40,000 generations from a laboratory population of \textit{Escherichia coli}. Although adaptation decelerated sharply, genomic evolution was nearly constant for 20,000 generations. Such clock-like regularity is usually viewed as the signature of neutral evolution, but several lines of evidence indicate that almost all of these mutations were beneficial. This same population later evolved an elevated mutation rate and accumulated hundreds of additional mutations dominated by a neutral signature. Thus, the coupling between genomic and adaptive evolution is complex and can be counterintuitive even in a constant environment. In particular, beneficial substitutions were surprisingly uniform over time, whereas neutral substitutions were highly variable.},
keywords = {Fitness Trajectories, Genome Evolution, Genotypes and Phenotypes, Mutation Rates, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
Philippe N; Pelosi L; Lenski R E; Schneider D
Evolution of Penicillin-Binding Protein 2 Concentration and Cell Shape during a Long-Term Experiment with Escherichia coli Journal Article
Journal of Bacteriology, 191 (3), pp. 909-921, 2009, ISSN: 0021-9193.
Abstract | Links | BibTeX | Altmetric | Tags: Cell Morphology, Genotypes and Phenotypes
@article{nokey,
title = {Evolution of Penicillin-Binding Protein 2 Concentration and Cell Shape during a Long-Term Experiment with \textit{Escherichia coli}},
author = {Nadège Philippe and Ludovic Pelosi and Richard E. Lenski and Dominique Schneider
},
url = {https://journals.asm.org/doi/full/10.1128/JB.01419-08},
doi = {10.1128/JB.01419-08},
issn = {0021-9193},
year = {2009},
date = {2009-02-01},
urldate = {2009-02-01},
journal = {Journal of Bacteriology},
volume = {191},
number = {3},
pages = {909-921},
abstract = {Peptidoglycan is the major component of the bacterial cell wall and is involved in osmotic protection and in determining cell shape. Cell shape potentially influences many processes, including nutrient uptake as well as cell survival and growth. Peptidoglycan is a dynamic structure that changes during the growth cycle. Penicillin-binding proteins (PBPs) catalyze the final stages of peptidoglycan synthesis. Although PBPs are biochemically and physiologically well characterized, their broader effects, especially their effects on organismal fitness, are not well understood. In a long-term experiment, 12 populations of \textit{Escherichia coli} having a common ancestor were allowed to evolve for more than 40,000 generations in a defined environment. We previously identified mutations in the \textit{pbpA} operon in one-half of these populations; this operon encodes PBP2 and RodA proteins that are involved in cell wall elongation. In this study, we characterized the effects of two of these mutations on competitive fitness and other phenotypes. By constructing and performing competition experiments with strains that are isogenic except for the \textit{pbpA} alleles, we showed that both mutations that evolved were beneficial in the environment used for the long-term experiment and that these mutations caused parallel phenotypic changes. In particular, they reduced the cellular concentration of PBP2, thereby generating spherical cells with an increased volume. In contrast to their fitness-enhancing effect in the environment where they evolved, both mutations decreased cellular resistance to osmotic stress. Moreover, one mutation reduced fitness during prolonged stationary phase. Therefore, alteration of the PBP2 concentration contributed to physiological trade-offs and ecological specialization during experimental evolution.},
keywords = {Cell Morphology, Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}
2008
Ostrowski E A; Woods R J; Lenski R E
The genetic basis of parallel and divergent phenotypic responses in evolving populations of Escherichia coli Journal Article
Proceedings of the Royal Society B: Biological Sciences, 275 (1632), pp. 277–284, 2008, ISSN: 0962-8452.
Abstract | Links | BibTeX | Altmetric | Tags: Descendant Experiments, Genotypes and Phenotypes
@article{Ostrowski2008,
title = {The genetic basis of parallel and divergent phenotypic responses in evolving populations of \textit{Escherichia coli}},
author = {Elizabeth A. Ostrowski and Robert J. Woods and Richard E. Lenski},
url = {https://royalsocietypublishing.org/doi/10.1098/rspb.2007.1244},
doi = {10.1098/rspb.2007.1244},
issn = {0962-8452},
year = {2008},
date = {2008-02-01},
urldate = {2008-02-01},
journal = {Proceedings of the Royal Society B: Biological Sciences},
volume = {275},
number = {1632},
pages = {277--284},
abstract = {Pleiotropy plays a central role in theories of adaptation, but little is known about the distribution of pleiotropic effects associated with different adaptive mutations. Previously, we described the phenotypic effects of a collection of independently arising beneficial mutations in \textit{Escherichia coli}. We quantified their fitness effects in the glucose environment in which they evolved and their pleiotropic effects in five novel resource environments. Here we use a candidate gene approach to associate the phenotypic effects of the mutations with the underlying genetic changes. Among our collection of 27 adaptive mutants, we identified a total of 21 mutations (18 of which were unique) encompassing five different loci or gene regions. There was limited resolution to distinguish among loci based on their fitness effects in the glucose environment, demonstrating widespread parallelism in the direct response to selection. However, substantial heterogeneity in mutant effects was revealed when we examined their pleiotropic effects on fitness in the five novel environments. Substitutions in the same locus clustered together phenotypically, indicating concordance between molecular and phenotypic measures of divergence.},
keywords = {Descendant Experiments, Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}
Cooper T F; Remold S K; Lenski R E; Schneider D
Expression Profiles Reveal Parallel Evolution of Epistatic Interactions Involving the CRP Regulon in Escherichia coli Journal Article
PLoS Genetics, 4 (2), pp. e35, 2008, ISSN: 1553-7404.
Abstract | Links | BibTeX | Altmetric | Tags: Genotypes and Phenotypes, Parallelism and Divergence
@article{Cooper2008,
title = {Expression Profiles Reveal Parallel Evolution of Epistatic Interactions Involving the CRP Regulon in \textit{Escherichia coli}},
author = {Tim F. Cooper and Susanna K. Remold and Richard E. Lenski and Dominique Schneider},
editor = {David S Guttman},
url = {https://dx.plos.org/10.1371/journal.pgen.0040035},
doi = {10.1371/journal.pgen.0040035},
issn = {1553-7404},
year = {2008},
date = {2008-02-01},
urldate = {2008-02-01},
journal = {PLoS Genetics},
volume = {4},
number = {2},
pages = {e35},
abstract = {The extent and nature of epistatic interactions between mutations are issues of fundamental importance in evolutionary biology. However, they are difficult to study and their influence on adaptation remains poorly understood. Here, we use a systems-level approach to examine epistatic interactions that arose during the evolution of \textit{Escherichia coli} in a defined environment. We used expression arrays to compare the effect on global patterns of gene expression of deleting a central regulatory gene, \textit{crp}. Effects were measured in two lineages that had independently evolved for 20,000 generations and in their common ancestor. We found that deleting \textit{crp} had a much more dramatic effect on the expression profile of the two evolved lines than on the ancestor. Because the sequence of the \textit{crp} gene was unchanged during evolution, these differences indicate epistatic interactions between \textit{crp} and mutations at other loci that accumulated during evolution. Moreover, a striking degree of parallelism was observed between the two independently evolved lines; 115 genes that were not \textit{crp}-dependent in the ancestor became dependent on \textit{crp} in both evolved lines. An analysis of changes in \textit{crp} dependence of well-characterized regulons identified a number of regulatory genes as candidates for harboring beneficial mutations that could account for these parallel expression changes. Mutations within three of these genes have previously been found and shown to contribute to fitness. Overall, these findings indicate that epistasis has been important in the adaptive evolution of these lines, and they provide new insight into the types of genetic changes through which epistasis can evolve. More generally, we demonstrate that expression profiles can be profitably used to investigate epistatic interactions. },
keywords = {Genotypes and Phenotypes, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
2006
Cooper T F; Morby A P; Gunn A; Schneider D
Effect of random and hub gene disruptions on environmental and mutational robustness in Escherichia coli Journal Article
BMC Genomics, 7 (1), pp. 237, 2006, ISSN: 1471-2164.
Abstract | Links | BibTeX | Altmetric | Tags: Genotypes and Phenotypes
@article{Cooper2006,
title = {Effect of random and hub gene disruptions on environmental and mutational robustness in \textit{Escherichia coli}},
author = {Tim F. Cooper and Andrew P. Morby and Annabel Gunn and Dominique Schneider},
url = {https://bmcgenomics.biomedcentral.com/articles/10.1186/1471-2164-7-237},
doi = {10.1186/1471-2164-7-237},
issn = {1471-2164},
year = {2006},
date = {2006-12-01},
urldate = {2006-12-01},
journal = {BMC Genomics},
volume = {7},
number = {1},
pages = {237},
abstract = {Background
Genome-wide profiling has allowed the regulatory interaction networks of many organisms to be visualised and the pattern of connections between genes to be studied. These networks are non-random, following a power-law distribution with a small number of well-connected 'hubs' and many genes with only one or a few connections. Theoretical work predicts that power-law networks display several unique properties. One of the most biologically interesting of these is an intrinsic robustness to disturbance such that removal of a random gene will have little effect on network function. Conversely, targeted removal of a hub gene is expected to have a large effect.
Results
We compared the response of \textit{Escherichia coli} to environmental and mutational stress following disruption of random or hub genes. We found that disruption of random genes had less effect on robustness to environmental stress than did the targeted disruption of hub genes. In contrast, random disruption strains were slightly less robust to the effect of mutational stress than were hub disruption strains. When we compared the effect of each disruption on environmental and mutational stress, we found a negative relationship, such that strains that were more environmentally robust tended to be less robust to mutational stress.
Conclusion
Our results demonstrate that mutant strains of \textit{E. coli} respond differently to stress, depending on whether random or hub genes are disrupted. This difference indicates that the power-law distribution of regulatory interactions has biological significance, making random disruptions less deleterious to organisms facing environmental stress. That \textit{E. coli} can reduce the effect of environmental stress without reducing the phenotypic effect of additional mutations, indicates that robustness and evolvability need not be antagonistic.},
keywords = {Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}
Genome-wide profiling has allowed the regulatory interaction networks of many organisms to be visualised and the pattern of connections between genes to be studied. These networks are non-random, following a power-law distribution with a small number of well-connected 'hubs' and many genes with only one or a few connections. Theoretical work predicts that power-law networks display several unique properties. One of the most biologically interesting of these is an intrinsic robustness to disturbance such that removal of a random gene will have little effect on network function. Conversely, targeted removal of a hub gene is expected to have a large effect.
Results
We compared the response of Escherichia coli to environmental and mutational stress following disruption of random or hub genes. We found that disruption of random genes had less effect on robustness to environmental stress than did the targeted disruption of hub genes. In contrast, random disruption strains were slightly less robust to the effect of mutational stress than were hub disruption strains. When we compared the effect of each disruption on environmental and mutational stress, we found a negative relationship, such that strains that were more environmentally robust tended to be less robust to mutational stress.
Conclusion
Our results demonstrate that mutant strains of E. coli respond differently to stress, depending on whether random or hub genes are disrupted. This difference indicates that the power-law distribution of regulatory interactions has biological significance, making random disruptions less deleterious to organisms facing environmental stress. That E. coli can reduce the effect of environmental stress without reducing the phenotypic effect of additional mutations, indicates that robustness and evolvability need not be antagonistic.
Pelosi L; Kühn L; Guetta D; Garin J; Geiselmann J; Lenski R E; Schneider D
Parallel Changes in Global Protein Profiles During Long-Term Experimental Evolution in Escherichia coli Journal Article
Genetics, 173 (4), pp. 1851-1869, 2006, ISSN: 1943-2631.
Abstract | Links | BibTeX | Altmetric | Tags: Genotypes and Phenotypes, Parallelism and Divergence
@article{nokey,
title = {Parallel Changes in Global Protein Profiles During Long-Term Experimental Evolution in \textit{Escherichia coli}},
author = {Ludovic Pelosi and Lauriane Kühn and Dorian Guetta and Jérôme Garin and Johannes Geiselmann and Richard E. Lenski and Dominique Schneider},
url = {https://academic.oup.com/genetics/article/173/4/1851/6061068},
doi = {10.1534/genetics.105.049619},
issn = {1943-2631},
year = {2006},
date = {2006-08-01},
urldate = {2006-08-01},
journal = {Genetics},
volume = {173},
number = {4},
pages = {1851-1869},
abstract = {Twelve populations of \textit{Escherichia coli} evolved in and adapted to a glucose-limited environment from a common ancestor. We used two-dimensional protein electrophoresis to compare two evolved clones, isolated from independently derived populations after 20,000 generations. Exceptional parallelism was detected. We compared the observed changes in protein expression profiles with previously characterized global transcription profiles of the same clones; this is the first time such a comparison has been made in an evolutionary context where these changes are often quite subtle. The two methodologies exhibited some remarkable similarities that highlighted two different levels of parallel regulatory changes that were beneficial during the evolution experiment. First, at the higher level, both methods revealed extensive parallel changes in the same global regulatory network, reflecting the involvement of beneficial mutations in genes that control the ppGpp regulon. Second, both methods detected expression changes of identical gene sets that reflected parallel changes at a lower level of gene regulation. The protein profiles led to the discovery of beneficial mutations affecting the \textit{malT} gene, with strong genetic parallelism across independently evolved populations. Functional and evolutionary analyses of these mutations revealed parallel phenotypic decreases in the maltose regulon expression and a high level of polymorphism at this locus in the evolved populations.},
keywords = {Genotypes and Phenotypes, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
Woods R J; Schneider D; Winkworth C L; Riley M A; Lenski R E
Tests of parallel molecular evolution in a long-term experiment with Escherichia coli Journal Article
Proceedings of the National Academy of Sciences of the United States of America, 103 (24), pp. 9107–9112, 2006, ISSN: 0027-8424.
Abstract | Links | BibTeX | Altmetric | Tags: Genome Evolution, Genotypes and Phenotypes, Parallelism and Divergence
@article{Woods2006,
title = {Tests of parallel molecular evolution in a long-term experiment with \textit{Escherichia coli}},
author = {Robert J. Woods and Dominique Schneider and Cynthia L. Winkworth and Margaret A. Riley and Richard E. Lenski},
url = {http://www.pnas.org/cgi/doi/10.1073/pnas.0602917103},
doi = {10.1073/pnas.0602917103},
issn = {0027-8424},
year = {2006},
date = {2006-06-01},
urldate = {2006-06-01},
journal = {Proceedings of the National Academy of Sciences of the United States of America},
volume = {103},
number = {24},
pages = {9107--9112},
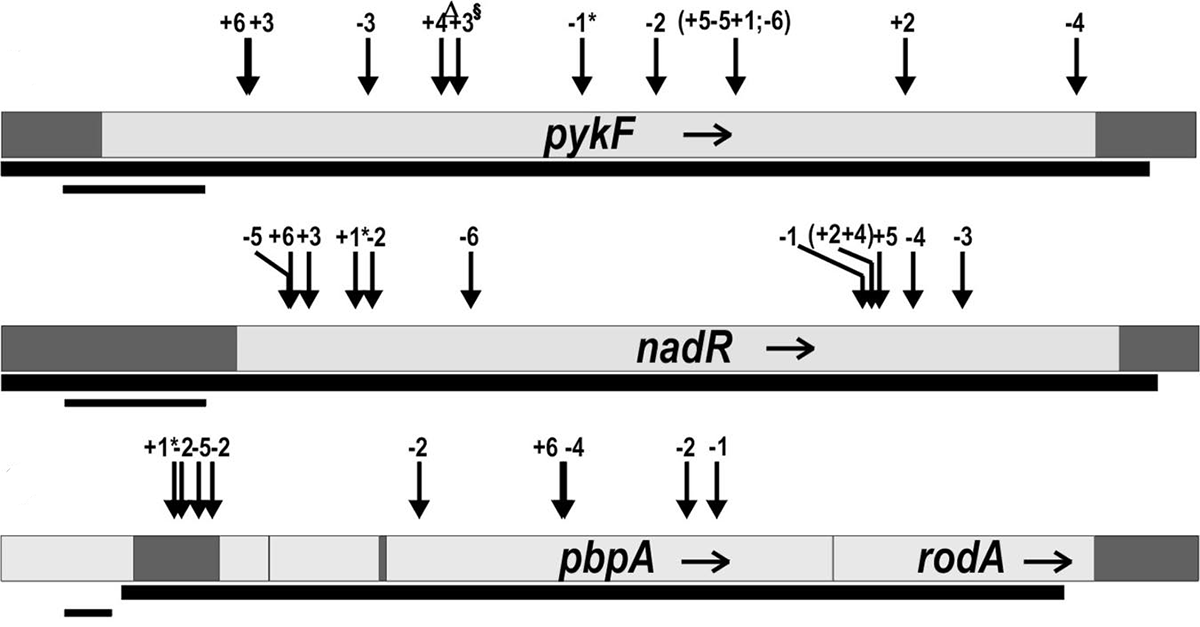
abstract = {The repeatability of evolutionary change is difficult to quantify because only a single outcome can usually be observed for any precise set of circumstances. In this study, however, we have quantified the frequency of parallel and divergent genetic changes in 12 initially identical populations of \textit{Escherichia coli} that evolved in identical environments for 20,000 cell generations. Unlike previous analyses in which candidate genes were identified based on parallel phenotypic changes, here we sequenced four loci (\textit{pykF}, \textit{nadR}, \textit{pbpA}-\textit{rodA}, and \textit{hokB}/\textit{sokB}) in which mutations of unknown effect had been discovered in one population, and then we compared the substitution pattern in these "blind" candidate genes with the pattern found in 36 randomly chosen genes. Two candidate genes, \textit{pykF} and \textit{nadR}, had substitutions in all 11 other populations, and the other 2 in several populations. There were very few cases, however, in which the exact same mutations were substituted, in contrast to the findings from conceptually related work performed with evolving virus populations. No random genes had any substitutions except in four populations that evolved defects in DNA repair. Tests of four different statistical aspects of the pattern of molecular evolution all indicate that adaptation by natural selection drove the parallel changes in these candidate genes.},
keywords = {Genome Evolution, Genotypes and Phenotypes, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
2005
Crozat E; Philippe N; Lenski R E; Geiselmann J; Schneider D
Long-Term Experimental Evolution in Escherichia coli. XII. DNA Topology as a Key Target of Selection Journal Article
Genetics, 169 (2), pp. 523-532, 2005, ISSN: 1943-2631.
Abstract | Links | BibTeX | Altmetric | Tags: Genotypes and Phenotypes, Parallelism and Divergence
@article{nokey,
title = {Long-Term Experimental Evolution in \textit{Escherichia coli}. XII. DNA Topology as a Key Target of Selection},
author = {Estelle Crozat and Nadège Philippe and Richard E. Lenski and Johannes Geiselmann and Dominique Schneider},
url = {https://academic.oup.com/genetics/article/169/2/523/6060318},
doi = {10.1534/genetics.104.035717},
issn = {1943-2631},
year = {2005},
date = {2005-02-01},
urldate = {2005-02-01},
journal = {Genetics},
volume = {169},
number = {2},
pages = {523-532},
abstract = {The genetic bases of adaptation are being investigated in 12 populations of \textit{Escherichia coli}, founded from a common ancestor and serially propagated for 20,000 generations, during which time they achieved substantial fitness gains. Each day, populations alternated between active growth and nutrient exhaustion. DNA supercoiling in bacteria is influenced by nutritional state, and DNA topology helps coordinate the overall pattern of gene expression in response to environmental changes. We therefore examined whether the genetic controls over supercoiling might have changed during the evolution experiment. Parallel changes in topology occurred in most populations, with the level of DNA supercoiling increasing, usually in the first 2000 generations. Two mutations in the \textit{topA} and \textit{fis} genes that control supercoiling were discovered in a population that served as the focus for further investigation. Moving the mutations, alone and in combination, into the ancestral background had an additive effect on supercoiling, and together they reproduced the net change in DNA topology observed in this population. Moreover, both mutations were beneficial in competition experiments. Clonal interference involving other beneficial DNA topology mutations was also detected. These findings define a new class of fitness-enhancing mutations and indicate that the control of DNA supercoiling can be a key target of selection in evolving bacterial populations.},
keywords = {Genotypes and Phenotypes, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
2003
Cooper T F; Rozen D E; Lenski R E
Parallel changes in gene expression after 20,000 generations of evolution in Escherichia coli Journal Article
Proceedings of the National Academy of Sciences of the United States of America, 100 (3), pp. 1072–1077, 2003, ISSN: 0027-8424.
Abstract | Links | BibTeX | Altmetric | Tags: Genotypes and Phenotypes, Parallelism and Divergence
@article{Cooper2003,
title = {Parallel changes in gene expression after 20,000 generations of evolution in \textit{Escherichia coli}},
author = {Tim F. Cooper and Daniel E. Rozen and Richard E. Lenski},
url = {http://www.pnas.org/cgi/doi/10.1073/pnas.0334340100},
doi = {10.1073/pnas.0334340100},
issn = {0027-8424},
year = {2003},
date = {2003-02-01},
urldate = {2003-02-01},
journal = {Proceedings of the National Academy of Sciences of the United States of America},
volume = {100},
number = {3},
pages = {1072--1077},
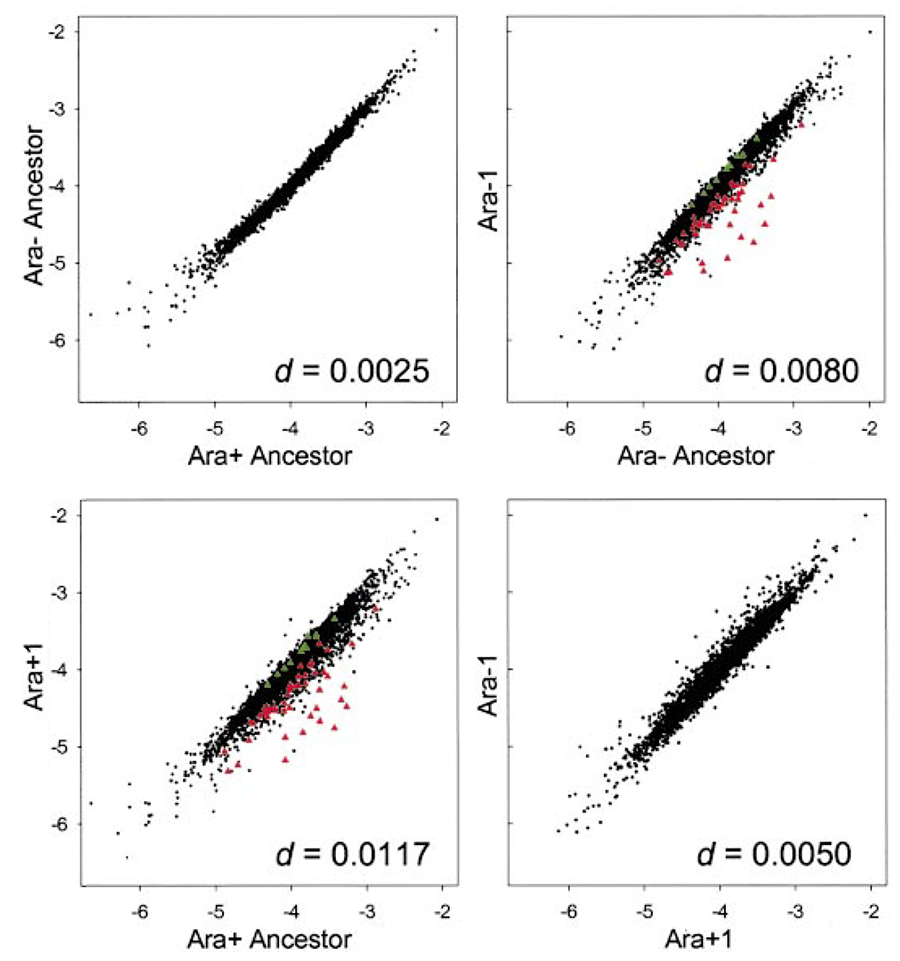
abstract = {Twelve populations of \textit{Escherichia coli}, derived from a common ancestor, evolved in a glucose-limited medium for 20,000 generations. Here we use DNA expression arrays to examine whether gene-expression profiles in two populations evolved in parallel, which would indicate adaptation, and to gain insight into the mechanisms underlying their adaptation. We compared the expression profile of the ancestor to that of clones sampled from both populations after 20,000 generations. The expression of 59 genes had changed significantly in both populations. Remarkably, all 59 were changed in the same direction relative to the ancestor. Many of these genes were members of the cAMP-cAMP receptor protein (CRP) and guanosine tetraphosphate (ppGpp) regulons. Sequencing of several genes controlling the effectors of these regulons found a nonsynonymous mutation in \textit{spoT} in one population. Moving this mutation into the ancestral background showed that it increased fitness and produced many of the expression changes manifest after 20,000 generations. The same mutation had no effect on fitness when introduced into the other evolved population, indicating that a mutation of similar effect was present already. Our study demonstrates the utility of expression arrays for addressing evolutionary issues including the quantitative measurement of parallel evolution in independent lineages and the identification of beneficial mutations.},
keywords = {Genotypes and Phenotypes, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
2001
Cooper V S; Schneider D; Blot M; Lenski R E
Mechanisms causing rapid and parallel losses of ribose catabolism in evolving populations of Escherichia coli B Journal Article
Journal of Bacteriology, 183 (9), pp. 2834-2841, 2001, ISSN: 0021-9193.
Abstract | Links | BibTeX | Altmetric | Tags: Correlated Responses, Genotypes and Phenotypes
@article{nokey,
title = {Mechanisms causing rapid and parallel losses of ribose catabolism in evolving populations of \textit{Escherichia coli} B},
author = {Vaughn S. Cooper and Dominique Schneider and Michel Blot and Richard E. Lenski},
url = {https://journals.asm.org/doi/full/10.1128/JB.183.9.2834-2841.2001},
doi = {10.1128/JB.183.9.2834-2841.2001},
issn = {0021-9193},
year = {2001},
date = {2001-05-01},
urldate = {2001-05-01},
journal = {Journal of Bacteriology},
volume = {183},
number = {9},
pages = {2834-2841},
abstract = {Twelve populations of \textit{Escherichia coli} B all lost d -ribose catabolic function during 2,000 generations of evolution in glucose minimal medium. We sought to identify the population genetic processes and molecular genetic events that caused these rapid and parallel losses. Seven independent Rbs ^{−} mutants were isolated, and their competitive fitnesses were measured relative to that of their Rbs ^{+} progenitor. These Rbs ^{−} mutants were all about 1 to 2% more fit than the progenitor. A fluctuation test revealed an unusually high rate, about 5 × 10 ^{−5} per cell generation, of mutation from Rbs ^{+} to Rbs ^{−}, which contributed to rapid fixation. At the molecular level, the loss of ribose catabolic function involved the deletion of part or all of the ribose operon (\textit{rbs} genes). The physical extent of the deletion varied between mutants, but each deletion was associated with an IS\textit{150} element located immediately upstream of the \textit{rbs} operon. The deletions apparently involved transposition into various locations within the \textit{rbs} operon; recombination between the new IS\textit{150} copy and the one upstream of the \textit{rbs} operon then led to the deletion of the intervening sequence. To confirm that the beneficial fitness effect was caused by deletion of the \textit{rbs} operon (and not some undetected mutation elsewhere), we used P1 transduction to restore the functional \textit{rbs} operon to two Rbs^{−} mutants, and we constructed another Rbs^{−} strain by gene replacement with a deletion not involving IS\textit{150}. All three of these new constructs confirmed that Rbs^{−} mutants have a competitive advantage relative to their Rbs^{+} counterparts in glucose minimal medium. The rapid and parallel evolutionary losses of ribose catabolic function thus involved both (i) an unusually high mutation rate, such that Rbs^{−} mutants appeared repeatedly in all populations, and (ii) a selective advantage in glucose minimal medium that drove these mutants to fixation.},
keywords = {Correlated Responses, Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}
2000
Schneider D; Duperchy E; Coursange E; Lenski R E; Blot M
Long-Term Experimental Evolution in Escherichia coli. IX. Characterization of Insertion Sequence-Mediated Mutations and Rearrangements Journal Article
Genetics, 156 (2), pp. 477–488, 2000, ISSN: 1943-2631.
Abstract | Links | BibTeX | Altmetric | Tags: Genome Evolution, Genotypes and Phenotypes
@article{Schneider2000,
title = {Long-Term Experimental Evolution in \textit{Escherichia coli}. IX. Characterization of Insertion Sequence-Mediated Mutations and Rearrangements},
author = {Dominique Schneider and Esther Duperchy and Evelyne Coursange and Richard E. Lenski and Michel Blot},
url = {https://academic.oup.com/genetics/article/156/2/477/6051340},
doi = {10.1093/genetics/156.2.477},
issn = {1943-2631},
year = {2000},
date = {2000-10-01},
urldate = {2000-10-01},
journal = {Genetics},
volume = {156},
number = {2},
pages = {477--488},
abstract = {As part of a long-term evolution experiment, two populations of \textit{Escherichia coli} B adapted to a glucose minimal medium for 10,000 generations. In both populations, multiple IS-associated mutations arose that then went to fixation. We identify the affected genetic loci and characterize the molecular events that produced nine of these mutations. All nine were IS-mediated events, including simple insertions as well as recombination between homologous elements that generated inversions and deletions. Sequencing DNA adjacent to the insertions indicates that the affected genes are involved in central metabolism (knockouts of \textit{pykF} and \textit{nadR}), cell wall synthesis (adjacent to the promoter of \textit{pbpA-rodA}), and ill-defined functions (knockouts of \textit{hokB-sokB} and \textit{yfcU}). These genes are candidates for manipulation and competition experiments to determine whether the mutations were beneficial or merely hitchhiked to fixation.},
keywords = {Genome Evolution, Genotypes and Phenotypes},
pubstate = {published},
tppubtype = {article}
}
1997
Sniegowski P D; Gerrish P J; Lenski R E
Evolution of high mutation rates in experimental populations of E. coli Journal Article
Nature, 387 (6634), pp. 703–705, 1997, ISSN: 0028-0836.
Abstract | Links | BibTeX | Altmetric | Tags: Genotypes and Phenotypes, Mutation Rates
@article{Sniegowski1997,
title = {Evolution of high mutation rates in experimental populations of \textit{E. coli}},
author = {Paul D. Sniegowski and Philip J. Gerrish and Richard E. Lenski},
url = {http://www.nature.com/articles/42701},
doi = {10.1038/42701},
issn = {0028-0836},
year = {1997},
date = {1997-06-01},
urldate = {1997-06-01},
journal = {Nature},
volume = {387},
number = {6634},
pages = {703--705},
abstract = {Most mutations are likely to be deleterious, and so the spontaneous mutation rate is generally held at a very low value. Nonetheless, evolutionary theory predicts that high mutation rates can evolve under certain circumstances. Empirical observations have previously been limited to short-term studies of the fates of mutator strains deliberately introduced into laboratory populations of \textit{Escherichia coli}, and to the effects of intense selective events on mutator frequencies in \textit{E. coli}. Here we report the rise of spontaneously originated mutators in populations of \textit{E. coli} undergoing long-term adaptation to a new environment. Our results corroborate computer simulations of mutator evolution in adapting clonal populations, and may help to explain observations that associate high mutation rates with emerging pathogens and with certain cancers.},
keywords = {Genotypes and Phenotypes, Mutation Rates},
pubstate = {published},
tppubtype = {article}
}
1996
Travisano M; Lenski R E
Long-Term Experimental Evolution in Escherichia coli. IV. Targets of Selection and the Specificity of Adaptation Journal Article
Genetics, 143 (1), pp. 15–26, 1996, ISSN: 1943-2631.
Abstract | Links | BibTeX | Altmetric | Tags: Correlated Responses, Genotypes and Phenotypes, Parallelism and Divergence
@article{Travisano1996,
title = {Long-Term Experimental Evolution in \textit{Escherichia coli}. IV. Targets of Selection and the Specificity of Adaptation},
author = {Michael Travisano and Richard E. Lenski},
url = {https://academic.oup.com/genetics/article/143/1/15/6016887},
doi = {10.1093/genetics/143.1.15},
issn = {1943-2631},
year = {1996},
date = {1996-05-01},
urldate = {1996-05-01},
journal = {Genetics},
volume = {143},
number = {1},
pages = {15--26},
abstract = {This study investigates the physiological manifestation of adaptive evolutionary change in 12 replicate populations of \textit{Escherichia coli} that were propagated for 2000 generations in a glucose-limited environment. Representative genotypes from each population were assayed for fitness relative to their common ancestor in the experimental glucose environment and in 11 novel single-nutrient environments. After 2000 generations, the 12 derived genotypes had diverged into at least six distinct phenotypic classes. The nutrients were classified into four groups based upon their uptake physiology. All 12 derived genotypes improved in fitness by similar amounts in the glucose environment, and this pattern of parallel fitness gains was also seen in those novel environments where the limiting nutrient shared uptake mechanisms with glucose. Fitness showed little or no consistent improvement, but much greater genetic variation, in novel environments where the limiting nutrient differed from glucose in its uptake mechanisms. This pattern of fitness variation in the novel nutrient environments suggests that the independently derived genotypes adapted to the glucose environment by similar, but not identical, changes in the physiological mechanisms for moving glucose across both the inner and outer membranes.},
keywords = {Correlated Responses, Genotypes and Phenotypes, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}
1995
Travisano M; Vasi F K; Lenski R E
Long-Term Experimental Evolution in Escherichia coli. III. Variation Among Replicate Populations in Correlated Responses to Novel Environments Journal Article
Evolution, 49 (1), pp. 189–200, 1995.
Abstract | Links | BibTeX | Altmetric | Tags: Correlated Responses, Genotypes and Phenotypes, Parallelism and Divergence
@article{Travisano1995b,
title = {Long-Term Experimental Evolution in \textit{Escherichia coli}. III. Variation Among Replicate Populations in Correlated Responses to Novel Environments},
author = {Michael Travisano and Farida K. Vasi and Richard E. Lenski},
url = {https://onlinelibrary.wiley.com/doi/10.1111/j.1558-5646.1995.tb05970.x},
doi = {https://doi.org/10.1111/j.1558-5646.1995.tb05970.x},
year = {1995},
date = {1995-01-01},
urldate = {1995-01-01},
journal = {Evolution},
volume = {49},
number = {1},
pages = {189--200},
abstract = {Twelve populations of \textit{Escherichia coli} were founded from a single clone and propagated for 2000 gen- erations in identical glucose-limited environments. During this time, the mean fitnesses of the evolving populations relative to their common ancestor improved greatly, but their fitnesses relative to one another diverged only slightly. Although the populations showed similar fitness increases, they may have done so by different underlying adaptations, or they may have diverged in other respects by random genetic drift. Therefore, we examined the relative fitnesses of independently derived genotypes in two other sugars, maltose and lactose, to determine whether they were ho- mogeneous or heterogeneous in these environments. The genetic variation among the derived lines in fitness on maltose and lactose was more than 100-times greater than their variation in fitness on glucose. Moreover, the glucose-adapted genotypes, on average, showed significant adaptation to lactose, but not to maltose. That pathways for use of maltose and glucose are virtually identical in \textit{E. coli}, except for their distinct mechanisms of uptake, suggests that the derived genotypes have adapted primarily by improved glucose transport. From consideration of the number of generations of divergence, the mutation rate in \textit{E. coli}, and the proportion of its genome required for growth on maltose (but not glucose), we hypothesize that pleiotropy involving the selected alleles, rather than random genetic drift of alleles at other loci, was the major cause of the variation among the derived genotypes in fitness on these other sugars.},
keywords = {Correlated Responses, Genotypes and Phenotypes, Parallelism and Divergence},
pubstate = {published},
tppubtype = {article}
}